diff --git a/.Rbuildignore b/.Rbuildignore
index 1da9fa4..1e87b4d 100644
--- a/.Rbuildignore
+++ b/.Rbuildignore
@@ -39,3 +39,4 @@
^man/figures/relabel-workflow\.png$
^man/figures/sample-browser\.png$
^man/figures/settings-dialog\.png$
+^man/figures/class-review\.png$
diff --git a/DESCRIPTION b/DESCRIPTION
index 6c26539..8cb3961 100644
--- a/DESCRIPTION
+++ b/DESCRIPTION
@@ -1,6 +1,6 @@
Package: ClassiPyR
Title: A Shiny App for Manual Image Classification and Validation of IFCB Data
-Version: 0.1.1
+Version: 0.2.0
Authors@R: c(
person("Anders", "Torstensson", email = "anders.torstensson@smhi.se", role = c("aut", "cre"),
comment = c("Swedish Meteorological and Hydrological Institute", ORCID = "0000-0002-8283-656X")),
@@ -9,9 +9,10 @@ Authors@R: c(
)
Description: A Shiny application for manual classification and validation of
Imaging FlowCytobot (IFCB) plankton images. Supports loading classifications
- from CSV files or MATLAB classifier output, drag-select for batch operations,
- and exports classifications in MATLAB-compatible format for use with the
- 'ifcb-analysis' toolbox.
+ from CSV, HDF5 (.h5), or MATLAB classifier output, directly from remote
+ IFCB Dashboard instances, or via live CNN prediction through a Gradio API.
+ Features drag-select for batch operations and exports classifications in
+ MATLAB-compatible format for use with the 'ifcb-analysis' toolbox.
License: MIT + file LICENSE
Encoding: UTF-8
LazyData: true
@@ -20,7 +21,8 @@ Imports:
shinyjs,
shinyFiles,
bslib,
- iRfcb,
+ curl,
+ iRfcb (>= 0.8.1),
dplyr,
DT,
jsonlite,
@@ -30,6 +32,7 @@ Imports:
Suggests:
testthat (>= 3.0.0),
covr,
+ hdf5r,
knitr,
rmarkdown,
pkgdown
diff --git a/NAMESPACE b/NAMESPACE
index 9577f2a..7c6d311 100644
--- a/NAMESPACE
+++ b/NAMESPACE
@@ -3,12 +3,19 @@
export(copy_images_to_class_folders)
export(create_empty_changes_log)
export(create_new_classifications)
+export(download_dashboard_adc)
+export(download_dashboard_autoclass)
+export(download_dashboard_image_single)
+export(download_dashboard_images)
+export(download_dashboard_images_bulk)
+export(download_dashboard_images_individual)
export(export_all_db_to_mat)
export(export_all_db_to_png)
export(export_db_to_mat)
export(export_db_to_png)
export(filter_to_extracted)
export(get_config_dir)
+export(get_dashboard_cache_dir)
export(get_db_path)
export(get_default_db_dir)
export(get_file_index_path)
@@ -16,24 +23,33 @@ export(get_sample_paths)
export(get_settings_path)
export(import_all_mat_to_db)
export(import_mat_to_db)
+export(import_png_folder_to_db)
export(init_python_env)
export(is_valid_sample_name)
export(list_annotated_samples_db)
+export(list_annotation_metadata_db)
+export(list_classes_db)
+export(list_dashboard_bins)
export(load_annotations_db)
+export(load_class_annotations_db)
export(load_class_list)
export(load_file_index)
export(load_from_classifier_mat)
export(load_from_csv)
export(load_from_db)
+export(load_from_h5)
export(load_from_mat)
+export(parse_dashboard_url)
export(read_roi_dimensions)
export(rescan_file_index)
export(run_app)
export(sanitize_string)
export(save_annotations_db)
+export(save_class_review_changes_db)
export(save_file_index)
export(save_sample_annotations)
export(save_validation_statistics)
+export(scan_png_class_folder)
export(update_annotator)
importFrom(DBI,dbConnect)
importFrom(DBI,dbDisconnect)
@@ -43,9 +59,13 @@ importFrom(DBI,dbWriteTable)
importFrom(DT,renderDT)
importFrom(RSQLite,SQLite)
importFrom(bslib,bs_theme)
+importFrom(curl,curl_fetch_disk)
+importFrom(curl,curl_fetch_memory)
+importFrom(curl,new_handle)
importFrom(dplyr,filter)
importFrom(iRfcb,ifcb_annotate_samples)
importFrom(iRfcb,ifcb_create_manual_file)
+importFrom(iRfcb,ifcb_download_dashboard_data)
importFrom(iRfcb,ifcb_extract_pngs)
importFrom(iRfcb,ifcb_get_mat_variable)
importFrom(jsonlite,fromJSON)
diff --git a/NEWS.md b/NEWS.md
index 2db4d50..3189cca 100644
--- a/NEWS.md
+++ b/NEWS.md
@@ -1,3 +1,27 @@
+# ClassiPyR 0.2.0
+
+## New features
+
+- **Live Prediction**: Added a "Predict" button in Sample Mode that classifies all images in the loaded sample using a remote CNN model via `iRfcb::ifcb_classify_images()`. Configure the Gradio API URL and model in Settings > Live Prediction. The model dropdown is populated dynamically from the Gradio server. Predictions respect the classification threshold setting, skip manually reclassified images, and new class names from the model are added to the class list automatically. A per-image progress bar shows classification progress.
+- **IFCB Dashboard support**: Connect directly to remote IFCB Dashboard instances (e.g. `https://habon-ifcb.whoi.edu/`) without downloading data locally. Toggle between "Local Folders" and "IFCB Dashboard" in Settings, enter a Dashboard URL (with optional `?dataset=` parameter), and browse samples from the API. Images are downloaded on demand and cached locally. Optionally load dashboard auto-classifications for validation mode. Supports MAT export by downloading ADC files on demand, with graceful fallback to SQLite-only when ADC is unavailable.
+- **Dashboard class review optimization**: Class review mode with a dashboard source now downloads individual PNG images instead of entire zip archives, making it much faster when reviewing a single class across many samples.
+- **Configurable dashboard download settings**: Dashboard mode now exposes parallel downloads, sleep time, timeout, and max retries in an "Advanced Download Settings" section in Settings. Previously these were hardcoded.
+- **Local classification files in dashboard mode**: The Classification Folder setting is now available in dashboard mode. When configured, local CSV/H5/MAT classification files take priority over dashboard auto-classifications, with dashboard autoclass as a fallback.
+- New exported functions: `parse_dashboard_url()`, `list_dashboard_bins()`, `download_dashboard_images()`, `download_dashboard_images_bulk()`, `download_dashboard_image_single()`, `download_dashboard_images_individual()`, `download_dashboard_adc()`, `download_dashboard_autoclass()`, and `get_dashboard_cache_dir()` for programmatic dashboard access.
+- **Class Review Mode**: View and reclassify all annotated images of a specific class across the entire database. Switch to class review via the mode toggle in the sidebar, select a class, and load all matching images from all samples at once. Changes are saved as row-level updates to the database.
+- New exported functions: `list_classes_db()`, `load_class_annotations_db()`, and `save_class_review_changes_db()` for programmatic class review operations.
+- Added **Import PNG → SQLite** button in Settings > Import / Export. Imports annotations from a folder of PNG images organized in class-name subfolders (e.g. exported by ClassiPyR or other tools). Folder names follow the iRfcb convention where trailing `_NNN` suffixes are stripped.
+- When importing PNG folders with class names not in the current class list, a **class mapping dialog** lets users remap unmatched classes to existing ones or add them as new classes.
+- Overwrite warning dialog shown when imported samples already exist in the database.
+- New exported functions: `scan_png_class_folder()` for scanning PNG class folder structures, and `import_png_folder_to_db()` for programmatic bulk import.
+- **HDF5 classification support**: Load classifications from `.h5` files produced by [iRfcb](https://github.com/EuropeanIFCBGroup/iRfcb) (>= 0.8.0). Requires the optional `hdf5r` package.
+- **Classification threshold toggle**: New "Apply classification threshold" checkbox in Settings controls whether thresholded or raw predictions are used, for all classification formats (CSV, H5, MAT).
+- **Skip class from PNG export**: New option in Settings to exclude a specific class (e.g. "unclassified") from PNG output.
+
+## UI improvements
+
+- The **class list editor** now shows the number of annotated images per class in parentheses, queried from the SQLite database.
+
# ClassiPyR 0.1.1
## New features
diff --git a/R/dashboard.R b/R/dashboard.R
new file mode 100644
index 0000000..7be086a
--- /dev/null
+++ b/R/dashboard.R
@@ -0,0 +1,563 @@
+# Dashboard functions for ClassiPyR
+#
+# Functions for fetching sample lists and images from remote IFCB Dashboard
+# instances (e.g., https://habon-ifcb.whoi.edu/).
+
+#' @importFrom jsonlite fromJSON
+#' @importFrom curl curl_fetch_memory curl_fetch_disk new_handle
+#' @importFrom iRfcb ifcb_download_dashboard_data
+NULL
+
+#' Get persistent cache directory for dashboard downloads
+#'
+#' Returns the path to the dashboard cache directory. During R CMD check,
+#' uses a temporary directory.
+#'
+#' @return Path to the dashboard cache directory
+#' @export
+#' @examples
+#' cache_dir <- get_dashboard_cache_dir()
+#' print(cache_dir)
+get_dashboard_cache_dir <- function() {
+ if (nzchar(Sys.getenv("_R_CHECK_PACKAGE_NAME_", ""))) {
+ return(file.path(tempdir(), "ClassiPyR", "dashboard"))
+ }
+ file.path(tools::R_user_dir("ClassiPyR", "cache"), "dashboard")
+}
+
+#' Parse an IFCB Dashboard URL
+#'
+#' Extracts the base URL and optional dataset name from a Dashboard URL.
+#'
+#' @param url Character. A Dashboard URL, e.g.
+#' \code{"https://habon-ifcb.whoi.edu/"} or
+#' \code{"https://habon-ifcb.whoi.edu/timeline?dataset=tangosund"}.
+#' @return A list with \code{base_url} (without trailing slash) and
+#' \code{dataset_name} (character or NULL).
+#' @export
+#' @examples
+#' parse_dashboard_url("https://habon-ifcb.whoi.edu/")
+#' parse_dashboard_url("https://habon-ifcb.whoi.edu/timeline?dataset=tangosund")
+parse_dashboard_url <- function(url) {
+ if (is.null(url) || !is.character(url) || length(url) != 1 || !nzchar(url)) {
+ stop("url must be a non-empty character string")
+ }
+ if (!grepl("^https?://", url)) {
+ stop("url must start with http:// or https://")
+ }
+
+ # Extract dataset from query parameter ?dataset=xxx
+ dataset_name <- NULL
+ query_match <- regmatches(url, regexpr("[?&]dataset=([^&#]+)", url))
+ if (length(query_match) == 1 && nzchar(query_match)) {
+ dataset_name <- sub("^[?&]dataset=", "", query_match)
+ }
+
+ # Strip query string and path components (timeline, etc.) to get base URL
+ base_url <- sub("[?].*$", "", url)
+ base_url <- sub("/timeline/?$", "", base_url)
+ base_url <- sub("/+$", "", base_url)
+
+ list(base_url = base_url, dataset_name = dataset_name)
+}
+
+#' List bins from an IFCB Dashboard
+#'
+#' Fetches the bin list from the Dashboard API. This is a vendored copy of
+#' \code{iRfcb::ifcb_list_dashboard_bins()} from the development version that
+#' supports the \code{dataset_name} parameter.
+#'
+#' @param base_url Character. Base URL (e.g. \code{"https://habon-ifcb.whoi.edu"}).
+#' @param dataset_name Optional character. Dataset slug (e.g. \code{"tangosund"}).
+#' @return Character vector of bin (sample) names.
+#' @export
+#' @examples
+#' \donttest{
+#' bins <- list_dashboard_bins("https://ifcb-data.whoi.edu", "mvco")
+#' }
+# TODO: Replace with iRfcb::ifcb_list_dashboard_bins() once iRfcb >= 0.9.0
+# ships dataset_name support.
+list_dashboard_bins <- function(base_url, dataset_name = NULL) {
+ base_url <- sub("/+$", "", base_url)
+
+ api_url <- paste0(base_url, "/api/list_bins")
+
+ if (!is.null(dataset_name) && nzchar(dataset_name)) {
+ dataset_name <- utils::URLencode(dataset_name, reserved = TRUE)
+ api_url <- paste0(api_url, "?dataset=", dataset_name)
+ }
+
+ response <- tryCatch(
+ curl::curl_fetch_memory(api_url,
+ handle = curl::new_handle(httpheader = c(Accept = "application/json"))),
+ error = function(e) stop("Failed to connect to IFCB Dashboard API: ", e$message)
+ )
+
+ if (response$status_code != 200) {
+ stop("API request failed [", response$status_code, "]: ", api_url)
+ }
+
+ json_content <- rawToChar(response$content)
+ Encoding(json_content) <- "UTF-8"
+
+ parsed <- tryCatch(
+ jsonlite::fromJSON(json_content, flatten = TRUE),
+ error = function(e) stop("Failed to parse JSON content: ", e$message)
+ )
+
+ # The API returns a list with one element containing a data frame with a
+ # "pid" column (or similar). Extract the sample names.
+ if (is.data.frame(parsed)) {
+ bins <- parsed[[1]]
+ } else if (is.list(parsed) && length(parsed) > 0) {
+ first <- parsed[[1]]
+ if (is.data.frame(first)) {
+ bins <- first[[1]]
+ } else {
+ bins <- as.character(first)
+ }
+ } else {
+ bins <- as.character(parsed)
+ }
+
+ as.character(bins)
+}
+
+#' Download and extract PNG images from the Dashboard
+#'
+#' Downloads a zip file of PNG images for a sample from the Dashboard.
+#' Extracts into the cache directory. Skips re-download if PNGs already exist.
+#'
+#' @param base_url Character. Dashboard base URL.
+#' @param sample_name Character. Sample name (bin PID).
+#' @param cache_dir Character. Cache directory. Defaults to
+#' \code{\link{get_dashboard_cache_dir}()}.
+#' @param parallel_downloads Integer. Number of parallel downloads.
+#' @param sleep_time Numeric. Seconds to sleep between download batches.
+#' @param multi_timeout Numeric. Timeout in seconds for multi-file downloads.
+#' @param max_retries Integer. Maximum number of retry attempts.
+#' @return Path to the folder containing extracted PNGs, or NULL on failure.
+#' @export
+download_dashboard_images <- function(base_url, sample_name,
+ cache_dir = get_dashboard_cache_dir(),
+ parallel_downloads = 5, sleep_time = 2,
+ multi_timeout = 120, max_retries = 3) {
+ # Expected path structure: cache_dir/sample_name/sample_name/*.png
+ png_folder <- file.path(cache_dir, sample_name)
+ png_subfolder <- file.path(png_folder, sample_name)
+
+ # Check if PNGs already exist in cache
+ if (dir.exists(png_subfolder)) {
+ existing_pngs <- list.files(png_subfolder, pattern = "\\.png$")
+ if (length(existing_pngs) > 0) {
+ return(png_folder)
+ }
+ }
+
+ dir.create(cache_dir, recursive = TRUE, showWarnings = FALSE)
+
+ # Build the dashboard URL for download
+ # ifcb_download_dashboard_data expects a URL with a path component
+ dashboard_url <- paste0(sub("/+$", "", base_url), "/")
+
+ tryCatch({
+ ifcb_download_dashboard_data(
+ dashboard_url = dashboard_url,
+ samples = sample_name,
+ file_types = "zip",
+ dest_dir = cache_dir,
+ parallel_downloads = parallel_downloads,
+ sleep_time = sleep_time,
+ multi_timeout = multi_timeout,
+ max_retries = max_retries,
+ quiet = TRUE
+ )
+
+ # The download saves to cache_dir/DYYYYMMDD/sample_name.zip
+ # Find the zip file
+ date_part <- substr(sample_name, 1, 9)
+ zip_path <- file.path(cache_dir, date_part, paste0(sample_name, ".zip"))
+
+ if (!file.exists(zip_path)) {
+ # Try alternate location (directly in cache_dir)
+ zip_path <- file.path(cache_dir, paste0(sample_name, ".zip"))
+ }
+
+ if (!file.exists(zip_path)) {
+ warning("Zip file not found after download for: ", sample_name)
+ return(NULL)
+ }
+
+ # Extract to the expected folder structure
+ dir.create(png_subfolder, recursive = TRUE, showWarnings = FALSE)
+ utils::unzip(zip_path, exdir = png_subfolder)
+
+ # Clean up zip file
+ unlink(zip_path)
+ # Also clean up the date folder if empty
+ date_folder <- file.path(cache_dir, date_part)
+ if (dir.exists(date_folder) && length(list.files(date_folder)) == 0) {
+ unlink(date_folder, recursive = TRUE)
+ }
+
+ png_folder
+ }, error = function(e) {
+ warning("Failed to download images for ", sample_name, ": ", e$message)
+ NULL
+ })
+}
+
+#' Download ADC file from the Dashboard
+#'
+#' Downloads the ADC file for a sample from the Dashboard on demand.
+#'
+#' @param base_url Character. Dashboard base URL.
+#' @param sample_name Character. Sample name.
+#' @param cache_dir Character. Cache directory.
+#' @param parallel_downloads Integer. Number of parallel downloads.
+#' @param sleep_time Numeric. Seconds to sleep between download batches.
+#' @param multi_timeout Numeric. Timeout in seconds for multi-file downloads.
+#' @param max_retries Integer. Maximum number of retry attempts.
+#' @return Path to the downloaded ADC file, or NULL on failure.
+#' @export
+download_dashboard_adc <- function(base_url, sample_name,
+ cache_dir = get_dashboard_cache_dir(),
+ parallel_downloads = 5, sleep_time = 2,
+ multi_timeout = 120, max_retries = 3) {
+ date_part <- substr(sample_name, 1, 9)
+ adc_path <- file.path(cache_dir, date_part, paste0(sample_name, ".adc"))
+
+ if (file.exists(adc_path)) {
+ return(adc_path)
+ }
+
+ dashboard_url <- paste0(sub("/+$", "", base_url), "/")
+
+ tryCatch({
+ ifcb_download_dashboard_data(
+ dashboard_url = dashboard_url,
+ samples = sample_name,
+ file_types = "adc",
+ dest_dir = cache_dir,
+ parallel_downloads = parallel_downloads,
+ sleep_time = sleep_time,
+ multi_timeout = multi_timeout,
+ max_retries = max_retries,
+ quiet = TRUE
+ )
+
+ if (file.exists(adc_path)) {
+ return(adc_path)
+ }
+
+ # Try alternate location
+ alt_path <- file.path(cache_dir, paste0(sample_name, ".adc"))
+ if (file.exists(alt_path)) {
+ return(alt_path)
+ }
+
+ NULL
+ }, error = function(e) {
+ warning("Failed to download ADC for ", sample_name, ": ", e$message)
+ NULL
+ })
+}
+
+#' Download and parse autoclass scores from the Dashboard
+#'
+#' Downloads \code{_class_scores.csv} for a sample and extracts the winning
+#' class (column with max score) per ROI.
+#'
+#' @param base_url Character. Dashboard base URL.
+#' @param sample_name Character. Sample name.
+#' @param cache_dir Character. Cache directory.
+#' @param parallel_downloads Integer. Number of parallel downloads.
+#' @param sleep_time Numeric. Seconds to sleep between download batches.
+#' @param multi_timeout Numeric. Timeout in seconds for multi-file downloads.
+#' @param max_retries Integer. Maximum number of retry attempts.
+#' @return Data frame with columns \code{file_name}, \code{class_name},
+#' \code{score}, or NULL on failure.
+#' @export
+download_dashboard_autoclass <- function(base_url, sample_name,
+ cache_dir = get_dashboard_cache_dir(),
+ parallel_downloads = 5, sleep_time = 2,
+ multi_timeout = 120, max_retries = 3) {
+ # The dashboard URL needs to include the dataset path for autoclass
+ dashboard_url <- paste0(sub("/+$", "", base_url), "/")
+
+ tryCatch({
+ ifcb_download_dashboard_data(
+ dashboard_url = dashboard_url,
+ samples = sample_name,
+ file_types = "autoclass",
+ dest_dir = cache_dir,
+ parallel_downloads = parallel_downloads,
+ sleep_time = sleep_time,
+ multi_timeout = multi_timeout,
+ max_retries = max_retries,
+ quiet = TRUE
+ )
+
+ # Find the downloaded CSV file - may have a version suffix
+ csv_pattern <- paste0("^", sample_name, "_class.*\\.csv$")
+ csv_files <- list.files(cache_dir, pattern = csv_pattern, recursive = TRUE,
+ full.names = TRUE)
+
+ if (length(csv_files) == 0) {
+ return(NULL)
+ }
+
+ csv_path <- csv_files[1]
+
+ # Parse the score matrix CSV
+ # Rows = ROIs, columns = class names, values = scores
+ scores <- utils::read.csv(csv_path, check.names = FALSE)
+
+ if (nrow(scores) == 0 || ncol(scores) < 2) {
+ return(NULL)
+ }
+
+ # The first column is typically the ROI identifier
+ # Check if the first column looks like a ROI ID (numeric or sample_NNNNN)
+ first_col <- scores[[1]]
+ class_cols <- if (is.numeric(first_col) || all(grepl("^\\d+$|_\\d+$", as.character(first_col)))) {
+ # First column is ROI identifier, class scores start at column 2
+ names(scores)[-1]
+ } else {
+ names(scores)
+ }
+
+ score_matrix <- as.matrix(scores[, class_cols, drop = FALSE])
+
+ # Extract winning class per ROI
+ max_idx <- apply(score_matrix, 1, which.max)
+ max_scores <- apply(score_matrix, 1, max)
+ winning_classes <- class_cols[max_idx]
+
+ # Extract file names from pid column (e.g., "D20190402T200352_IFCB010_00001")
+ # or fall back to sequential ROI numbers if pid is not available
+ if (is.character(first_col) && all(grepl("_\\d+$", first_col))) {
+ # pid column contains full identifiers — use them directly
+ file_names <- paste0(first_col, ".png")
+ } else if (is.numeric(first_col) || all(grepl("^\\d+$", as.character(first_col)))) {
+ # First column is numeric ROI numbers
+ roi_numbers <- as.integer(first_col)
+ file_names <- sprintf("%s_%05d.png", sample_name, roi_numbers)
+ } else {
+ # Fallback: sequential numbering
+ file_names <- sprintf("%s_%05d.png", sample_name, seq_len(nrow(scores)))
+ }
+
+ data.frame(
+ file_name = file_names,
+ class_name = winning_classes,
+ score = max_scores,
+ stringsAsFactors = FALSE
+ )
+ }, error = function(e) {
+ warning("Failed to download autoclass for ", sample_name, ": ", e$message)
+ NULL
+ })
+}
+
+#' Bulk download zip archives for multiple samples from the Dashboard
+#'
+#' Downloads zip files for all specified samples in a single batched call
+#' to \code{\link[iRfcb]{ifcb_download_dashboard_data}}, leveraging its
+#' built-in parallel download support. Samples already cached are skipped.
+#' After download, zips are extracted and cleaned up.
+#'
+#' @param base_url Character. Dashboard base URL.
+#' @param sample_names Character vector. Sample names to download.
+#' @param cache_dir Character. Cache directory.
+#' @param parallel_downloads Integer. Number of parallel downloads.
+#' @param sleep_time Numeric. Seconds to sleep between download batches.
+#' @param multi_timeout Numeric. Timeout in seconds for multi-file downloads.
+#' @param max_retries Integer. Maximum number of retry attempts.
+#' @return Character vector of sample names that were successfully downloaded
+#' or already cached.
+#' @export
+download_dashboard_images_bulk <- function(base_url, sample_names,
+ cache_dir = get_dashboard_cache_dir(),
+ parallel_downloads = 5, sleep_time = 2,
+ multi_timeout = 120, max_retries = 3) {
+ dir.create(cache_dir, recursive = TRUE, showWarnings = FALSE)
+ dashboard_url <- paste0(sub("/+$", "", base_url), "/")
+
+ # Determine which samples need downloading (not already cached)
+ needs_download <- vapply(sample_names, function(sn) {
+ png_subfolder <- file.path(cache_dir, sn, sn)
+ !(dir.exists(png_subfolder) &&
+ length(list.files(png_subfolder, pattern = "\\.png$")) > 0)
+ }, logical(1))
+
+ to_download <- sample_names[needs_download]
+
+ if (length(to_download) > 0) {
+ tryCatch({
+ ifcb_download_dashboard_data(
+ dashboard_url = dashboard_url,
+ samples = to_download,
+ file_types = "zip",
+ dest_dir = cache_dir,
+ parallel_downloads = parallel_downloads,
+ sleep_time = sleep_time,
+ multi_timeout = multi_timeout,
+ max_retries = max_retries,
+ quiet = TRUE
+ )
+ }, error = function(e) {
+ warning("Bulk zip download failed: ", e$message)
+ })
+
+ # Extract each downloaded zip into the expected folder structure
+ for (sn in to_download) {
+ date_part <- substr(sn, 1, 9)
+ zip_path <- file.path(cache_dir, date_part, paste0(sn, ".zip"))
+
+ if (!file.exists(zip_path)) {
+ zip_path <- file.path(cache_dir, paste0(sn, ".zip"))
+ }
+ if (!file.exists(zip_path)) next
+
+ png_subfolder <- file.path(cache_dir, sn, sn)
+ dir.create(png_subfolder, recursive = TRUE, showWarnings = FALSE)
+ tryCatch(utils::unzip(zip_path, exdir = png_subfolder), error = function(e) NULL)
+ unlink(zip_path)
+
+ # Clean up empty date folder
+ date_folder <- file.path(cache_dir, date_part)
+ if (dir.exists(date_folder) && length(list.files(date_folder)) == 0) {
+ unlink(date_folder, recursive = TRUE)
+ }
+ }
+ }
+
+ # Return all samples that are now cached
+ cached_ok <- vapply(sample_names, function(sn) {
+ png_subfolder <- file.path(cache_dir, sn, sn)
+ dir.exists(png_subfolder) &&
+ length(list.files(png_subfolder, pattern = "\\.png$")) > 0
+ }, logical(1))
+
+ sample_names[cached_ok]
+}
+
+#' Download a single PNG image from the Dashboard
+#'
+#' Downloads one PNG from the Dashboard's \code{/data/} endpoint.
+#' The image is saved to \code{dest_dir/sample_name/file_name}.
+#'
+#' @param base_url Character. Dashboard base URL.
+#' @param sample_name Character. Sample name (bin PID).
+#' @param roi_number Integer. ROI number to download.
+#' @param dest_dir Character. Destination directory.
+#' @param max_retries Integer. Maximum number of retry attempts.
+#' @param timeout Numeric. Request timeout in seconds.
+#' @return File path to the downloaded PNG, or NULL on failure.
+#' @export
+download_dashboard_image_single <- function(base_url, sample_name, roi_number,
+ dest_dir, max_retries = 3,
+ timeout = 15) {
+ file_name <- sprintf("%s_%05d.png", sample_name, roi_number)
+ dest_folder <- file.path(dest_dir, sample_name)
+ dest_path <- file.path(dest_folder, file_name)
+
+ if (file.exists(dest_path)) {
+ return(dest_path)
+ }
+
+ dir.create(dest_folder, recursive = TRUE, showWarnings = FALSE)
+
+ img_url <- paste0(sub("/+$", "", base_url), "/data/", file_name)
+
+ for (attempt in seq_len(max_retries)) {
+ result <- tryCatch({
+ h <- curl::new_handle()
+ curl::handle_setopt(h, connecttimeout = 10, timeout = timeout)
+ response <- curl::curl_fetch_disk(img_url, dest_path, handle = h)
+ if (response$status_code == 200 && file.exists(dest_path) &&
+ file.info(dest_path)$size > 0) {
+ return(dest_path)
+ }
+ # Non-200 status or empty file — no point retrying a 404
+ if (file.exists(dest_path)) unlink(dest_path)
+ if (response$status_code %in% c(404L, 410L)) return(NULL)
+ NULL
+ }, error = function(e) {
+ if (file.exists(dest_path)) unlink(dest_path)
+ NULL
+ })
+
+ if (!is.null(result)) return(result)
+ if (attempt < max_retries) Sys.sleep(0.5)
+ }
+
+ NULL
+}
+
+#' Download individual PNG images from the Dashboard
+#'
+#' Downloads specific PNG files from the Dashboard's \code{/data/} endpoint,
+#' one at a time. This is much faster than downloading entire zip archives
+#' when only a subset of ROIs are needed (e.g., class review mode).
+#'
+#' Samples that fail repeatedly are automatically skipped to avoid long
+#' waits when annotations reference samples not available on the dashboard.
+#'
+#' @param base_url Character. Dashboard base URL.
+#' @param file_names Character vector. PNG file names
+#' (e.g., \code{"D20240716T000431_IFCB134_00108.png"}).
+#' @param dest_dir Character. Destination directory.
+#' @param max_retries Integer. Maximum number of retry attempts per image.
+#' @param sample_fail_threshold Integer. After this many consecutive failures
+#' from the same sample, skip all remaining images from that sample.
+#' @return Character vector of successfully downloaded file names.
+#' @export
+download_dashboard_images_individual <- function(base_url, file_names, dest_dir,
+ max_retries = 3,
+ sample_fail_threshold = 2) {
+ dir.create(dest_dir, recursive = TRUE, showWarnings = FALSE)
+
+ succeeded <- character()
+ # Track consecutive failures per sample to skip unavailable samples early
+ sample_failures <- list()
+ skipped_samples <- character()
+
+ for (fname in file_names) {
+ # Parse sample_name and roi_number from file_name
+ parts <- regmatches(fname, regexec("^(.+)_(\\d+)\\.png$", fname))[[1]]
+ if (length(parts) < 3) next
+
+ sample_name <- parts[2]
+ roi_number <- as.integer(parts[3])
+
+ # Skip samples that have already been marked as unavailable
+ if (sample_name %in% skipped_samples) next
+
+ result <- download_dashboard_image_single(
+ base_url = base_url,
+ sample_name = sample_name,
+ roi_number = roi_number,
+ dest_dir = dest_dir,
+ max_retries = max_retries
+ )
+
+ if (!is.null(result)) {
+ succeeded <- c(succeeded, fname)
+ # Reset failure counter on success
+ sample_failures[[sample_name]] <- 0L
+ } else {
+ prev <- sample_failures[[sample_name]]
+ count <- (if (is.null(prev)) 0L else prev) + 1L
+ sample_failures[[sample_name]] <- count
+ if (count >= sample_fail_threshold) {
+ skipped_samples <- c(skipped_samples, sample_name)
+ warning("Skipping remaining images from ", sample_name,
+ " (", count, " consecutive failures)", call. = FALSE)
+ }
+ }
+ }
+
+ succeeded
+}
diff --git a/R/database.R b/R/database.R
index 536a218..31fd40a 100644
--- a/R/database.R
+++ b/R/database.R
@@ -601,6 +601,79 @@ export_db_to_png <- function(db_path, sample_name, roi_path, png_folder,
})
}
+#' Import annotations from a PNG class folder into the SQLite database
+#'
+#' Scans a folder of PNG images organized in class-name subfolders (via
+#' \code{\link{scan_png_class_folder}}) and imports the annotations into the
+#' database. An optional \code{class_mapping} named vector remaps class names
+#' before saving.
+#'
+#' @param png_folder Path to the top-level folder containing class subfolders
+#' @param db_path Path to the SQLite database file
+#' @param class2use Character vector of class names (preserves index order for
+#' .mat export)
+#' @param class_mapping Optional named character vector mapping scanned class
+#' names to target class names. Names are the source classes, values are the
+#' target classes. Classes not in the mapping are kept as-is.
+#' @param annotator Annotator name (defaults to \code{"imported"})
+#' @return Named list with counts: \code{success}, \code{failed}
+#' @export
+#' @examples
+#' \dontrun{
+#' db_path <- get_db_path("/data/manual")
+#' class2use <- c("Diatom", "Dinoflagellate", "Ciliate")
+#' result <- import_png_folder_to_db(
+#' "/data/png_export", db_path, class2use,
+#' class_mapping = c("OldName" = "NewName"),
+#' annotator = "Jane"
+#' )
+#' cat(result$success, "imported,", result$failed, "failed\n")
+#' }
+import_png_folder_to_db <- function(png_folder, db_path, class2use,
+ class_mapping = NULL,
+ annotator = "imported") {
+ scan_result <- scan_png_class_folder(png_folder)
+
+ counts <- list(success = 0L, failed = 0L)
+
+ if (nrow(scan_result$annotations) == 0) {
+ return(counts)
+ }
+
+ annotations <- scan_result$annotations
+
+ # Apply class mapping if provided
+ if (!is.null(class_mapping) && length(class_mapping) > 0) {
+ mapped <- class_mapping[annotations$class_name]
+ has_mapping <- !is.na(mapped)
+ annotations$class_name[has_mapping] <- mapped[has_mapping]
+ }
+
+ # Group by sample_name and save each sample
+
+ sample_names <- unique(annotations$sample_name)
+
+ for (sn in sample_names) {
+ sample_rows <- annotations[annotations$sample_name == sn, ]
+
+ classifications <- data.frame(
+ file_name = sample_rows$file_name,
+ class_name = sample_rows$class_name,
+ stringsAsFactors = FALSE
+ )
+
+ ok <- save_annotations_db(db_path, sn, classifications, class2use,
+ annotator)
+ if (isTRUE(ok)) {
+ counts$success <- counts$success + 1L
+ } else {
+ counts$failed <- counts$failed + 1L
+ }
+ }
+
+ counts
+}
+
#' Bulk export all annotated samples from SQLite to class-organized PNGs
#'
#' Exports every annotated sample in the database to PNG images organized
@@ -650,3 +723,273 @@ export_all_db_to_png <- function(db_path, png_folder, roi_path_map,
counts
}
+
+#' List all classes with counts in the annotations database
+#'
+#' Queries the database for distinct class names and their annotation counts.
+#' Useful for populating class review mode dropdowns. Optional filters restrict
+#' results to annotations matching a given year, month, or instrument.
+#'
+#' @param db_path Path to the SQLite database file
+#' @param year Optional year filter (e.g. \code{"2023"}). When not \code{"all"}
+#' or \code{NULL}, restricts to sample names starting with \code{DYYYY}.
+#' @param month Optional month filter (e.g. \code{"03"}). When not \code{"all"}
+#' or \code{NULL}, restricts to sample names with that month at positions 6-7.
+#' @param instrument Optional instrument filter (e.g. \code{"IFCB134"}). When
+#' not \code{"all"} or \code{NULL}, restricts to sample names ending with
+#' \code{_INSTRUMENT}.
+#' @param annotator Optional annotator name filter (e.g. \code{"Jane"}). When
+#' not \code{"all"} or \code{NULL}, restricts to annotations by that annotator.
+#' @return Data frame with columns \code{class_name} and \code{count}, ordered
+#' alphabetically by class name. Returns an empty data frame if the database
+#' does not exist or has no annotations.
+#' @export
+#' @examples
+#' \dontrun{
+#' db_path <- get_db_path("/data/manual")
+#' classes <- list_classes_db(db_path)
+#' classes_2023 <- list_classes_db(db_path, year = "2023")
+#' }
+list_classes_db <- function(db_path, year = NULL, month = NULL,
+ instrument = NULL, annotator = NULL) {
+ empty <- data.frame(class_name = character(), count = integer(),
+ stringsAsFactors = FALSE)
+
+ if (!file.exists(db_path)) {
+ return(empty)
+ }
+
+ con <- dbConnect(SQLite(), db_path)
+ on.exit(dbDisconnect(con), add = TRUE)
+
+ tables <- dbGetQuery(con, "SELECT name FROM sqlite_master WHERE type='table'")
+ if (!"annotations" %in% tables$name) {
+ return(empty)
+ }
+
+ where <- build_sample_filter_clause(year, month, instrument,
+ annotator = annotator)
+
+ sql <- paste0(
+ "SELECT class_name, COUNT(*) AS count FROM annotations",
+ where$clause,
+ " GROUP BY class_name ORDER BY class_name"
+ )
+
+ if (length(where$params) > 0) {
+ dbGetQuery(con, sql, params = where$params)
+ } else {
+ dbGetQuery(con, sql)
+ }
+}
+
+#' Load all annotations for a specific class from the database
+#'
+#' Returns every annotation matching \code{class_name}, with a computed
+#' \code{file_name} column for gallery display. Optional filters restrict
+#' results by year, month, or instrument.
+#'
+#' @param db_path Path to the SQLite database file
+#' @param class_name Class name to load
+#' @param year Optional year filter (e.g. \code{"2023"})
+#' @param month Optional month filter (e.g. \code{"03"})
+#' @param instrument Optional instrument filter (e.g. \code{"IFCB134"})
+#' @param annotator Optional annotator name filter (e.g. \code{"Jane"})
+#' @return Data frame with columns \code{sample_name}, \code{roi_number},
+#' \code{class_name}, and \code{file_name}. Returns \code{NULL} if no
+#' annotations match.
+#' @export
+#' @examples
+#' \dontrun{
+#' db_path <- get_db_path("/data/manual")
+#' diatoms <- load_class_annotations_db(db_path, "Diatom")
+#' diatoms_2023 <- load_class_annotations_db(db_path, "Diatom", year = "2023")
+#' }
+load_class_annotations_db <- function(db_path, class_name, year = NULL,
+ month = NULL, instrument = NULL,
+ annotator = NULL) {
+ if (!file.exists(db_path)) {
+ return(NULL)
+ }
+
+ con <- dbConnect(SQLite(), db_path)
+ on.exit(dbDisconnect(con), add = TRUE)
+
+ where <- build_sample_filter_clause(year, month, instrument,
+ annotator = annotator)
+ params <- c(list(class_name), where$params)
+
+ rows <- dbGetQuery(con, paste0(
+ "SELECT sample_name, roi_number, class_name FROM annotations WHERE class_name = ?",
+ if (nzchar(where$clause)) gsub("^ WHERE ", " AND ", where$clause),
+ " ORDER BY sample_name, roi_number"
+ ), params = params)
+
+ if (nrow(rows) == 0) {
+ return(NULL)
+ }
+
+ rows$file_name <- sprintf("%s_%05d.png", rows$sample_name, rows$roi_number)
+ rows
+}
+
+#' Save class review changes to the database
+#'
+#' Performs row-level UPDATEs for reclassified images identified during class
+#' review mode. Only the changed rows are updated; other annotations for the
+#' same samples are left untouched.
+#'
+#' @param db_path Path to the SQLite database file
+#' @param changes_df Data frame with columns \code{sample_name},
+#' \code{roi_number}, and \code{new_class_name}
+#' @param annotator Annotator name
+#' @return Integer count of rows updated
+#' @export
+#' @examples
+#' \dontrun{
+#' db_path <- get_db_path("/data/manual")
+#' changes <- data.frame(
+#' sample_name = "D20230101T120000_IFCB134",
+#' roi_number = 5L,
+#' new_class_name = "Ciliate"
+#' )
+#' save_class_review_changes_db(db_path, changes, "Jane")
+#' }
+save_class_review_changes_db <- function(db_path, changes_df, annotator) {
+ if (is.null(changes_df) || nrow(changes_df) == 0) {
+ return(0L)
+ }
+
+ con <- dbConnect(SQLite(), db_path)
+ on.exit(dbDisconnect(con), add = TRUE)
+
+ init_db_schema(con)
+
+ timestamp <- format(Sys.time(), "%Y-%m-%d %H:%M:%S")
+ updated <- 0L
+
+ tryCatch({
+ dbExecute(con, "BEGIN TRANSACTION")
+
+ for (i in seq_len(nrow(changes_df))) {
+ n <- dbExecute(con,
+ "UPDATE annotations SET class_name = ?, annotator = ?, timestamp = ?, is_manual = 1 WHERE sample_name = ? AND roi_number = ?",
+ params = list(
+ changes_df$new_class_name[i],
+ annotator,
+ timestamp,
+ changes_df$sample_name[i],
+ changes_df$roi_number[i]
+ )
+ )
+ updated <- updated + as.integer(n)
+ }
+
+ dbExecute(con, "COMMIT")
+ updated
+ }, error = function(e) {
+ tryCatch(dbExecute(con, "ROLLBACK"), error = function(e2) NULL)
+ warning("Failed to save class review changes: ", e$message)
+ 0L
+ })
+}
+
+#' List distinct years, months, and instruments from annotations
+#'
+#' Extracts metadata from sample names in the annotations table for use as
+#' filter options. Sample names follow the IFCB naming convention
+#' \code{DYYYYMMDDTHHMMSS_INSTRUMENT}.
+#'
+#' @param db_path Path to the SQLite database file
+#' @return A list with character vectors: \code{years}, \code{months},
+#' \code{instruments}, and \code{annotators}. Returns empty vectors if the
+#' database does not exist or has no annotations.
+#' @export
+#' @examples
+#' \dontrun{
+#' db_path <- get_db_path("/data/manual")
+#' meta <- list_annotation_metadata_db(db_path)
+#' meta$years # e.g. c("2022", "2023")
+#' meta$months # e.g. c("01", "06", "12")
+#' meta$instruments # e.g. c("IFCB134", "IFCB135")
+#' meta$annotators # e.g. c("Jane", "imported")
+#' }
+list_annotation_metadata_db <- function(db_path) {
+ empty <- list(years = character(), months = character(),
+ instruments = character(), annotators = character())
+
+ if (!file.exists(db_path)) {
+ return(empty)
+ }
+
+ con <- dbConnect(SQLite(), db_path)
+ on.exit(dbDisconnect(con), add = TRUE)
+
+ tables <- dbGetQuery(con, "SELECT name FROM sqlite_master WHERE type='table'")
+ if (!"annotations" %in% tables$name) {
+ return(empty)
+ }
+
+ samples <- dbGetQuery(con,
+ "SELECT DISTINCT sample_name FROM annotations"
+ )$sample_name
+
+ annotators <- sort(dbGetQuery(con,
+ "SELECT DISTINCT annotator FROM annotations WHERE annotator IS NOT NULL"
+ )$annotator)
+
+ if (length(samples) == 0) {
+ return(list(years = character(), months = character(),
+ instruments = character(), annotators = annotators))
+ }
+
+ years <- sort(unique(substr(samples, 2, 5)))
+ months <- sort(unique(substr(samples, 6, 7)))
+ instruments <- sort(unique(sub(".*_", "", samples)))
+
+ list(years = years, months = months, instruments = instruments,
+ annotators = annotators)
+}
+
+# Build WHERE clause fragments for sample_name filtering
+#
+# @param year Year string or "all"/NULL
+# @param month Month string or "all"/NULL
+# @param instrument Instrument string or "all"/NULL
+# @return List with `clause` (SQL fragment starting with " WHERE " or "") and
+# `params` (list of bind values)
+# @keywords internal
+build_sample_filter_clause <- function(year = NULL, month = NULL,
+ instrument = NULL,
+ annotator = NULL) {
+ conditions <- character()
+ params <- list()
+
+ if (!is.null(year) && year != "all") {
+ conditions <- c(conditions, "sample_name LIKE ?")
+ params <- c(params, list(paste0("D", year, "%")))
+ }
+
+ if (!is.null(month) && month != "all") {
+ conditions <- c(conditions, "sample_name LIKE ?")
+ params <- c(params, list(paste0("D____", month, "%")))
+ }
+
+ if (!is.null(instrument) && instrument != "all") {
+ conditions <- c(conditions, "sample_name LIKE ?")
+ params <- c(params, list(paste0("%_", instrument)))
+ }
+
+ if (!is.null(annotator) && annotator != "all") {
+ conditions <- c(conditions, "annotator = ?")
+ params <- c(params, list(annotator))
+ }
+
+ clause <- if (length(conditions) > 0) {
+ paste0(" WHERE ", paste(conditions, collapse = " AND "))
+ } else {
+ ""
+ }
+
+ list(clause = clause, params = params)
+}
diff --git a/R/sample_loading.R b/R/sample_loading.R
index 0f97e9f..6066db9 100644
--- a/R/sample_loading.R
+++ b/R/sample_loading.R
@@ -15,9 +15,12 @@ NULL
#' \item{class_name}{Predicted class name (e.g., `Diatom`).}
#' }
#'
-#' An optional column may also be included:
+#' Optional columns may also be included:
#' \describe{
#' \item{score}{Classification confidence value between 0 and 1.}
+#' \item{class_name_auto}{Raw (unthresholded) class prediction. When
+#' \code{use_threshold = FALSE} and this column exists, its values are
+#' used as \code{class_name}.}
#' }
#'
#' The CSV file must be named after the sample it describes
@@ -25,6 +28,9 @@ NULL
#' Folder configured in the app (subfolders are searched recursively).
#'
#' @param csv_path Path to classification CSV file
+#' @param use_threshold Logical, whether to use the threshold-filtered
+#' \code{class_name} column (default \code{TRUE}) or the raw
+#' \code{class_name_auto} column when available.
#' @return Data frame with classifications. Expected columns: `file_name`,
#' `class_name`, and optionally `score`.
#' @export
@@ -34,9 +40,15 @@ NULL
#' classifications <- load_from_csv("/path/to/D20230101T120000_IFCB134.csv")
#' head(classifications)
#' }
-load_from_csv <- function(csv_path) {
+load_from_csv <- function(csv_path, use_threshold = TRUE) {
classifications <- utils::read.csv(csv_path, stringsAsFactors = FALSE)
+ # When threshold is off and class_name_auto exists, use raw predictions
+
+ if (!use_threshold && "class_name_auto" %in% names(classifications)) {
+ classifications$class_name <- classifications$class_name_auto
+ }
+
# Strip trailing 3-digit suffix from class names (e.g., "Diatom_001" -> "Diatom")
# This matches iRfcb behavior where class folders may include numeric suffixes
classifications$class_name <- sub("_\\d{3}$", "", classifications$class_name)
@@ -55,6 +67,80 @@ load_from_csv <- function(csv_path) {
classifications
}
+#' Load classifications from HDF5 classifier output file
+#'
+#' Reads an HDF5 classifier output file (from iRfcb 0.8.0+) and extracts
+#' class predictions. Requires the \pkg{hdf5r} package.
+#'
+#' @param h5_path Path to classifier H5 file (matching pattern *_class*.h5)
+#' @param sample_name Sample name (e.g., "D20220522T000439_IFCB134")
+#' @param roi_dimensions Data frame from \code{\link{read_roi_dimensions}}
+#' @param use_threshold Logical, whether to use the threshold-filtered
+#' \code{class_name} dataset (default \code{TRUE}) or the raw
+#' \code{class_name_auto} dataset.
+#' @return Data frame with columns: file_name, class_name, score, width, height,
+#' roi_area
+#' @export
+#' @examples
+#' \dontrun{
+#' dims <- read_roi_dimensions("/data/raw/2022/D20220522/D20220522T000439_IFCB134.adc")
+#' classifications <- load_from_h5(
+#' h5_path = "/data/classified/D20220522T000439_IFCB134_class.h5",
+#' sample_name = "D20220522T000439_IFCB134",
+#' roi_dimensions = dims,
+#' use_threshold = TRUE
+#' )
+#' head(classifications)
+#' }
+load_from_h5 <- function(h5_path, sample_name, roi_dimensions, use_threshold = TRUE) {
+ if (!requireNamespace("hdf5r", quietly = TRUE)) {
+ stop("Package 'hdf5r' is required to read H5 classification files. ",
+ "Install it with: install.packages('hdf5r')")
+ }
+
+ h5 <- hdf5r::H5File$new(h5_path, "r")
+ on.exit(h5$close_all(), add = TRUE)
+
+ roi_numbers <- h5[["roi_numbers"]]$read()
+
+ if (use_threshold) {
+ class_names <- h5[["class_name"]]$read()
+ } else {
+ class_names <- h5[["class_name_auto"]]$read()
+ }
+
+ class_names[is.na(class_names)] <- "unclassified"
+
+ # Extract per-ROI max score from output_scores matrix (num_classes x num_rois)
+ output_scores <- h5[["output_scores"]]$read()
+ scores <- apply(output_scores, 2, max)
+
+ # Match ROI dimensions
+ roi_data <- lapply(roi_numbers, function(rn) {
+ idx <- which(roi_dimensions$roi_number == rn)
+ if (length(idx) > 0) {
+ list(width = roi_dimensions$width[idx],
+ height = roi_dimensions$height[idx],
+ area = roi_dimensions$area[idx])
+ } else {
+ list(width = NA_real_, height = NA_real_, area = NA_real_)
+ }
+ })
+
+ classifications <- data.frame(
+ file_name = sprintf("%s_%05d.png", sample_name, roi_numbers),
+ class_name = class_names,
+ score = scores,
+ width = vapply(roi_data, `[[`, numeric(1), "width"),
+ height = vapply(roi_data, `[[`, numeric(1), "height"),
+ roi_area = vapply(roi_data, `[[`, numeric(1), "area"),
+ stringsAsFactors = FALSE
+ )
+
+ # Sort by area (descending)
+ classifications[order(-classifications$roi_area), ]
+}
+
#' Load classifications from SQLite database
#'
#' Reads annotations for a sample from the SQLite database and returns a data
@@ -257,6 +343,111 @@ create_new_classifications <- function(sample_name, roi_dimensions) {
classifications[order(-classifications$roi_area), ]
}
+#' Scan a PNG folder with class subfolders
+#'
+#' Scans a directory containing PNG images organized into class-name
+#' subfolders (e.g. as exported by \code{\link{export_db_to_png}} or other
+#' tools). Folder names follow the iRfcb convention where a trailing 3-digit
+#' suffix is stripped (e.g. \code{Diatom_001} becomes \code{Diatom}).
+#'
+#' @param png_folder Path to the top-level folder containing class subfolders
+#' @return A list with components:
+#' \describe{
+#' \item{annotations}{Data frame with columns \code{sample_name},
+#' \code{roi_number}, \code{file_name}, and \code{class_name}}
+#' \item{classes_found}{Character vector of unique class names found}
+#' \item{sample_names}{Character vector of unique sample names found}
+#' }
+#' @export
+#' @examples
+#' \dontrun{
+#' result <- scan_png_class_folder("/data/png_export")
+#' head(result$annotations)
+#' result$classes_found
+#' result$sample_names
+#' }
+scan_png_class_folder <- function(png_folder) {
+ if (!dir.exists(png_folder)) {
+ stop("PNG folder does not exist: ", png_folder)
+ }
+
+ subdirs <- list.dirs(png_folder, recursive = FALSE, full.names = TRUE)
+
+ if (length(subdirs) == 0) {
+ return(list(
+ annotations = data.frame(
+ sample_name = character(),
+ roi_number = integer(),
+ file_name = character(),
+ class_name = character(),
+ stringsAsFactors = FALSE
+ ),
+ classes_found = character(),
+ sample_names = character()
+ ))
+ }
+
+ all_rows <- list()
+ seen_rois <- list()
+
+ for (subdir in subdirs) {
+ class_name <- sub("_\\d{3}$", "", basename(subdir))
+ png_files <- list.files(subdir, pattern = "\\.png$", full.names = FALSE)
+
+ for (fn in png_files) {
+ # Parse sample_name and roi_number from filename
+ # Expected format: SampleName_NNNNN.png (5-digit ROI number)
+ m <- regmatches(fn, regexec("^(.+)_(\\d{5})\\.png$", fn))[[1]]
+ if (length(m) < 3) {
+ warning("Skipping file with unexpected name format: ", fn)
+ next
+ }
+
+ sample_name <- m[2]
+ roi_number <- as.integer(m[3])
+ roi_key <- paste0(sample_name, "_", roi_number)
+
+ if (!is.null(seen_rois[[roi_key]])) {
+ warning(sprintf("Duplicate ROI %s found in class '%s' (already in '%s'), using first occurrence",
+ roi_key, class_name, seen_rois[[roi_key]]))
+ next
+ }
+ seen_rois[[roi_key]] <- class_name
+
+ all_rows[[length(all_rows) + 1L]] <- data.frame(
+ sample_name = sample_name,
+ roi_number = roi_number,
+ file_name = fn,
+ class_name = class_name,
+ stringsAsFactors = FALSE
+ )
+ }
+ }
+
+ if (length(all_rows) == 0) {
+ return(list(
+ annotations = data.frame(
+ sample_name = character(),
+ roi_number = integer(),
+ file_name = character(),
+ class_name = character(),
+ stringsAsFactors = FALSE
+ ),
+ classes_found = character(),
+ sample_names = character()
+ ))
+ }
+
+ annotations <- do.call(rbind, all_rows)
+ rownames(annotations) <- NULL
+
+ list(
+ annotations = annotations,
+ classes_found = sort(unique(annotations$class_name)),
+ sample_names = sort(unique(annotations$sample_name))
+ )
+}
+
#' Filter classifications to only include extracted images
#'
#' Filters a classifications data frame to only include ROIs that have
diff --git a/R/utils.R b/R/utils.R
index d8c3717..4505f3a 100644
--- a/R/utils.R
+++ b/R/utils.R
@@ -136,12 +136,16 @@ load_file_index <- function() {
#' If folder paths are not provided, they are read from saved settings.
#'
#' @param roi_folder Path to ROI data folder. If NULL, read from saved settings.
-#' @param csv_folder Path to classification folder (CSV/MAT). If NULL, read from saved settings.
+#' @param csv_folder Path to classification folder (CSV/H5/MAT). If NULL, read from saved settings.
#' @param output_folder Path to output folder for MAT annotations. If NULL, read from saved settings.
#' @param verbose If TRUE, print progress messages. Default TRUE.
#' @param db_folder Path to the database folder for SQLite annotations. If NULL,
#' read from saved settings; if not found in settings, defaults to
#' \code{\link{get_default_db_dir}()}.
+#' @param data_source Either \code{"local"} (default) for local folder scanning,
+#' or \code{"dashboard"} to fetch the sample list from a remote IFCB Dashboard.
+#' @param dashboard_url When \code{data_source = "dashboard"}, the full Dashboard
+#' URL (e.g. \code{"https://habon-ifcb.whoi.edu/timeline?dataset=tangosund"}).
#' @return Invisibly returns the file index list, or NULL if roi_folder is invalid.
#' @export
#' @examples
@@ -156,12 +160,17 @@ load_file_index <- function() {
#' output_folder = "/data/ifcb/manual"
#' )
#'
+#' # Scan from a remote Dashboard
+#' rescan_file_index(data_source = "dashboard",
+#' dashboard_url = "https://habon-ifcb.whoi.edu/timeline?dataset=tangosund")
+#'
#' # Use in a cron job:
#' # Rscript -e 'ClassiPyR::rescan_file_index()'
#' }
rescan_file_index <- function(roi_folder = NULL, csv_folder = NULL,
output_folder = NULL, verbose = TRUE,
- db_folder = NULL) {
+ db_folder = NULL, data_source = "local",
+ dashboard_url = NULL) {
# Read from saved settings if not provided
if (is.null(roi_folder) || is.null(csv_folder) || is.null(output_folder) ||
is.null(db_folder)) {
@@ -183,6 +192,58 @@ rescan_file_index <- function(roi_folder = NULL, csv_folder = NULL,
db_folder <- get_default_db_dir()
}
+ # Dashboard mode: fetch sample list from remote API
+ if (identical(data_source, "dashboard")) {
+ if (is.null(dashboard_url) || !nzchar(dashboard_url)) {
+ if (verbose) message("Dashboard URL not set")
+ return(invisible(NULL))
+ }
+
+ parsed <- parse_dashboard_url(dashboard_url)
+ if (verbose) message("Fetching bin list from: ", parsed$base_url)
+
+ bins <- tryCatch(
+ list_dashboard_bins(parsed$base_url, parsed$dataset_name),
+ error = function(e) {
+ if (verbose) message("Failed to list dashboard bins: ", e$message)
+ character()
+ }
+ )
+
+ sample_names <- as.character(bins)
+ if (verbose) message(" Found ", length(sample_names), " samples")
+
+ if (length(sample_names) == 0) {
+ if (verbose) message("No samples found on dashboard.")
+ return(invisible(NULL))
+ }
+
+ # Check DB for existing annotations
+ db_path <- get_db_path(db_folder)
+ annotated_db <- list_annotated_samples_db(db_path)
+ annotated <- annotated_db[annotated_db %in% sample_names]
+
+ index_data <- list(
+ data_source = "dashboard",
+ dashboard_url = dashboard_url,
+ dashboard_base_url = parsed$base_url,
+ dashboard_dataset = parsed$dataset_name,
+ sample_names = sample_names,
+ classified_samples = character(),
+ annotated_samples = annotated,
+ roi_path_map = list(),
+ csv_path_map = list(),
+ classifier_mat_files = list(),
+ classifier_h5_files = list(),
+ timestamp = as.character(Sys.time())
+ )
+
+ save_file_index(index_data)
+ if (verbose) message("File index saved to: ", get_file_index_path())
+
+ return(invisible(index_data))
+ }
+
# Validate ROI folder
roi_valid <- !is.null(roi_folder) && length(roi_folder) == 1 &&
!isTRUE(is.na(roi_folder)) && nzchar(roi_folder) && dir.exists(roi_folder)
@@ -221,6 +282,7 @@ rescan_file_index <- function(roi_folder = NULL, csv_folder = NULL,
# Scan classification files
classified <- character()
mat_file_map <- list()
+ h5_file_map <- list()
csv_map <- list()
if (csv_valid) {
@@ -248,9 +310,21 @@ rescan_file_index <- function(roi_folder = NULL, csv_folder = NULL,
}
}
+ h5_files <- list.files(csv_folder, pattern = "_class.*\\.h5$",
+ recursive = TRUE, full.names = TRUE)
+
+ for (h5_file in h5_files) {
+ h5_basename <- basename(h5_file)
+ sample_from_h5 <- sub("_class.*\\.h5$", "", h5_basename)
+ if (sample_from_h5 %in% sample_names) {
+ h5_file_map[[sample_from_h5]] <- h5_file
+ }
+ }
+
mat_samples <- names(mat_file_map)
+ h5_samples <- names(h5_file_map)
csv_matched <- csv_sample_names[csv_sample_names %in% sample_names]
- classified <- unique(c(csv_matched, mat_samples))
+ classified <- unique(c(csv_matched, h5_samples, mat_samples))
if (verbose) message(" Found ", length(classified), " classified samples")
}
@@ -286,6 +360,7 @@ rescan_file_index <- function(roi_folder = NULL, csv_folder = NULL,
roi_path_map = roi_map,
csv_path_map = csv_map,
classifier_mat_files = mat_file_map,
+ classifier_h5_files = h5_file_map,
timestamp = as.character(Sys.time())
)
diff --git a/README.md b/README.md
index 65a6a1d..d07276d 100644
--- a/README.md
+++ b/README.md
@@ -16,6 +16,9 @@ A Shiny application for manual (human) image classification and validation of Im
## Features
- **Dual Mode**: Validate existing classifications or annotate from scratch
+- **Class Review**: Review and reclassify all images of a specific class across the entire database
+- **IFCB Dashboard**: Work directly with remote IFCB Dashboard instances - no local data files needed
+- **Live Prediction**: One-click CNN classification via a remote Gradio API using [iRfcb](https://github.com/EuropeanIFCBGroup/iRfcb)
- **Multiple Formats**: Load from CSV or MATLAB classifier output
- **SQLite Storage**: Annotations stored in a local SQLite database by default - no Python needed
- **Efficient Workflow**: Drag-select, batch relabeling, class filtering
diff --git a/_pkgdown.yml b/_pkgdown.yml
index dda7f38..fe49626 100644
--- a/_pkgdown.yml
+++ b/_pkgdown.yml
@@ -48,10 +48,12 @@ reference:
- load_class_list
- load_from_classifier_mat
- load_from_csv
+ - load_from_h5
- load_from_mat
- load_from_db
- create_new_classifications
- filter_to_extracted
+ - scan_png_class_folder
- title: Sample Saving
desc: Functions for saving annotations and exporting images
contents:
@@ -66,6 +68,7 @@ reference:
- save_annotations_db
- load_annotations_db
- list_annotated_samples_db
+ - list_annotation_metadata_db
- update_annotator
- import_mat_to_db
- import_all_mat_to_db
@@ -73,6 +76,10 @@ reference:
- export_all_db_to_mat
- export_db_to_png
- export_all_db_to_png
+ - import_png_folder_to_db
+ - list_classes_db
+ - load_class_annotations_db
+ - save_class_review_changes_db
- title: File Index Cache
desc: Functions for managing the file index cache for faster startup
contents:
@@ -80,6 +87,18 @@ reference:
- load_file_index
- save_file_index
- rescan_file_index
+- title: Dashboard
+ desc: Functions for working with remote IFCB Dashboard instances
+ contents:
+ - parse_dashboard_url
+ - list_dashboard_bins
+ - download_dashboard_images
+ - download_dashboard_images_bulk
+ - download_dashboard_image_single
+ - download_dashboard_images_individual
+ - download_dashboard_adc
+ - download_dashboard_autoclass
+ - get_dashboard_cache_dir
- title: Utilities
desc: Helper functions for IFCB data processing
contents:
diff --git a/codecov.yml b/codecov.yml
index 02eff64..512fa26 100644
--- a/codecov.yml
+++ b/codecov.yml
@@ -13,7 +13,4 @@ ignore:
- "tests/"
- "docs/"
-comment:
- layout: "reach,diff,flags,files"
- behavior: default
- require_changes: true
+comment: false
diff --git a/inst/CITATION b/inst/CITATION
index 8b93f47..a277ae8 100644
--- a/inst/CITATION
+++ b/inst/CITATION
@@ -9,12 +9,12 @@ bibentry(
comment = c(ORCID = "0000-0002-8283-656X"))
),
year = "2026",
- note = "R package version 0.1.1",
+ note = "R package version 0.2.0",
url = "https://doi.org/10.5281/zenodo.18414999",
textVersion = paste(
"Torstensson, A. (2026).",
"ClassiPyR: A Shiny Application for Manual Image Classification and Validation of IFCB Data.",
- "R package version 0.1.1.",
+ "R package version 0.2.0.",
"https://doi.org/10.5281/zenodo.18414999"
)
)
diff --git a/inst/app/server.R b/inst/app/server.R
index de99b4c..17285fe 100644
--- a/inst/app/server.R
+++ b/inst/app/server.R
@@ -60,7 +60,13 @@ server <- function(input, output, session) {
resource_path_name = NULL, # Session-specific Shiny resource path for images
is_loading = FALSE, # TRUE while loading/saving operations in progress
measure_mode = FALSE, # TRUE when measure tool is active
- pending_sample_select = NULL # Pending sample selection for dropdown update
+ pending_sample_select = NULL, # Pending sample selection for dropdown update
+
+ # Class review mode state
+ class_review_mode = FALSE, # TRUE when in class review mode
+ class_review_class = NULL, # Currently reviewed class name
+ class_review_samples = character(), # Unique sample names in class review
+ class_review_original = NULL # Original classifications snapshot for diff
)
# Settings file for persistence (uses R_user_dir for CRAN compliance)
@@ -104,7 +110,17 @@ server <- function(input, output, session) {
class2use_path = NULL, # Path to class2use file for auto-loading
python_venv_path = NULL, # NULL = use ./venv in working directory
save_format = "sqlite", # "sqlite" (default), "mat", or "both"
- export_statistics = TRUE # Write validation statistics CSV files
+ export_statistics = TRUE, # Write validation statistics CSV files
+ skip_class_png = "", # Class name to exclude from PNG export
+ data_source = "local", # "local" or "dashboard"
+ dashboard_url = "", # IFCB Dashboard URL
+ dashboard_autoclass = FALSE, # Use dashboard auto-classifications for validation
+ gradio_url = "", # Gradio API URL for CNN classification
+ prediction_model = "", # Model name for live prediction
+ dashboard_parallel_downloads = 5,
+ dashboard_sleep_time = 2,
+ dashboard_multi_timeout = 120,
+ dashboard_max_retries = 3
)
if (file.exists(settings_file)) {
@@ -160,7 +176,17 @@ server <- function(input, output, session) {
auto_sync = saved_settings$auto_sync,
python_venv_path = saved_settings$python_venv_path,
save_format = saved_settings$save_format,
- export_statistics = saved_settings$export_statistics
+ export_statistics = saved_settings$export_statistics,
+ skip_class_png = saved_settings$skip_class_png,
+ data_source = saved_settings$data_source,
+ dashboard_url = saved_settings$dashboard_url,
+ dashboard_autoclass = saved_settings$dashboard_autoclass,
+ gradio_url = saved_settings$gradio_url,
+ prediction_model = saved_settings$prediction_model,
+ dashboard_parallel_downloads = saved_settings$dashboard_parallel_downloads,
+ dashboard_sleep_time = saved_settings$dashboard_sleep_time,
+ dashboard_multi_timeout = saved_settings$dashboard_multi_timeout,
+ dashboard_max_retries = saved_settings$dashboard_max_retries
)
# Initialize class dropdown with default class list on startup
@@ -173,10 +199,11 @@ server <- function(input, output, session) {
# Store all sample names and their classification status
all_samples <- reactiveVal(character())
- classified_samples <- reactiveVal(character()) # Auto-classified (CSV or classifier MAT)
+ classified_samples <- reactiveVal(character()) # Auto-classified (CSV, H5, or classifier MAT)
annotated_samples <- reactiveVal(character()) # Manually annotated (has .mat in output folder)
- # Store mapping of sample names to classifier MAT file paths
+ # Store mapping of sample names to classifier MAT/H5 file paths
classifier_mat_files <- reactiveVal(list())
+ classifier_h5_files <- reactiveVal(list())
# Path maps: sample_name -> full file path (discovered during scan)
roi_path_map <- reactiveVal(list())
csv_path_map <- reactiveVal(list())
@@ -221,25 +248,93 @@ server <- function(input, output, session) {
size = "l",
easyClose = TRUE,
- # ── Folder Paths ──────────────────────────────────────────────
- h5("Folder Paths"),
+ # ── Data Source ─────────────────────────────────────────────
+ h5("Data Source"),
+
+ radioButtons("cfg_data_source", NULL,
+ choices = c("Local Folders" = "local",
+ "IFCB Dashboard" = "dashboard"),
+ selected = config$data_source, inline = TRUE),
+
+ # Dashboard settings (visible when data source is "dashboard")
+ conditionalPanel(
+ condition = "input.cfg_data_source == 'dashboard'",
+ textInput("cfg_dashboard_url", "Dashboard URL",
+ value = config$dashboard_url, width = "100%",
+ placeholder = "https://habon-ifcb.whoi.edu/timeline?dataset=tangosund"),
+ tags$small(class = "text-muted", style = "display: block; margin-bottom: 10px;",
+ "Enter the IFCB Dashboard URL. Dataset can be specified via ?dataset=name."),
+ checkboxInput("cfg_dashboard_autoclass", "Use dashboard auto-classifications",
+ value = config$dashboard_autoclass),
+ tags$small(class = "text-muted", style = "display: block; margin-bottom: 15px;",
+ "When enabled, downloads auto-classification scores from the dashboard for validation mode."),
+
+ # Advanced download settings (collapsible)
+ tags$details(
+ tags$summary(style = "cursor: pointer; margin-bottom: 10px; color: #666;",
+ "Advanced Download Settings"),
+ fluidRow(
+ column(6, numericInput("cfg_dashboard_parallel_downloads",
+ "Parallel Downloads",
+ value = config$dashboard_parallel_downloads,
+ min = 1, max = 20, step = 1)),
+ column(6, numericInput("cfg_dashboard_sleep_time",
+ "Sleep Time (seconds)",
+ value = config$dashboard_sleep_time,
+ min = 0, max = 30, step = 0.5))
+ ),
+ fluidRow(
+ column(6, numericInput("cfg_dashboard_multi_timeout",
+ "Download Timeout (seconds)",
+ value = config$dashboard_multi_timeout,
+ min = 10, max = 600, step = 10)),
+ column(6, numericInput("cfg_dashboard_max_retries",
+ "Max Retries",
+ value = config$dashboard_max_retries,
+ min = 1, max = 10, step = 1))
+ ),
+ tags$small(class = "text-muted", style = "display: block; margin-bottom: 15px;",
+ "Settings for zip/ADC/autoclass downloads from the dashboard.")
+ )
+ ),
+
+ # ── Classification Folder (both modes) ──────────────────────────
+ h5("Classification"),
div(
- style = "display: flex; gap: 5px; align-items: flex-end; margin-bottom: 15px;",
+ style = "display: flex; gap: 5px; align-items: flex-end; margin-bottom: 5px;",
div(style = "flex: 1;",
- textInput("cfg_csv_folder", "Classification Folder (CSV/MAT)",
+ textInput("cfg_csv_folder", "Classification Folder (CSV/H5/MAT)",
value = config$csv_folder, width = "100%")),
shinyDirButton("browse_csv_folder", "Browse", "Select Classification Folder",
class = "btn-outline-secondary", style = "margin-bottom: 15px;")
),
- div(
- style = "display: flex; gap: 5px; align-items: flex-end; margin-bottom: 15px;",
- div(style = "flex: 1;",
- textInput("cfg_roi_folder", "ROI Data Folder",
- value = config$roi_folder, width = "100%")),
- shinyDirButton("browse_roi_folder", "Browse", "Select ROI Data Folder",
- class = "btn-outline-secondary", style = "margin-bottom: 15px;")
+ conditionalPanel(
+ condition = "input.cfg_data_source == 'dashboard'",
+ tags$small(class = "text-muted", style = "display: block; margin-bottom: 5px;",
+ "Optional. Use local classification files instead of dashboard auto-classifications.")
+ ),
+
+ checkboxInput("cfg_use_threshold", "Apply classification threshold",
+ value = config$use_threshold),
+ tags$small(class = "text-muted", style = "display: block; margin-bottom: 15px;",
+ "When enabled, classifications below the confidence threshold are marked as 'unclassified'."),
+
+ # ── Folder Paths (local mode only) ──────────────────────────────
+ conditionalPanel(
+ condition = "input.cfg_data_source == 'local'",
+
+ h5("ROI Data"),
+
+ div(
+ style = "display: flex; gap: 5px; align-items: flex-end; margin-bottom: 15px;",
+ div(style = "flex: 1;",
+ textInput("cfg_roi_folder", "ROI Data Folder",
+ value = config$roi_folder, width = "100%")),
+ shinyDirButton("browse_roi_folder", "Browse", "Select ROI Data Folder",
+ class = "btn-outline-secondary", style = "margin-bottom: 15px;")
+ )
),
div(
@@ -320,41 +415,60 @@ server <- function(input, output, session) {
# ── Import / Export ────────────────────────────────────────────
h5("Import / Export"),
+ tags$label("Import to SQLite", style = "font-weight: 600; display: block; margin-bottom: 5px;"),
div(
- style = "display: flex; gap: 10px; margin-bottom: 8px;",
- actionButton("import_mat_to_db_btn", "Import .mat \u2192 SQLite",
- icon = icon("database"), class = "btn-outline-secondary btn-sm"),
- actionButton("export_db_to_mat_btn", "Export SQLite \u2192 .mat",
- icon = icon("file-export"), class = "btn-outline-secondary btn-sm"),
- actionButton("export_db_to_png_btn", "Export SQLite \u2192 PNG",
- icon = icon("image"), class = "btn-outline-secondary btn-sm")
+ style = "display: flex; gap: 10px; margin-bottom: 5px;",
+ actionButton("import_mat_to_db_btn", ".mat \u2192 SQLite",
+ icon = icon("file-import"), class = "btn-outline-secondary btn-sm"),
+ actionButton("import_png_to_db_btn", "PNG \u2192 SQLite",
+ icon = icon("file-import"), class = "btn-outline-secondary btn-sm")
),
- tags$small(class = "text-muted",
- "Bulk import/export all annotated samples between storage formats.",
- "PNG export extracts images into class-name subfolders."),
+ tags$small(class = "text-muted", style = "display: block; margin-bottom: 10px;",
+ "Bulk import annotated samples from .mat files or PNG class folders."),
+ tags$label("Export from SQLite", style = "font-weight: 600; display: block; margin-bottom: 5px;"),
div(
- style = "margin-top: 8px;",
+ style = "display: flex; gap: 10px; margin-bottom: 5px;",
+ actionButton("export_db_to_mat_btn", "SQLite \u2192 .mat",
+ icon = icon("file-export"), class = "btn-outline-secondary btn-sm"),
+ actionButton("export_db_to_png_btn", "SQLite \u2192 PNG",
+ icon = icon("file-export"), class = "btn-outline-secondary btn-sm")
+ ),
+ div(
+ style = "margin-bottom: 5px;",
textInput("cfg_skip_class_png", "Skip class in PNG export",
- value = if (!is.null(rv$class2use) && length(rv$class2use) > 0) rv$class2use[1] else "",
+ value = if (nzchar(config$skip_class_png)) config$skip_class_png
+ else if (!is.null(rv$class2use) && length(rv$class2use) > 0) rv$class2use[1]
+ else "",
width = "250px"),
tags$small(class = "text-muted",
"Images with this class are excluded from PNG export.",
- "Pre-filled with the first class in your class list.",
"Leave empty to export all classes.")
),
hr(),
+ # ── Live Prediction ─────────────────────────────────────────
+ h5("Live Prediction"),
+
+ textInput("cfg_gradio_url", "Gradio API URL",
+ value = config$gradio_url, width = "100%",
+ placeholder = "https://irfcb-classify.hf.space"),
+ tags$small(class = "text-muted", style = "display: block; margin-bottom: 10px;",
+ "Enter Gradio API URL for CNN classification. Example: https://irfcb-classify.hf.space"),
+
+ selectInput("cfg_prediction_model", "Prediction Model",
+ choices = if (nzchar(config$prediction_model)) config$prediction_model else NULL,
+ selected = if (nzchar(config$prediction_model)) config$prediction_model else NULL,
+ width = "100%"),
+ tags$small(class = "text-muted", style = "display: block; margin-bottom: 15px;",
+ "Select a CNN model for classification. Models are fetched from the Gradio API."),
+
+ hr(),
+
# ── IFCB Options ──────────────────────────────────────────────
h5("IFCB Options"),
- checkboxInput("cfg_use_threshold", "Apply classification threshold",
- value = config$use_threshold),
- tags$small(class = "text-muted",
- "Only applies to ifcb-analysis MATLAB classifier output (*_class*.mat).",
- "When enabled, classifications below the confidence threshold are marked as 'unclassified'."),
-
div(
style = "display: flex; gap: 10px; align-items: center; margin-top: 10px;",
numericInput("cfg_pixels_per_micron", "Pixels per micron",
@@ -429,9 +543,35 @@ server <- function(input, output, session) {
}
})
-
+ # Fetch available models when Gradio URL changes in settings (debounced)
+ gradio_url_debounced <- debounce(reactive(input$cfg_gradio_url), 1500)
+
+ observeEvent(gradio_url_debounced(), {
+ url <- gradio_url_debounced()
+ if (is.null(url) || !nzchar(url)) {
+ updateSelectInput(session, "cfg_prediction_model", choices = character(0))
+ return()
+ }
+ tryCatch({
+ models <- iRfcb::ifcb_classify_models(url)
+ if (length(models) > 0) {
+ # Preserve current selection if still valid
+ current <- config$prediction_model
+ selected <- if (nzchar(current) && current %in% models) current else models[1]
+ updateSelectInput(session, "cfg_prediction_model",
+ choices = models, selected = selected)
+ } else {
+ updateSelectInput(session, "cfg_prediction_model", choices = character(0))
+ showNotification("No models found at the provided URL.", type = "warning")
+ }
+ }, error = function(e) {
+ updateSelectInput(session, "cfg_prediction_model", choices = character(0))
+ showNotification(paste("Could not fetch models:", e$message), type = "error")
+ })
+ })
+
# Class count display
-
+
output$class_count_text <- renderText({
if (is.null(rv$class2use)) {
"No class list loaded"
@@ -446,7 +586,11 @@ server <- function(input, output, session) {
title = "Class List Editor",
size = "l",
easyClose = TRUE,
-
+
+ tags$head(tags$style(HTML(
+ ".modal-dialog.modal-lg { max-width: 1200px; }"
+ ))),
+
tags$div(
class = "alert alert-warning",
style = "font-size: 12px; padding: 8px;",
@@ -544,21 +688,33 @@ server <- function(input, output, session) {
"No classes defined yet. Add classes using the form below or edit the text area."
))
}
-
+
classes <- rv$class2use
indices <- seq_along(classes)
-
+
+ # Get image counts per class from the database
+ counts <- tryCatch({
+ db_path <- get_db_path(config$db_folder)
+ if (file.exists(db_path)) {
+ classes_df <- list_classes_db(db_path)
+ setNames(classes_df$count, classes_df$class_name)
+ } else {
+ NULL
+ }
+ }, error = function(e) NULL)
+
# Create data frame for sorting
df <- data.frame(idx = indices, cls = classes, stringsAsFactors = FALSE)
-
+
if (rv$class_sort_mode == "alpha") {
df <- df[order(df$cls), ]
}
-
+
class_lines <- mapply(function(idx, cls) {
- tags$div(sprintf("%3d: %s", idx, cls))
+ count <- if (!is.null(counts) && cls %in% names(counts)) counts[[cls]] else 0L
+ tags$div(sprintf("%3d: %s (%d)", idx, cls, count))
}, df$idx, df$cls, SIMPLIFY = FALSE)
-
+
tagList(class_lines)
})
@@ -684,8 +840,10 @@ server <- function(input, output, session) {
# Check if folder paths actually changed (to avoid spurious resets)
roi_changed <- !identical(config$roi_folder, input$cfg_roi_folder)
csv_changed <- !identical(config$csv_folder, input$cfg_csv_folder)
- paths_changed <- roi_changed || csv_changed
-
+ data_source_changed <- !identical(config$data_source, input$cfg_data_source)
+ dashboard_url_changed <- !identical(config$dashboard_url, input$cfg_dashboard_url)
+ paths_changed <- roi_changed || csv_changed || data_source_changed || dashboard_url_changed
+
config$csv_folder <- input$cfg_csv_folder
config$roi_folder <- input$cfg_roi_folder
config$output_folder <- input$cfg_output_folder
@@ -696,6 +854,16 @@ server <- function(input, output, session) {
config$auto_sync <- input$cfg_auto_sync
config$save_format <- input$cfg_save_format
config$export_statistics <- input$cfg_export_statistics
+ config$skip_class_png <- input$cfg_skip_class_png
+ config$data_source <- input$cfg_data_source
+ config$dashboard_url <- input$cfg_dashboard_url
+ config$dashboard_autoclass <- input$cfg_dashboard_autoclass
+ config$dashboard_parallel_downloads <- if (!is.null(input$cfg_dashboard_parallel_downloads)) input$cfg_dashboard_parallel_downloads else 5
+ config$dashboard_sleep_time <- if (!is.null(input$cfg_dashboard_sleep_time)) input$cfg_dashboard_sleep_time else 2
+ config$dashboard_multi_timeout <- if (!is.null(input$cfg_dashboard_multi_timeout)) input$cfg_dashboard_multi_timeout else 120
+ config$dashboard_max_retries <- if (!is.null(input$cfg_dashboard_max_retries)) input$cfg_dashboard_max_retries else 3
+ config$gradio_url <- input$cfg_gradio_url
+ config$prediction_model <- input$cfg_prediction_model
# Persist settings to file for next session
# python_venv_path is kept from config (set via run_app() or previous save)
@@ -710,13 +878,23 @@ server <- function(input, output, session) {
auto_sync = input$cfg_auto_sync,
save_format = input$cfg_save_format,
export_statistics = input$cfg_export_statistics,
+ skip_class_png = input$cfg_skip_class_png,
class2use_path = rv$class2use_path,
- python_venv_path = config$python_venv_path
+ python_venv_path = config$python_venv_path,
+ data_source = input$cfg_data_source,
+ dashboard_url = input$cfg_dashboard_url,
+ dashboard_autoclass = input$cfg_dashboard_autoclass,
+ dashboard_parallel_downloads = config$dashboard_parallel_downloads,
+ dashboard_sleep_time = config$dashboard_sleep_time,
+ dashboard_multi_timeout = config$dashboard_multi_timeout,
+ dashboard_max_retries = config$dashboard_max_retries,
+ gradio_url = input$cfg_gradio_url,
+ prediction_model = input$cfg_prediction_model
))
-
+
removeModal()
showNotification("Settings saved.", type = "message")
-
+
# Only trigger sample rescan if folder paths actually changed
if (paths_changed) {
cache_path <- get_file_index_path()
@@ -727,6 +905,154 @@ server <- function(input, output, session) {
}
})
+ # ============================================================================
+ # LIVE PREDICTION
+ # ============================================================================
+
+ # Render predict button conditionally
+ output$predict_btn_ui <- renderUI({
+ has_config <- nzchar(config$gradio_url) && nzchar(config$prediction_model)
+ has_sample <- !is.null(rv$current_sample)
+
+ if (!has_config) return(NULL)
+
+ actionButton("predict_btn", "Predict",
+ icon = icon("robot"),
+ class = if (has_sample) "btn-info" else "btn-outline-secondary",
+ width = "100%",
+ disabled = if (!has_sample) "disabled" else NULL)
+ })
+
+ # Predict observer
+ observeEvent(input$predict_btn, {
+ req(rv$current_sample, rv$temp_png_folder, config$gradio_url, config$prediction_model)
+
+ # Validate Gradio URL scheme
+ if (!grepl("^https?://", config$gradio_url)) {
+ showNotification("Gradio URL must start with http:// or https://", type = "error")
+ return()
+ }
+
+ # Get PNG files from temp folder (PNGs live in a sample-name subfolder)
+ png_folder <- file.path(rv$temp_png_folder, rv$current_sample)
+ if (!dir.exists(png_folder)) {
+ # Fallback: try the temp folder directly (e.g. dashboard cache)
+ png_folder <- rv$temp_png_folder
+ }
+ png_files <- list.files(png_folder, pattern = "\\.png$", full.names = TRUE)
+ if (length(png_files) == 0) {
+ showNotification("No PNG images found in the loaded sample.", type = "warning")
+ return()
+ }
+
+ # Identify images manually reclassified by the user (skip these)
+ manually_changed <- character()
+ if (!is.null(rv$original_classifications) && !is.null(rv$classifications)) {
+ merged <- merge(
+ rv$classifications[, c("file_name", "class_name")],
+ rv$original_classifications[, c("file_name", "class_name")],
+ by = "file_name", suffixes = c("_current", "_original")
+ )
+ changed_rows <- merged$class_name_current != merged$class_name_original
+ manually_changed <- merged$file_name[changed_rows]
+ }
+
+ # Filter to only non-reclassified images
+ all_filenames <- basename(png_files)
+ files_to_predict <- png_files[!all_filenames %in% manually_changed]
+
+ if (length(files_to_predict) == 0) {
+ showNotification("All images have been manually reclassified. Nothing to predict.",
+ type = "message")
+ return()
+ }
+
+ # Run classification with per-image progress
+ n_total <- length(files_to_predict)
+ result_list <- vector("list", n_total)
+ failed <- 0L
+
+ withProgress(message = "Classifying images...", value = 0, {
+ for (i in seq_len(n_total)) {
+ setProgress(value = (i - 1) / n_total,
+ detail = paste0(i, " / ", n_total))
+ tryCatch({
+ result_list[[i]] <- iRfcb::ifcb_classify_images(
+ png_file = files_to_predict[i],
+ gradio_url = config$gradio_url,
+ model_name = config$prediction_model,
+ verbose = FALSE
+ )
+ }, error = function(e) {
+ failed <<- failed + 1L
+ })
+ }
+ setProgress(1, detail = "Done")
+ })
+
+ predictions <- do.call(rbind, Filter(Negate(is.null), result_list))
+
+ if (is.null(predictions) || nrow(predictions) == 0) {
+ showNotification("Prediction failed for all images.", type = "error")
+ return()
+ }
+ if (failed > 0) {
+ showNotification(paste("Warning:", failed, "images failed to classify."),
+ type = "warning")
+ }
+
+ # Apply threshold logic
+ if (!config$use_threshold && "class_name_auto" %in% names(predictions)) {
+ predictions$class_name <- predictions$class_name_auto
+ }
+
+ # Merge predictions into rv$classifications
+ cls <- rv$classifications
+ for (i in seq_len(nrow(predictions))) {
+ fname <- predictions$file_name[i]
+ idx <- which(cls$file_name == fname)
+ if (length(idx) == 1) {
+ cls$class_name[idx] <- predictions$class_name[i]
+ if ("score" %in% names(cls)) {
+ cls$score[idx] <- predictions$score[i]
+ }
+ }
+ }
+ rv$classifications <- cls
+
+ # Update original_classifications so predictions become the new baseline
+ rv$original_classifications <- rv$classifications
+
+ # Switch to validation mode since we now have auto-classifications
+ rv$is_annotation_mode <- FALSE
+
+ # Add any new classes from predictions to class2use
+ new_classes <- setdiff(unique(predictions$class_name), rv$class2use)
+ new_classes <- new_classes[!is.na(new_classes) & nzchar(new_classes)]
+ if (length(new_classes) > 0) {
+ rv$class2use <- c(rv$class2use, new_classes)
+ # Update relabel dropdown
+ sorted_classes <- sort(rv$class2use)
+ updateSelectizeInput(session, "new_class_quick",
+ choices = sorted_classes,
+ selected = character(0))
+ }
+
+ # Update class filter dropdown
+ available_classes <- sort(unique(rv$classifications$class_name))
+ updateSelectInput(session, "class_filter",
+ choices = build_class_filter_choices(available_classes),
+ selected = "all")
+
+ n_predicted <- nrow(predictions)
+ n_skipped <- length(manually_changed)
+ msg <- paste0("Predicted ", n_predicted, " images.")
+ if (n_skipped > 0) {
+ msg <- paste0(msg, " Skipped ", n_skipped, " manually reclassified images.")
+ }
+ showNotification(msg, type = "message", duration = 8)
+ })
+
# Import .mat -> SQLite bulk handler
observeEvent(input$import_mat_to_db_btn, {
if (is.null(config$output_folder) || config$output_folder == "") {
@@ -812,95 +1138,414 @@ server <- function(input, output, session) {
type = "error")
return()
}
- if (is.null(config$output_folder) || config$output_folder == "") {
- showNotification("Output folder is not configured. Set it in Settings first.",
- type = "error")
- return()
- }
db_path <- get_db_path(config$db_folder)
- current_roi_map <- roi_path_map()
+ is_dashboard <- identical(config$data_source, "dashboard")
- if (length(current_roi_map) == 0) {
- showNotification("No ROI file index available. Click Sync first.",
- type = "error")
- return()
- }
+ if (is_dashboard) {
+ # Dashboard mode: copy PNGs from cache instead of extracting from ROI
+ samples <- list_annotated_samples_db(db_path)
+ if (length(samples) == 0) {
+ showNotification("No annotated samples in database.", type = "warning")
+ return()
+ }
+
+ cache_dir <- get_dashboard_cache_dir()
+ parsed <- parse_dashboard_url(config$dashboard_url)
+
+ skip <- if (!is.null(config$skip_class_png) && nzchar(config$skip_class_png)) {
+ config$skip_class_png
+ } else {
+ NULL
+ }
+
+ counts <- list(success = 0L, failed = 0L, skipped = 0L)
+
+ # Bulk download all needed zip archives in one batched call
+ # (uses parallel_downloads internally, much faster than one-at-a-time)
+ withProgress(message = "Downloading images from dashboard...", value = 0, {
+ cached_samples <- download_dashboard_images_bulk(
+ parsed$base_url, samples, cache_dir,
+ parallel_downloads = config$dashboard_parallel_downloads,
+ sleep_time = config$dashboard_sleep_time,
+ multi_timeout = config$dashboard_multi_timeout,
+ max_retries = config$dashboard_max_retries)
+ })
+
+ withProgress(message = "Copying PNGs to class folders...", value = 0, {
+ con <- dbConnect(SQLite(), db_path)
+ on.exit(dbDisconnect(con), add = TRUE)
+
+ for (sn in samples) {
+ rows <- dbGetQuery(con,
+ "SELECT roi_number, class_name FROM annotations WHERE sample_name = ? ORDER BY roi_number",
+ params = list(sn))
+
+ if (nrow(rows) == 0) {
+ counts$skipped <- counts$skipped + 1L
+ next
+ }
+
+ if (!is.null(skip)) {
+ rows <- rows[!rows$class_name %in% skip, ]
+ if (nrow(rows) == 0) next
+ }
- skip <- if (!is.null(input$cfg_skip_class_png) && nzchar(input$cfg_skip_class_png)) {
- input$cfg_skip_class_png
+ src_dir <- file.path(cache_dir, sn, sn)
+ if (!sn %in% cached_samples || !dir.exists(src_dir)) {
+ counts$skipped <- counts$skipped + 1L
+ next
+ }
+
+ ok <- tryCatch({
+ for (cls in unique(rows$class_name)) {
+ cls_rois <- rows$roi_number[rows$class_name == cls]
+ dest <- file.path(config$png_output_folder, cls)
+ dir.create(dest, recursive = TRUE, showWarnings = FALSE)
+ for (rn in cls_rois) {
+ fname <- sprintf("%s_%05d.png", sn, rn)
+ src <- file.path(src_dir, fname)
+ if (file.exists(src)) {
+ file.copy(src, file.path(dest, fname), overwrite = TRUE)
+ }
+ }
+ }
+ TRUE
+ }, error = function(e) FALSE)
+
+ if (isTRUE(ok)) {
+ counts$success <- counts$success + 1L
+ } else {
+ counts$failed <- counts$failed + 1L
+ }
+
+ incProgress(1 / length(samples))
+ }
+ })
+
+ showNotification(
+ sprintf("PNG export complete: %d exported, %d failed, %d skipped.",
+ counts$success, counts$failed, counts$skipped),
+ type = if (counts$failed > 0) "warning" else "message",
+ duration = 8
+ )
} else {
- NULL
- }
+ # Local mode: extract from ROI files
+ if (is.null(config$output_folder) || config$output_folder == "") {
+ showNotification("Output folder is not configured. Set it in Settings first.",
+ type = "error")
+ return()
+ }
- withProgress(message = "Exporting PNGs from SQLite...", {
- result <- export_all_db_to_png(db_path, config$png_output_folder,
- current_roi_map, skip_class = skip)
- })
+ current_roi_map <- roi_path_map()
- showNotification(
- sprintf("PNG export complete: %d exported, %d failed, %d skipped (ROI not found).",
- result$success, result$failed, result$skipped),
- type = if (result$failed > 0) "warning" else "message",
- duration = 8
- )
- })
+ if (length(current_roi_map) == 0) {
+ showNotification("No ROI file index available. Click Sync first.",
+ type = "error")
+ return()
+ }
- # ============================================================================
- # UI Outputs - Warnings and Indicators
- # ============================================================================
-
- output$cache_age_text <- renderUI({
- invalidateLater(60000)
- ts <- last_sync_time()
- if (!is.null(ts)) {
- cache_time <- as.POSIXct(ts)
- age_secs <- as.numeric(difftime(Sys.time(), cache_time, units = "secs"))
- age_text <- if (age_secs < 60) {
- "just now"
- } else if (age_secs < 3600) {
- paste0(round(age_secs / 60), " min ago")
- } else if (age_secs < 86400) {
- paste0(round(age_secs / 3600), " hours ago")
+ skip <- if (!is.null(config$skip_class_png) && nzchar(config$skip_class_png)) {
+ config$skip_class_png
} else {
- paste0(round(age_secs / 86400), " days ago")
+ NULL
}
- div(
- style = "font-size: 11px; color: #999; margin-bottom: 5px;",
- icon("clock", style = "margin-right: 3px;"),
- paste0("Last folder sync: ", age_text)
+
+ withProgress(message = "Exporting PNGs from SQLite...", {
+ result <- export_all_db_to_png(db_path, config$png_output_folder,
+ current_roi_map, skip_class = skip)
+ })
+
+ showNotification(
+ sprintf("PNG export complete: %d exported, %d failed, %d skipped (ROI not found).",
+ result$success, result$failed, result$skipped),
+ type = if (result$failed > 0) "warning" else "message",
+ duration = 8
)
}
})
-
- output$python_warning <- renderUI({
- needs_python <- config$save_format %in% c("mat", "both")
- if (!python_available && needs_python) {
+
+ # ============================================================================
+ # Import PNG -> SQLite (multi-step flow)
+ # ============================================================================
+
+ # Temporary storage for the PNG import flow
+ png_import_state <- reactiveValues(
+ scan_result = NULL,
+ class_mapping = NULL,
+ png_folder = NULL
+ )
+
+ # Step 1: Button click - show folder picker dialog
+ observeEvent(input$import_png_to_db_btn, {
+ showModal(modalDialog(
+ title = "Import PNG \u2192 SQLite",
+ size = "m",
+ easyClose = TRUE,
div(
- class = "alert alert-warning",
- style = "margin-top: 10px; padding: 8px; font-size: 12px;",
- "Python not available. Saving .mat files will not work. ",
- "Switch to SQLite storage format in Settings, or install Python: ",
- "run ifcb_py_install() in R console. ",
- "MAT files are only needed for ",
- tags$a(href = "https://github.com/hsosik/ifcb-analysis", target = "_blank", "ifcb-analysis"),
- " compatibility."
+ style = "display: flex; gap: 5px; align-items: flex-end; margin-bottom: 15px;",
+ div(style = "flex: 1;",
+ textInput("cfg_png_import_folder", "PNG Import Folder",
+ value = config$png_output_folder, width = "100%")),
+ shinyDirButton("browse_png_import_folder", "Browse", "Select PNG Import Folder",
+ class = "btn-outline-secondary", style = "margin-bottom: 15px;")
+ ),
+ tags$small(class = "text-muted",
+ "Select a folder containing PNG images organized in class-name subfolders.",
+ "Folder names follow iRfcb convention (trailing _NNN suffix is stripped)."),
+ footer = tagList(
+ modalButton("Cancel"),
+ actionButton("scan_png_folder_btn", "Scan Folder", class = "btn-primary")
)
- }
+ ))
})
-
- # Send pixels_per_micron to JavaScript for measure tool
- observe({
- session$sendCustomMessage("updatePixelsPerMicron", config$pixels_per_micron)
+
+ # Set up shinyFiles dir chooser for PNG import folder
+ shinyDirChoose(input, "browse_png_import_folder",
+ roots = make_dynamic_roots("cfg_png_import_folder"), session = session)
+
+ observeEvent(input$browse_png_import_folder, {
+ if (!is.integer(input$browse_png_import_folder)) {
+ folder <- parseDirPath(get_browse_volumes(input$cfg_png_import_folder),
+ input$browse_png_import_folder)
+ if (length(folder) > 0) {
+ updateTextInput(session, "cfg_png_import_folder", value = as.character(folder))
+ }
+ }
})
-
- # Loading overlay (shown during load/save operations)
- output$loading_overlay <- renderUI({
- if (rv$is_loading) {
- div(
- class = "loading-overlay",
- div(
- style = "text-align: center;",
+
+ # Step 1b: Scan the folder
+ observeEvent(input$scan_png_folder_btn, {
+ png_folder <- input$cfg_png_import_folder
+ if (is.null(png_folder) || !nzchar(png_folder) || !dir.exists(png_folder)) {
+ showNotification("Please select a valid folder.", type = "error")
+ return()
+ }
+
+ withProgress(message = "Scanning PNG folder...", {
+ scan_result <- tryCatch(
+ scan_png_class_folder(png_folder),
+ error = function(e) {
+ showNotification(paste("Scan failed:", e$message), type = "error")
+ NULL
+ }
+ )
+ })
+
+ if (is.null(scan_result) || nrow(scan_result$annotations) == 0) {
+ showNotification("No valid PNG images found in the selected folder.", type = "error")
+ return()
+ }
+
+ png_import_state$scan_result <- scan_result
+ png_import_state$png_folder <- png_folder
+ png_import_state$class_mapping <- NULL
+
+ # Check for class mismatches
+ unmatched <- setdiff(scan_result$classes_found, rv$class2use)
+
+ if (length(unmatched) > 0) {
+ # Step 2: Show class mapping dialog
+ mapping_inputs <- lapply(unmatched, function(cls) {
+ choices <- c("Add as new" = "__add_new__", setNames(rv$class2use, rv$class2use))
+ div(
+ style = "display: flex; gap: 10px; align-items: center; margin-bottom: 5px;",
+ tags$span(style = "flex: 0 0 200px; font-weight: bold;", cls),
+ div(style = "flex: 1;",
+ selectInput(paste0("png_map_", gsub("[^a-zA-Z0-9]", "_", cls)),
+ label = NULL, choices = choices, width = "100%"))
+ )
+ })
+
+ showModal(modalDialog(
+ title = "Map Unmatched Classes",
+ size = "l",
+ easyClose = FALSE,
+ p(sprintf("Found %d class(es) not in your current class list. Map them to existing classes or add as new:",
+ length(unmatched))),
+ p(sprintf("Scanned: %d images across %d samples in %d classes.",
+ nrow(scan_result$annotations),
+ length(scan_result$sample_names),
+ length(scan_result$classes_found))),
+ hr(),
+ tagList(mapping_inputs),
+ footer = tagList(
+ modalButton("Cancel"),
+ actionButton("confirm_png_class_mapping_btn", "Continue", class = "btn-primary")
+ )
+ ))
+ } else {
+ # No mismatches - proceed to overwrite check (Step 3)
+ check_png_import_overwrite()
+ }
+ })
+
+ # Step 2b: Process class mapping
+ observeEvent(input$confirm_png_class_mapping_btn, {
+ scan_result <- png_import_state$scan_result
+ if (is.null(scan_result)) return()
+
+ unmatched <- setdiff(scan_result$classes_found, rv$class2use)
+ class_mapping <- character()
+ new_classes <- character()
+
+ for (cls in unmatched) {
+ input_id <- paste0("png_map_", gsub("[^a-zA-Z0-9]", "_", cls))
+ mapped_to <- input[[input_id]]
+ if (!is.null(mapped_to) && mapped_to != "__add_new__") {
+ class_mapping[cls] <- mapped_to
+ } else {
+ new_classes <- c(new_classes, cls)
+ }
+ }
+
+ png_import_state$class_mapping <- if (length(class_mapping) > 0) class_mapping else NULL
+
+ # Add new classes to class2use
+ if (length(new_classes) > 0) {
+ rv$class2use <- c(rv$class2use, new_classes)
+ showNotification(
+ sprintf("Added %d new class(es): %s", length(new_classes),
+ paste(new_classes, collapse = ", ")),
+ type = "message", duration = 5
+ )
+ }
+
+ check_png_import_overwrite()
+ })
+
+ # Step 3: Check for existing samples and show overwrite warning
+ check_png_import_overwrite <- function() {
+ scan_result <- png_import_state$scan_result
+ if (is.null(scan_result)) return()
+
+ db_path <- get_db_path(config$db_folder)
+ existing <- list_annotated_samples_db(db_path)
+ overlapping <- intersect(scan_result$sample_names, existing)
+
+ if (length(overlapping) > 0) {
+ showModal(modalDialog(
+ title = "Overwrite Existing Samples?",
+ size = "m",
+ easyClose = FALSE,
+ p(sprintf("%d of %d samples already have annotations in the database and will be overwritten:",
+ length(overlapping), length(scan_result$sample_names))),
+ div(
+ style = "max-height: 200px; overflow-y: auto; background: #f8f9fa; padding: 10px; border-radius: 4px;",
+ tags$ul(lapply(overlapping, tags$li))
+ ),
+ footer = tagList(
+ modalButton("Cancel"),
+ actionButton("confirm_png_overwrite_btn", "Overwrite & Import",
+ class = "btn-warning")
+ )
+ ))
+ } else {
+ # No overlap - proceed directly
+ execute_png_import()
+ }
+ }
+
+ observeEvent(input$confirm_png_overwrite_btn, {
+ execute_png_import()
+ })
+
+ # Step 4: Execute the import
+ execute_png_import <- function() {
+ removeModal()
+
+ scan_result <- png_import_state$scan_result
+ png_folder <- png_import_state$png_folder
+ if (is.null(scan_result) || is.null(png_folder)) return()
+
+ db_path <- get_db_path(config$db_folder)
+ annotator <- if (!is.null(input$annotator_name) && nzchar(input$annotator_name)) {
+ input$annotator_name
+ } else {
+ "imported"
+ }
+
+ withProgress(message = "Importing PNG annotations to SQLite...", {
+ result <- import_png_folder_to_db(
+ png_folder, db_path, rv$class2use,
+ class_mapping = png_import_state$class_mapping,
+ annotator = annotator
+ )
+ })
+
+ showNotification(
+ sprintf("PNG import complete: %d samples imported, %d failed.",
+ result$success, result$failed),
+ type = if (result$failed > 0) "warning" else "message",
+ duration = 8
+ )
+
+ # Trigger rescan to refresh sample list
+ if (result$success > 0) {
+ rescan_trigger(rescan_trigger() + 1)
+ }
+
+ # Clean up state
+ png_import_state$scan_result <- NULL
+ png_import_state$class_mapping <- NULL
+ png_import_state$png_folder <- NULL
+ }
+
+ # ============================================================================
+ # UI Outputs - Warnings and Indicators
+ # ============================================================================
+
+ output$cache_age_text <- renderUI({
+ invalidateLater(60000)
+ ts <- last_sync_time()
+ if (!is.null(ts)) {
+ cache_time <- as.POSIXct(ts)
+ age_secs <- as.numeric(difftime(Sys.time(), cache_time, units = "secs"))
+ age_text <- if (age_secs < 60) {
+ "just now"
+ } else if (age_secs < 3600) {
+ paste0(round(age_secs / 60), " min ago")
+ } else if (age_secs < 86400) {
+ paste0(round(age_secs / 3600), " hours ago")
+ } else {
+ paste0(round(age_secs / 86400), " days ago")
+ }
+ div(
+ style = "font-size: 11px; color: #999; margin-bottom: 5px;",
+ icon("clock", style = "margin-right: 3px;"),
+ paste0("Last synced ", age_text)
+ )
+ }
+ })
+
+ output$python_warning <- renderUI({
+ needs_python <- config$save_format %in% c("mat", "both")
+ if (!python_available && needs_python) {
+ div(
+ class = "alert alert-warning",
+ style = "margin-top: 10px; padding: 8px; font-size: 12px;",
+ "Python not available. Saving .mat files will not work. ",
+ "Switch to SQLite storage format in Settings, or install Python: ",
+ "run ifcb_py_install() in R console. ",
+ "MAT files are only needed for ",
+ tags$a(href = "https://github.com/hsosik/ifcb-analysis", target = "_blank", "ifcb-analysis"),
+ " compatibility."
+ )
+ }
+ })
+
+ # Send pixels_per_micron to JavaScript for measure tool
+ observe({
+ session$sendCustomMessage("updatePixelsPerMicron", config$pixels_per_micron)
+ })
+
+ # Loading overlay (shown during load/save operations)
+ output$loading_overlay <- renderUI({
+ if (rv$is_loading) {
+ div(
+ class = "loading-overlay",
+ div(
+ style = "text-align: center;",
div(class = "spinner-border text-primary", role = "status",
style = "width: 3rem; height: 3rem;"),
div(style = "margin-top: 10px; font-weight: bold;", "Loading...")
@@ -912,22 +1557,26 @@ server <- function(input, output, session) {
# Dynamic title with mode-based navbar coloring
output$dynamic_title <- renderUI({
# Determine mode class for navbar styling
- mode_class <- if (is.null(rv$current_sample)) {
+ mode_class <- if (rv$class_review_mode) {
+ "navbar-mode-class-review"
+ } else if (is.null(rv$current_sample)) {
"navbar-mode-none"
} else if (rv$is_annotation_mode) {
"navbar-mode-annotation"
} else {
"navbar-mode-validation"
}
-
+
+ all_mode_classes <- "navbar-mode-none navbar-mode-annotation navbar-mode-validation navbar-mode-class-review"
+
# Add JavaScript to apply class to navbar
tagList(
tags$script(HTML(sprintf("
$(document).ready(function() {
- $('.navbar').removeClass('navbar-mode-none navbar-mode-annotation navbar-mode-validation').addClass('%s');
+ $('.navbar').removeClass('%s').addClass('%s');
});
- $('.navbar').removeClass('navbar-mode-none navbar-mode-annotation navbar-mode-validation').addClass('%s');
- ", mode_class, mode_class))),
+ $('.navbar').removeClass('%s').addClass('%s');
+ ", all_mode_classes, mode_class, all_mode_classes, mode_class))),
div(
style = "display: flex; align-items: baseline; gap: 20px;",
actionLink(
@@ -942,7 +1591,25 @@ server <- function(input, output, session) {
})
output$mode_indicator_inline <- renderUI({
- if (is.null(rv$current_sample)) {
+ if (rv$class_review_mode) {
+ n_images <- if (!is.null(rv$classifications)) nrow(rv$classifications) else 0
+ n_samples <- length(rv$class_review_samples)
+ n_changed <- 0L
+ if (!is.null(rv$class_review_original) && !is.null(rv$classifications)) {
+ n_changed <- sum(rv$classifications$class_name != rv$class_review_original$class_name)
+ }
+ change_text <- if (n_changed > 0) sprintf(", %d changed", n_changed) else ""
+
+ span(
+ style = "font-size: 14px; color: white;",
+ tags$span(style = "font-weight: bold; margin-right: 8px;", "CLASS REVIEW"),
+ tags$span(rv$class_review_class),
+ tags$span(
+ style = "margin-left: 10px; opacity: 0.9;",
+ sprintf("(%d images, %d samples%s)", n_images, n_samples, change_text)
+ )
+ )
+ } else if (is.null(rv$current_sample)) {
span(
style = "font-size: 14px; color: white; font-weight: 500;",
"No sample loaded"
@@ -1016,13 +1683,21 @@ server <- function(input, output, session) {
}
adc_path <- sub("\\.roi$", ".adc", roi_path)
- # Find classification source (CSV or classifier MAT)
+ # Find classification source (CSV > H5 > MAT)
csv_path <- find_csv_file(sample_name)
+ classifier_h5_path <- find_classifier_h5(sample_name)
classifier_mat_path <- classifier_mat_files()[[sample_name]]
if (!is.null(csv_path)) {
- classifications <- load_from_csv(csv_path)
+ classifications <- load_from_csv(csv_path, use_threshold = config$use_threshold)
showNotification("Switched to Validation mode (CSV)", type = "message")
+ } else if (!is.null(classifier_h5_path)) {
+ roi_dims <- read_roi_dimensions(adc_path)
+ classifications <- load_from_h5(
+ classifier_h5_path, sample_name, roi_dims,
+ use_threshold = config$use_threshold
+ )
+ showNotification("Switched to Validation mode (H5)", type = "message")
} else if (!is.null(classifier_mat_path)) {
roi_dims <- read_roi_dimensions(adc_path)
classifications <- load_from_classifier_mat(
@@ -1216,62 +1891,110 @@ server <- function(input, output, session) {
roi_path_map(safe_list(index_data$roi_path_map))
csv_path_map(safe_list(index_data$csv_path_map))
classifier_mat_files(safe_list(index_data$classifier_mat_files))
-
+ classifier_h5_files(safe_list(index_data$classifier_h5_files))
+
years <- unique(substr(sample_names, 2, 5))
years <- sort(years)
first_year <- if (length(years) > 0) years[1] else "all"
updateSelectInput(session, "year_select",
choices = c("All" = "all", setNames(years, years)),
selected = first_year)
+
+ instruments <- unique(sub(".*_", "", sample_names))
+ instruments <- sort(instruments)
+ updateSelectInput(session, "instrument_select",
+ choices = c("All" = "all", setNames(instruments, instruments)),
+ selected = "all")
last_sync_time(index_data$timestamp)
TRUE
}
- # Scan for available ROI files and classification files (CSV and MAT)
+ # Scan for available ROI files and classification files (CSV, H5, and MAT)
# Uses disk cache for fast startup on subsequent launches
observe({
rescan_trigger() # Force dependency on rescan trigger
- roi_folder <- config$roi_folder
- csv_folder <- config$csv_folder
- output_folder <- config$output_folder
-
- # Validate folder paths before using them
- roi_valid <- !is.null(roi_folder) && length(roi_folder) == 1 && !isTRUE(is.na(roi_folder)) && nzchar(roi_folder) && dir.exists(roi_folder)
-
- if (!roi_valid) return()
-
- # Try loading from cache first
- cached <- load_file_index()
- cache_valid <- !is.null(cached) &&
- identical(cached$roi_folder, roi_folder) &&
- identical(cached$csv_folder, csv_folder) &&
- identical(cached$output_folder, output_folder)
-
- if (cache_valid) {
- populate_from_index(cached)
- return()
- }
-
- # When auto-sync is disabled, load stale cache if available
- auto_sync <- config$auto_sync
- if (!isTRUE(auto_sync) && !is.null(cached)) {
- populate_from_index(cached)
- return()
- }
-
- # Full scan with progress indicator (delegates to rescan_file_index)
- withProgress(message = "Syncing folders...", value = 0, {
- result <- rescan_file_index(
- roi_folder = roi_folder,
- csv_folder = csv_folder,
- output_folder = output_folder,
- verbose = FALSE
- )
- })
-
- if (!is.null(result)) {
- populate_from_index(result)
+ data_source <- config$data_source
+
+ if (identical(data_source, "dashboard")) {
+ # Dashboard mode: fetch sample list from remote API
+ dashboard_url <- config$dashboard_url
+ if (is.null(dashboard_url) || !nzchar(dashboard_url)) return()
+
+ # Try loading from cache first
+ cached <- load_file_index()
+ cache_valid <- !is.null(cached) &&
+ identical(cached$data_source, "dashboard") &&
+ identical(cached$dashboard_url, dashboard_url)
+
+ if (cache_valid) {
+ populate_from_index(cached)
+ return()
+ }
+
+ # When auto-sync is disabled, load stale cache if available
+ auto_sync <- config$auto_sync
+ if (!isTRUE(auto_sync) && !is.null(cached) &&
+ identical(cached$data_source, "dashboard")) {
+ populate_from_index(cached)
+ return()
+ }
+
+ withProgress(message = "Fetching dashboard samples...", value = 0, {
+ result <- rescan_file_index(
+ data_source = "dashboard",
+ dashboard_url = dashboard_url,
+ db_folder = config$db_folder,
+ verbose = FALSE
+ )
+ })
+
+ if (!is.null(result)) {
+ populate_from_index(result)
+ }
+ } else {
+ # Local mode: scan folders
+ roi_folder <- config$roi_folder
+ csv_folder <- config$csv_folder
+ output_folder <- config$output_folder
+
+ # Validate folder paths before using them
+ roi_valid <- !is.null(roi_folder) && length(roi_folder) == 1 && !isTRUE(is.na(roi_folder)) && nzchar(roi_folder) && dir.exists(roi_folder)
+
+ if (!roi_valid) return()
+
+ # Try loading from cache first
+ cached <- load_file_index()
+ cache_valid <- !is.null(cached) &&
+ identical(cached$roi_folder, roi_folder) &&
+ identical(cached$csv_folder, csv_folder) &&
+ identical(cached$output_folder, output_folder)
+
+ if (cache_valid) {
+ populate_from_index(cached)
+ return()
+ }
+
+ # When auto-sync is disabled, load stale cache if available
+ auto_sync <- config$auto_sync
+ if (!isTRUE(auto_sync) && !is.null(cached)) {
+ populate_from_index(cached)
+ return()
+ }
+
+ # Full scan with progress indicator (delegates to rescan_file_index)
+ withProgress(message = "Syncing folders...", value = 0, {
+ result <- rescan_file_index(
+ roi_folder = roi_folder,
+ csv_folder = csv_folder,
+ output_folder = output_folder,
+ verbose = FALSE
+ )
+ })
+
+ if (!is.null(result)) {
+ populate_from_index(result)
+ }
}
})
@@ -1295,37 +2018,57 @@ server <- function(input, output, session) {
rescan_trigger(rescan_trigger() + 1)
})
- # Helper function to update month choices based on year selection
+ # Helper function to update month and instrument choices based on year selection
update_month_choices <- function() {
samples <- all_samples()
if (length(samples) == 0) return()
-
+
year_val <- input$year_select
-
+
if (!is.null(year_val) && year_val != "all") {
# Filter to selected year
year_pattern <- paste0("^D", year_val)
year_samples <- samples[grepl(year_pattern, samples)]
-
+
# Extract months (characters 6-7 of sample name: DYYYYMMDD...)
months <- unique(substr(year_samples, 6, 7))
months <- sort(months)
-
+
# Create month names
month_names <- c("01" = "Jan", "02" = "Feb", "03" = "Mar", "04" = "Apr",
"05" = "May", "06" = "Jun", "07" = "Jul", "08" = "Aug",
"09" = "Sep", "10" = "Oct", "11" = "Nov", "12" = "Dec")
month_labels <- month_names[months]
-
+
# Auto-select first month for better UX with large sample lists
first_month <- if (length(months) > 0) months[1] else "all"
updateSelectInput(session, "month_select",
choices = c("All" = "all", setNames(months, month_labels)),
selected = first_month)
+
+ # Update instrument choices for selected year
+ instruments <- unique(sub(".*_", "", year_samples))
+ instruments <- sort(instruments)
+ current_instrument <- input$instrument_select
+ selected_instrument <- if (!is.null(current_instrument) && current_instrument %in% instruments) {
+ current_instrument
+ } else {
+ "all"
+ }
+ updateSelectInput(session, "instrument_select",
+ choices = c("All" = "all", setNames(instruments, instruments)),
+ selected = selected_instrument)
} else {
updateSelectInput(session, "month_select",
choices = c("All" = "all"),
selected = "all")
+
+ # Show all instruments when no year filter
+ instruments <- unique(sub(".*_", "", samples))
+ instruments <- sort(instruments)
+ updateSelectInput(session, "instrument_select",
+ choices = c("All" = "all", setNames(instruments, instruments)),
+ selected = "all")
}
}
@@ -1351,7 +2094,14 @@ server <- function(input, output, session) {
month_pattern <- paste0("^D\\d{4}", month_val)
samples <- samples[grepl(month_pattern, samples)]
}
-
+
+ # Filter by instrument
+ instrument_val <- input$instrument_select
+ if (!is.null(instrument_val) && instrument_val != "all") {
+ instrument_pattern <- paste0("_", instrument_val, "$")
+ samples <- samples[grepl(instrument_pattern, samples)]
+ }
+
# Filter by classification status
if (!is.null(status_val)) {
if (status_val == "classified") {
@@ -1476,7 +2226,11 @@ server <- function(input, output, session) {
observeEvent(input$month_select, {
update_sample_list()
}, ignoreInit = TRUE, ignoreNULL = TRUE)
-
+
+ observeEvent(input$instrument_select, {
+ update_sample_list()
+ }, ignoreInit = TRUE, ignoreNULL = TRUE)
+
observeEvent(input$sample_status_filter, {
update_sample_list()
}, ignoreInit = TRUE, ignoreNULL = TRUE)
@@ -1502,7 +2256,12 @@ server <- function(input, output, session) {
month_pattern <- paste0("^D\\d{4}", input$month_select)
samples <- samples[grepl(month_pattern, samples)]
}
-
+
+ if (!is.null(input$instrument_select) && input$instrument_select != "all") {
+ instrument_pattern <- paste0("_", input$instrument_select, "$")
+ samples <- samples[grepl(instrument_pattern, samples)]
+ }
+
if (!is.null(input$sample_status_filter)) {
if (input$sample_status_filter == "classified") {
samples <- samples[samples %in% classified & !samples %in% annotated]
@@ -1530,7 +2289,7 @@ server <- function(input, output, session) {
})
# ============================================================================
- # Helper: Find classification file (CSV or classifier MAT)
+ # Helper: Find classification file (CSV, H5, or classifier MAT)
# ============================================================================
find_csv_file <- function(sample_name) {
@@ -1550,6 +2309,15 @@ server <- function(input, output, session) {
}
return(NULL)
}
+
+ # Find classifier H5 file for a sample
+ find_classifier_h5 <- function(sample_name) {
+ h5_map <- classifier_h5_files()
+ if (sample_name %in% names(h5_map)) {
+ return(h5_map[[sample_name]])
+ }
+ return(NULL)
+ }
# ============================================================================
# Sample Loading
@@ -1577,6 +2345,15 @@ server <- function(input, output, session) {
tryCatch({
roi_path_for_save <- roi_path_map()[[rv$current_sample]]
adc_folder_for_save <- if (!is.null(roi_path_for_save)) dirname(roi_path_for_save) else NULL
+
+ # In dashboard mode, fall back to SQLite if no ADC folder
+ save_fmt_for_autosave <- config$save_format
+ if (identical(config$data_source, "dashboard") && is.null(adc_folder_for_save)) {
+ if (save_fmt_for_autosave %in% c("mat", "both")) {
+ save_fmt_for_autosave <- "sqlite"
+ }
+ }
+
saved <- save_sample_annotations(
sample_name = rv$current_sample,
classifications = rv$classifications,
@@ -1590,7 +2367,7 @@ server <- function(input, output, session) {
class2use = rv$class2use,
annotator = input$annotator_name,
adc_folder = adc_folder_for_save,
- save_format = config$save_format,
+ save_format = save_fmt_for_autosave,
db_folder = config$db_folder,
export_statistics = config$export_statistics
)
@@ -1611,40 +2388,261 @@ server <- function(input, output, session) {
# Main sample loading function
load_sample_data <- function(sample_name) {
req(rv$class2use)
-
- # Find classification files
- csv_path <- find_csv_file(sample_name)
- classifier_mat_path <- find_classifier_mat(sample_name)
- has_csv <- !is.null(csv_path)
- has_classifier_mat <- !is.null(classifier_mat_path)
-
- # Use discovered paths from scan (supports any folder structure)
- roi_path <- roi_path_map()[[sample_name]]
- if (is.null(roi_path) || !file.exists(roi_path)) {
- showNotification(paste("ROI file not found for:", sample_name), type = "error")
- return(FALSE)
- }
- adc_path <- sub("\\.roi$", ".adc", roi_path)
-
- # Check session cache first
- if (sample_name %in% names(rv$session_cache)) {
- return(load_from_cache(sample_name, roi_path))
- }
-
- tryCatch({
- annotation_mat_path <- file.path(config$output_folder, paste0(sample_name, ".mat"))
- db_path <- get_db_path(config$db_folder)
- has_db_annotation <- sample_name %in% list_annotated_samples_db(db_path)
- has_mat_annotation <- file.exists(annotation_mat_path)
- has_existing_annotation <- has_db_annotation || has_mat_annotation
- has_classification <- has_csv || has_classifier_mat
- # Track if sample has both modes available
- rv$has_both_modes <- has_existing_annotation && has_classification
- rv$using_manual_mode <- has_existing_annotation # Default to manual if available
+ is_dashboard <- identical(config$data_source, "dashboard")
- # Variable to hold mode message for notification (shown after filtering)
- mode_message <- NULL
+ # --- Dashboard mode ---
+ if (is_dashboard) {
+ # Check session cache first
+ if (sample_name %in% names(rv$session_cache)) {
+ return(load_from_cache(sample_name, NULL))
+ }
+
+ tryCatch({
+ parsed <- parse_dashboard_url(config$dashboard_url)
+ cache_dir <- get_dashboard_cache_dir()
+ db_path <- get_db_path(config$db_folder)
+ has_db_annotation <- sample_name %in% list_annotated_samples_db(db_path)
+
+ # Download PNG images from dashboard
+ png_folder <- withProgress(message = "Downloading images...", {
+ download_dashboard_images(parsed$base_url, sample_name, cache_dir,
+ parallel_downloads = config$dashboard_parallel_downloads,
+ sleep_time = config$dashboard_sleep_time,
+ multi_timeout = config$dashboard_multi_timeout,
+ max_retries = config$dashboard_max_retries)
+ })
+
+ if (is.null(png_folder)) {
+ showNotification(paste("Failed to download images for:", sample_name), type = "error")
+ return(FALSE)
+ }
+
+ # Download ADC on demand for dimensions (graceful fallback to NA)
+ adc_path <- download_dashboard_adc(parsed$base_url, sample_name, cache_dir,
+ parallel_downloads = config$dashboard_parallel_downloads,
+ sleep_time = config$dashboard_sleep_time,
+ multi_timeout = config$dashboard_multi_timeout,
+ max_retries = config$dashboard_max_retries)
+ roi_dims <- if (!is.null(adc_path) && file.exists(adc_path)) {
+ tryCatch(read_roi_dimensions(adc_path), error = function(e) NULL)
+ } else {
+ NULL
+ }
+
+ mode_message <- NULL
+ classifications <- NULL
+
+ if (has_db_annotation) {
+ # Load from existing DB annotations
+ if (is.null(roi_dims)) {
+ # Build minimal roi_dims from PNG files
+ png_files <- list.files(file.path(png_folder, sample_name), pattern = "\\.png$")
+ roi_numbers <- as.integer(gsub(".*_(\\d+)\\.png$", "\\1", png_files))
+ roi_dims <- data.frame(
+ roi_number = roi_numbers,
+ width = NA_real_, height = NA_real_, area = NA_real_
+ )
+ }
+ classifications <- load_from_db(db_path, sample_name, roi_dims)
+ rv$is_annotation_mode <- TRUE
+ rv$has_both_modes <- isTRUE(config$dashboard_autoclass)
+ rv$using_manual_mode <- TRUE
+ mode_message <- if (rv$has_both_modes) "Manual mode (switch available)" else "Resumed"
+
+ }
+
+ # Try local classification files if csv_folder is configured
+ if (is.null(classifications)) {
+ csv_folder <- config$csv_folder
+ has_csv_folder <- !is.null(csv_folder) && nzchar(csv_folder) && dir.exists(csv_folder)
+
+ if (has_csv_folder) {
+ # Search for classification files in csv_folder (recursive)
+ local_csv <- list.files(csv_folder, pattern = paste0("^", sample_name, "\\.csv$"),
+ recursive = TRUE, full.names = TRUE)
+ local_h5 <- list.files(csv_folder, pattern = paste0("^", sample_name, ".*\\.h5$"),
+ recursive = TRUE, full.names = TRUE)
+ local_mat <- list.files(csv_folder, pattern = paste0("^", sample_name, ".*\\.mat$"),
+ recursive = TRUE, full.names = TRUE)
+
+ if (length(local_csv) > 0) {
+ classifications <- load_from_csv(local_csv[1], use_threshold = config$use_threshold)
+ rv$is_annotation_mode <- FALSE
+ rv$has_both_modes <- FALSE
+ rv$using_manual_mode <- FALSE
+ threshold_text <- if (config$use_threshold) "with threshold" else "without threshold"
+ mode_message <- paste0("Validation mode (Local CSV, ", threshold_text, ")")
+ } else if (length(local_h5) > 0) {
+ classifications <- load_from_h5(
+ local_h5[1], sample_name, roi_dims,
+ use_threshold = config$use_threshold
+ )
+ rv$is_annotation_mode <- FALSE
+ rv$has_both_modes <- FALSE
+ rv$using_manual_mode <- FALSE
+ threshold_text <- if (config$use_threshold) "with threshold" else "without threshold"
+ mode_message <- paste0("Validation mode (Local H5, ", threshold_text, ")")
+ } else if (length(local_mat) > 0) {
+ classifications <- load_from_classifier_mat(
+ local_mat[1], sample_name, rv$class2use, roi_dims,
+ use_threshold = config$use_threshold
+ )
+ rv$is_annotation_mode <- FALSE
+ rv$has_both_modes <- FALSE
+ rv$using_manual_mode <- FALSE
+ threshold_text <- if (config$use_threshold) "with threshold" else "without threshold"
+ mode_message <- paste0("Validation mode (Local MAT, ", threshold_text, ")")
+ }
+ }
+ }
+
+ if (is.null(classifications) && isTRUE(config$dashboard_autoclass)) {
+ # Try dashboard autoclass for validation mode (fallback)
+ autoclass <- withProgress(message = "Downloading auto-classifications...", {
+ download_dashboard_autoclass(parsed$base_url, sample_name, cache_dir,
+ parallel_downloads = config$dashboard_parallel_downloads,
+ sleep_time = config$dashboard_sleep_time,
+ multi_timeout = config$dashboard_multi_timeout,
+ max_retries = config$dashboard_max_retries)
+ })
+
+ if (!is.null(autoclass) && nrow(autoclass) > 0) {
+ # Merge with ROI dimensions if available
+ if (!is.null(roi_dims)) {
+ roi_numbers <- as.integer(gsub(".*_(\\d+)\\.png$", "\\1", autoclass$file_name))
+ dim_data <- lapply(roi_numbers, function(rn) {
+ idx <- which(roi_dims$roi_number == rn)
+ if (length(idx) > 0) {
+ list(width = roi_dims$width[idx], height = roi_dims$height[idx],
+ area = roi_dims$area[idx])
+ } else {
+ list(width = NA_real_, height = NA_real_, area = NA_real_)
+ }
+ })
+ autoclass$width <- vapply(dim_data, `[[`, numeric(1), "width")
+ autoclass$height <- vapply(dim_data, `[[`, numeric(1), "height")
+ autoclass$roi_area <- vapply(dim_data, `[[`, numeric(1), "area")
+ } else {
+ autoclass$width <- NA_real_
+ autoclass$height <- NA_real_
+ autoclass$roi_area <- NA_real_
+ }
+
+ classifications <- autoclass
+ rv$is_annotation_mode <- FALSE
+ rv$has_both_modes <- FALSE
+ rv$using_manual_mode <- FALSE
+ mode_message <- "Validation mode (Dashboard autoclass)"
+ }
+ }
+
+ if (is.null(classifications)) {
+ # NEW ANNOTATION - no existing annotations or autoclass
+ png_files <- list.files(file.path(png_folder, sample_name), pattern = "\\.png$")
+ if (length(png_files) == 0) {
+ showNotification(paste("No images found for:", sample_name), type = "error")
+ return(FALSE)
+ }
+
+ roi_numbers <- as.integer(gsub(".*_(\\d+)\\.png$", "\\1", png_files))
+
+ if (is.null(roi_dims)) {
+ roi_dims <- data.frame(
+ roi_number = roi_numbers,
+ width = NA_real_, height = NA_real_, area = NA_real_
+ )
+ }
+
+ classifications <- create_new_classifications(sample_name, roi_dims)
+ rv$is_annotation_mode <- TRUE
+ rv$has_both_modes <- FALSE
+ rv$using_manual_mode <- TRUE
+ mode_message <- "New annotation"
+ }
+
+ # Store state
+ rv$original_classifications <- classifications
+ rv$classifications <- classifications
+ rv$current_sample <- sample_name
+ rv$selected_images <- character()
+ rv$current_page <- 1
+ rv$changes_log <- create_empty_changes_log()
+
+ # Set temp_png_folder to the parent (preserves path structure for gallery)
+ if (!is.null(rv$temp_png_folder) && dir.exists(rv$temp_png_folder) &&
+ !startsWith(rv$temp_png_folder, cache_dir)) {
+ unlink(rv$temp_png_folder, recursive = TRUE)
+ }
+ rv$temp_png_folder <- png_folder
+
+ # Filter to actually extracted PNGs
+ extracted_folder <- file.path(png_folder, sample_name)
+ if (dir.exists(extracted_folder)) {
+ extracted_files <- list.files(extracted_folder, pattern = "\\.png$")
+ rv$classifications <- rv$classifications[rv$classifications$file_name %in% extracted_files, ]
+ rv$original_classifications <- rv$original_classifications[
+ rv$original_classifications$file_name %in% extracted_files, ]
+ }
+
+ # Update class filter
+ available_classes <- sort(unique(rv$classifications$class_name))
+ unmatched <- setdiff(available_classes, c(rv$class2use, "unclassified"))
+ display_names <- sapply(available_classes, function(cls) {
+ if (cls %in% unmatched) paste0("\u26A0 ", cls) else cls
+ })
+ updateSelectInput(session, "class_filter",
+ choices = c("All" = "all", setNames(available_classes, display_names)))
+
+ if (!is.null(mode_message)) {
+ actual_count <- nrow(rv$classifications)
+ showNotification(paste0(mode_message, ": ", actual_count, " images"), type = "message")
+ }
+
+ return(TRUE)
+ }, error = function(e) {
+ showNotification(paste("Error loading sample:", e$message), type = "error")
+ return(FALSE)
+ })
+ }
+
+ # --- Local mode ---
+
+ # Find classification files (priority: CSV > H5 > MAT)
+ csv_path <- find_csv_file(sample_name)
+ classifier_h5_path <- find_classifier_h5(sample_name)
+ classifier_mat_path <- find_classifier_mat(sample_name)
+ has_csv <- !is.null(csv_path)
+ has_classifier_h5 <- !is.null(classifier_h5_path)
+ has_classifier_mat <- !is.null(classifier_mat_path)
+
+ # Use discovered paths from scan (supports any folder structure)
+ roi_path <- roi_path_map()[[sample_name]]
+ if (is.null(roi_path) || !file.exists(roi_path)) {
+ showNotification(paste("ROI file not found for:", sample_name), type = "error")
+ return(FALSE)
+ }
+ adc_path <- sub("\\.roi$", ".adc", roi_path)
+
+ # Check session cache first
+ if (sample_name %in% names(rv$session_cache)) {
+ return(load_from_cache(sample_name, roi_path))
+ }
+
+ tryCatch({
+ annotation_mat_path <- file.path(config$output_folder, paste0(sample_name, ".mat"))
+ db_path <- get_db_path(config$db_folder)
+ has_db_annotation <- sample_name %in% list_annotated_samples_db(db_path)
+ has_mat_annotation <- file.exists(annotation_mat_path)
+ has_existing_annotation <- has_db_annotation || has_mat_annotation
+ has_classification <- has_csv || has_classifier_h5 || has_classifier_mat
+
+ # Track if sample has both modes available
+ rv$has_both_modes <- has_existing_annotation && has_classification
+ rv$using_manual_mode <- has_existing_annotation # Default to manual if available
+
+ # Variable to hold mode message for notification (shown after filtering)
+ mode_message <- NULL
# Priority: Manual annotation > Classification > New annotation
# Within manual annotations: SQLite first (faster), then .mat fallback
@@ -1668,41 +2666,59 @@ server <- function(input, output, session) {
} else if (has_csv) {
# VALIDATION MODE - from CSV
- classifications <- load_from_csv(csv_path)
+ classifications <- load_from_csv(csv_path, use_threshold = config$use_threshold)
rv$is_annotation_mode <- FALSE
- mode_message <- "Validation mode (CSV)"
-
+ threshold_text <- if (config$use_threshold) "with threshold" else "without threshold"
+ mode_message <- paste0("Validation mode (CSV, ", threshold_text, ")")
+
+ } else if (has_classifier_h5) {
+ # VALIDATION MODE - from classifier H5 file
+ if (!file.exists(adc_path)) {
+ showNotification(paste("ADC file not found:", adc_path), type = "error")
+ return(FALSE)
+ }
+
+ roi_dims <- read_roi_dimensions(adc_path)
+ classifications <- load_from_h5(
+ classifier_h5_path, sample_name, roi_dims,
+ use_threshold = config$use_threshold
+ )
+ rv$is_annotation_mode <- FALSE
+
+ threshold_text <- if (config$use_threshold) "with threshold" else "without threshold"
+ mode_message <- paste0("Validation mode (H5, ", threshold_text, ")")
+
} else if (has_classifier_mat) {
# VALIDATION MODE - from classifier MAT file
if (!file.exists(adc_path)) {
showNotification(paste("ADC file not found:", adc_path), type = "error")
return(FALSE)
}
-
+
roi_dims <- read_roi_dimensions(adc_path)
classifications <- load_from_classifier_mat(
classifier_mat_path, sample_name, rv$class2use, roi_dims,
use_threshold = config$use_threshold
)
rv$is_annotation_mode <- FALSE
-
+
threshold_text <- if (config$use_threshold) "with threshold" else "without threshold"
mode_message <- paste0("Validation mode (MAT, ", threshold_text, ")")
-
+
} else {
# NEW ANNOTATION
if (!file.exists(adc_path)) {
showNotification(paste("ADC file not found:", adc_path), type = "error")
return(FALSE)
}
-
+
roi_dims <- read_roi_dimensions(adc_path)
classifications <- create_new_classifications(sample_name, roi_dims)
rv$is_annotation_mode <- TRUE
-
+
mode_message <- "New annotation"
}
-
+
# Store state
rv$original_classifications <- classifications
rv$classifications <- classifications
@@ -1710,7 +2726,7 @@ server <- function(input, output, session) {
rv$selected_images <- character()
rv$current_page <- 1
rv$changes_log <- create_empty_changes_log()
-
+
# Update class filter with warnings for unmatched classes
available_classes <- sort(unique(classifications$class_name))
unmatched <- setdiff(available_classes, c(rv$class2use, "unclassified"))
@@ -1719,12 +2735,12 @@ server <- function(input, output, session) {
})
updateSelectInput(session, "class_filter",
choices = c("All" = "all", setNames(available_classes, display_names)))
-
+
# Extract images (notification shown after filtering with correct count)
extract_sample_images(sample_name, roi_path, classifications, mode_message = mode_message)
-
+
return(TRUE)
-
+
}, error = function(e) {
showNotification(paste("Error loading sample:", e$message), type = "error")
return(FALSE)
@@ -1740,7 +2756,7 @@ server <- function(input, output, session) {
rv$current_sample <- sample_name
rv$selected_images <- character()
rv$is_annotation_mode <- cached$is_annotation_mode
-
+
available_classes <- sort(unique(rv$classifications$class_name))
unmatched <- setdiff(available_classes, c(rv$class2use, "unclassified"))
display_names <- sapply(available_classes, function(cls) {
@@ -1748,25 +2764,40 @@ server <- function(input, output, session) {
})
updateSelectInput(session, "class_filter",
choices = c("All" = "all", setNames(available_classes, display_names)))
-
- if (!is.null(rv$temp_png_folder) && dir.exists(rv$temp_png_folder)) {
- unlink(rv$temp_png_folder, recursive = TRUE)
+
+ is_dashboard <- identical(config$data_source, "dashboard")
+
+ if (is_dashboard) {
+ # Dashboard mode: use cached download dir (PNGs already extracted)
+ cache_dir <- get_dashboard_cache_dir()
+ png_folder <- file.path(cache_dir, sample_name)
+
+ if (!is.null(rv$temp_png_folder) && dir.exists(rv$temp_png_folder) &&
+ !startsWith(rv$temp_png_folder, cache_dir)) {
+ unlink(rv$temp_png_folder, recursive = TRUE)
+ }
+ rv$temp_png_folder <- png_folder
+ } else {
+ # Local mode: re-extract from ROI file
+ if (!is.null(rv$temp_png_folder) && dir.exists(rv$temp_png_folder)) {
+ unlink(rv$temp_png_folder, recursive = TRUE)
+ }
+
+ rv$temp_png_folder <- tempfile(pattern = "ifcb_validator_")
+ dir.create(rv$temp_png_folder, recursive = TRUE)
+
+ roi_numbers <- as.numeric(gsub(".*_(\\d+)\\.png$", "\\1", rv$classifications$file_name))
+
+ withProgress(message = "Extracting images...", {
+ ifcb_extract_pngs(
+ roi_file = roi_path,
+ out_folder = rv$temp_png_folder,
+ ROInumbers = roi_numbers,
+ verbose = FALSE
+ )
+ })
}
-
- rv$temp_png_folder <- tempfile(pattern = "ifcb_validator_")
- dir.create(rv$temp_png_folder, recursive = TRUE)
-
- roi_numbers <- as.numeric(gsub(".*_(\\d+)\\.png$", "\\1", rv$classifications$file_name))
-
- withProgress(message = "Extracting images...", {
- ifcb_extract_pngs(
- roi_file = roi_path,
- out_folder = rv$temp_png_folder,
- ROInumbers = roi_numbers,
- verbose = FALSE
- )
- })
-
+
n_changes <- nrow(rv$changes_log)
showNotification(paste("Restored from cache:", n_changes, "changes"), type = "message")
return(TRUE)
@@ -1843,6 +2874,13 @@ server <- function(input, output, session) {
rv$is_loading <- FALSE
enable_nav_buttons()
})
+
+ # Clear class review state when loading a sample
+ rv$class_review_mode <- FALSE
+ rv$class_review_class <- NULL
+ rv$class_review_samples <- character()
+ rv$class_review_original <- NULL
+
save_to_cache()
rv$pending_sample_select <- input$sample_select
load_sample_data(input$sample_select)
@@ -1854,7 +2892,7 @@ server <- function(input, output, session) {
if (!is.null(rv$current_sample)) {
save_to_cache()
}
-
+
# Reset all sample-related state
rv$current_sample <- NULL
rv$classifications <- NULL
@@ -1863,10 +2901,19 @@ server <- function(input, output, session) {
rv$selected_images <- character(0)
rv$is_annotation_mode <- FALSE
rv$has_both_modes <- FALSE
-
+
+ # Reset class review state
+ rv$class_review_mode <- FALSE
+ rv$class_review_class <- NULL
+ rv$class_review_samples <- character()
+ rv$class_review_original <- NULL
+
# Clear sample selection
updateSelectizeInput(session, "sample_select", selected = "")
-
+
+ # Reset app mode to sample mode
+ updateRadioButtons(session, "app_mode", selected = "sample")
+
# Clear any displayed content via JavaScript
shinyjs::runjs("$('.image-card').remove();")
})
@@ -2030,26 +3077,27 @@ server <- function(input, output, session) {
output$image_gallery <- renderUI({
req(paginated_images())
req(rv$temp_png_folder)
- req(rv$current_sample)
-
+ # Allow gallery to render in class review mode (no current_sample)
+ req(isTRUE(rv$class_review_mode) || !is.null(rv$current_sample))
+
p <- paginated_images()
images <- p$images
-
+
if (nrow(images) == 0) {
return(div(class = "alert alert-info", "No images to display"))
}
-
+
classes <- sort(unique(images$class_name))
-
+
class_panels <- lapply(classes, function(cls) {
class_images <- images %>% filter(class_name == cls)
-
+
image_cards <- lapply(seq_len(nrow(class_images)), function(i) {
img_row <- class_images[i, ]
img_file <- img_row$file_name
-
+
is_selected <- img_file %in% rv$selected_images
-
+
was_relabeled <- FALSE
original_class <- ""
orig_idx <- which(rv$original_classifications$file_name == img_file)
@@ -2057,7 +3105,7 @@ server <- function(input, output, session) {
original_class <- rv$original_classifications$class_name[orig_idx]
was_relabeled <- (original_class != img_row$class_name)
}
-
+
border_style <- if (is_selected) {
"border: 3px solid #007bff;"
} else if (was_relabeled) {
@@ -2065,13 +3113,20 @@ server <- function(input, output, session) {
} else {
"border: 1px solid #ddd;"
}
-
+
card_class <- if (is_selected) "image-card selected" else "image-card"
-
+
# Sanitize file names to prevent XSS
safe_img_file <- htmltools::htmlEscape(img_file)
- safe_sample <- htmltools::htmlEscape(rv$current_sample)
resource_path <- if (!is.null(rv$resource_path_name)) rv$resource_path_name else "temp_images"
+
+ # In class review mode, derive sample from file_name since current_sample is NULL
+ if (rv$class_review_mode) {
+ sample_for_img <- sub("_(\\d{5})\\.png$", "", img_file)
+ } else {
+ sample_for_img <- rv$current_sample
+ }
+ safe_sample <- htmltools::htmlEscape(sample_for_img)
img_src <- sprintf("%s/%s/%s", resource_path, safe_sample, safe_img_file)
div(
@@ -2092,8 +3147,12 @@ server <- function(input, output, session) {
"Not found"),
div(
- style = "font-size: 10px; text-align: center; margin-top: 3px;",
- gsub(".*_(\\d+)\\.png$", "ROI \\1", img_file),
+ style = "font-size: 10px; text-align: center; margin-top: 3px; word-break: break-all;",
+ if (isTRUE(rv$class_review_mode)) {
+ sub("\\.png$", "", img_file)
+ } else {
+ gsub(".*_(\\d+)\\.png$", "ROI \\1", img_file)
+ },
if (was_relabeled) {
tags$span(style = "color: #856404;",
paste0(" (was: ", gsub("_\\d+$", "", original_class), ")"))
@@ -2236,7 +3295,380 @@ server <- function(input, output, session) {
observeEvent(input$relabel_quick, {
do_relabel(input$new_class_quick)
})
-
+
+ # ============================================================================
+ # Class Review Mode
+ # ============================================================================
+
+ # Helper: update class review class list based on current filter values
+ update_cr_class_list <- function() {
+ db_path <- get_db_path(config$db_folder)
+ year <- input$cr_year_select
+ month <- input$cr_month_select
+ instrument <- input$cr_instrument_select
+
+ annotator <- input$cr_annotator_select
+
+ classes_df <- list_classes_db(db_path, year = year, month = month,
+ instrument = instrument,
+ annotator = annotator)
+
+ if (nrow(classes_df) == 0) {
+ updateSelectizeInput(session, "class_review_select",
+ choices = c("No classes found" = ""))
+ return()
+ }
+
+ choices <- setNames(
+ classes_df$class_name,
+ sprintf("%s (%d)", classes_df$class_name, classes_df$count)
+ )
+ updateSelectizeInput(session, "class_review_select", choices = choices)
+ }
+
+ # Helper: update month/instrument choices for class review filters
+ update_cr_month_choices <- function() {
+ db_path <- get_db_path(config$db_folder)
+ meta <- list_annotation_metadata_db(db_path)
+
+ year_val <- input$cr_year_select
+
+ month_names <- c("01" = "Jan", "02" = "Feb", "03" = "Mar", "04" = "Apr",
+ "05" = "May", "06" = "Jun", "07" = "Jul", "08" = "Aug",
+ "09" = "Sep", "10" = "Oct", "11" = "Nov", "12" = "Dec")
+
+ if (!is.null(year_val) && year_val != "all") {
+ # Get samples for this year to extract available months/instruments
+ con <- DBI::dbConnect(RSQLite::SQLite(), get_db_path(config$db_folder))
+ on.exit(DBI::dbDisconnect(con), add = TRUE)
+ year_samples <- DBI::dbGetQuery(con,
+ "SELECT DISTINCT sample_name FROM annotations WHERE sample_name LIKE ?",
+ params = list(paste0("D", year_val, "%"))
+ )$sample_name
+
+ months <- sort(unique(substr(year_samples, 6, 7)))
+ month_labels <- month_names[months]
+ updateSelectInput(session, "cr_month_select",
+ choices = c("All" = "all", setNames(months, month_labels)),
+ selected = "all")
+
+ instruments <- sort(unique(sub(".*_", "", year_samples)))
+ current_instrument <- input$cr_instrument_select
+ selected_instrument <- if (!is.null(current_instrument) && current_instrument %in% instruments) {
+ current_instrument
+ } else {
+ "all"
+ }
+ updateSelectInput(session, "cr_instrument_select",
+ choices = c("All" = "all", setNames(instruments, instruments)),
+ selected = selected_instrument)
+ } else {
+ months <- meta$months
+ month_labels <- month_names[months]
+ updateSelectInput(session, "cr_month_select",
+ choices = c("All" = "all", setNames(months, month_labels)),
+ selected = "all")
+
+ instruments <- meta$instruments
+ updateSelectInput(session, "cr_instrument_select",
+ choices = c("All" = "all", setNames(instruments, instruments)),
+ selected = "all")
+ }
+ }
+
+ # When entering class review mode, populate filters and class dropdown
+ observeEvent(input$app_mode, {
+ if (input$app_mode == "class_review") {
+ db_path <- get_db_path(config$db_folder)
+ meta <- list_annotation_metadata_db(db_path)
+
+ month_names <- c("01" = "Jan", "02" = "Feb", "03" = "Mar", "04" = "Apr",
+ "05" = "May", "06" = "Jun", "07" = "Jul", "08" = "Aug",
+ "09" = "Sep", "10" = "Oct", "11" = "Nov", "12" = "Dec")
+
+ updateSelectInput(session, "cr_year_select",
+ choices = c("All" = "all", setNames(meta$years, meta$years)),
+ selected = "all")
+ month_labels <- month_names[meta$months]
+ updateSelectInput(session, "cr_month_select",
+ choices = c("All" = "all", setNames(meta$months, month_labels)),
+ selected = "all")
+ updateSelectInput(session, "cr_instrument_select",
+ choices = c("All" = "all", setNames(meta$instruments, meta$instruments)),
+ selected = "all")
+ updateSelectInput(session, "cr_annotator_select",
+ choices = c("All" = "all", setNames(meta$annotators, meta$annotators)),
+ selected = "all")
+
+ update_cr_class_list()
+ } else {
+ # Leaving class review mode — clear state
+ rv$class_review_mode <- FALSE
+ rv$class_review_class <- NULL
+ rv$class_review_samples <- character()
+ rv$class_review_original <- NULL
+ }
+ }, ignoreInit = TRUE)
+
+ # Cascading filter updates for class review
+ observeEvent(input$cr_year_select, {
+ req(input$app_mode == "class_review")
+ update_cr_month_choices()
+ update_cr_class_list()
+ }, ignoreInit = TRUE)
+
+ observeEvent(input$cr_month_select, {
+ req(input$app_mode == "class_review")
+ update_cr_class_list()
+ }, ignoreInit = TRUE)
+
+ observeEvent(input$cr_instrument_select, {
+ req(input$app_mode == "class_review")
+ update_cr_class_list()
+ }, ignoreInit = TRUE)
+
+ observeEvent(input$cr_annotator_select, {
+ req(input$app_mode == "class_review")
+ update_cr_class_list()
+ }, ignoreInit = TRUE)
+
+ # Class review info output
+ output$class_review_info <- renderUI({
+ if (!rv$class_review_mode || is.null(rv$classifications)) return(NULL)
+
+ n_images <- nrow(rv$classifications)
+ n_samples <- length(rv$class_review_samples)
+ n_changed <- 0L
+
+ if (!is.null(rv$class_review_original)) {
+ n_changed <- sum(rv$classifications$class_name != rv$class_review_original$class_name)
+ }
+
+ div(
+ style = "font-size: 12px; color: #666; margin-bottom: 8px;",
+ sprintf("%d images from %d samples", n_images, n_samples),
+ if (n_changed > 0) tags$span(
+ style = "color: #dc3545; font-weight: bold; margin-left: 5px;",
+ sprintf("(%d changed)", n_changed)
+ )
+ )
+ })
+
+ # Load class for review
+ observeEvent(input$load_class_review, {
+ req(input$class_review_select, input$class_review_select != "")
+
+ class_name <- input$class_review_select
+ db_path <- get_db_path(config$db_folder)
+
+ rv$is_loading <- TRUE
+ disable_nav_buttons()
+ on.exit({
+ rv$is_loading <- FALSE
+ enable_nav_buttons()
+ })
+
+ # Save current sample work if in sample mode
+ if (!rv$class_review_mode && !is.null(rv$current_sample)) {
+ save_to_cache()
+ }
+
+ # Query all annotations for this class (with filters)
+ annotations <- load_class_annotations_db(db_path, class_name,
+ year = input$cr_year_select,
+ month = input$cr_month_select,
+ instrument = input$cr_instrument_select,
+ annotator = input$cr_annotator_select)
+
+ if (is.null(annotations) || nrow(annotations) == 0) {
+ showNotification(paste("No annotations found for class:", class_name),
+ type = "warning")
+ return()
+ }
+
+ # Get unique samples and their ROI paths
+ unique_samples <- unique(annotations$sample_name)
+ current_roi_map <- roi_path_map()
+ is_dashboard <- identical(config$data_source, "dashboard")
+
+ # Create temp folder for extracted PNGs
+ if (!is.null(rv$temp_png_folder) && dir.exists(rv$temp_png_folder)) {
+ if (!is_dashboard || !startsWith(rv$temp_png_folder, get_dashboard_cache_dir())) {
+ unlink(rv$temp_png_folder, recursive = TRUE)
+ }
+ }
+ rv$temp_png_folder <- tempfile(pattern = "ifcb_class_review_")
+ dir.create(rv$temp_png_folder, recursive = TRUE)
+
+ # Extract PNGs per sample
+ missing_samples <- character()
+ extracted_files <- character()
+
+ withProgress(message = paste("Loading", class_name, "images..."),
+ value = 0, {
+
+ if (is_dashboard) {
+ # Dashboard mode: download individual PNGs directly (much faster than zip)
+ parsed <- parse_dashboard_url(config$dashboard_url)
+ all_file_names <- annotations$file_name
+
+ downloaded <- download_dashboard_images_individual(
+ base_url = parsed$base_url,
+ file_names = all_file_names,
+ dest_dir = rv$temp_png_folder,
+ max_retries = config$dashboard_max_retries
+ )
+
+ extracted_files <- downloaded
+ downloaded_samples <- unique(gsub("_\\d+\\.png$", "", downloaded))
+ missing_samples <- setdiff(unique_samples, downloaded_samples)
+ }
+
+ for (idx in seq_along(unique_samples)) {
+ sn <- unique_samples[idx]
+
+ if (is_dashboard) {
+ # Already handled above in batch
+ incProgress(1 / length(unique_samples))
+ next
+ } else {
+ # Local mode: extract from ROI file
+ roi_path <- current_roi_map[[sn]]
+
+ if (is.null(roi_path) || !file.exists(roi_path)) {
+ missing_samples <- c(missing_samples, sn)
+ next
+ }
+
+ sample_rois <- annotations$roi_number[annotations$sample_name == sn]
+
+ tryCatch({
+ ifcb_extract_pngs(
+ roi_file = roi_path,
+ out_folder = rv$temp_png_folder,
+ ROInumbers = sample_rois,
+ verbose = FALSE
+ )
+
+ # Check which files were actually extracted
+ sample_dir <- file.path(rv$temp_png_folder, sn)
+ if (dir.exists(sample_dir)) {
+ files <- list.files(sample_dir, pattern = "\\.png$")
+ extracted_files <- c(extracted_files, files)
+ }
+ }, error = function(e) {
+ missing_samples <<- c(missing_samples, sn)
+ })
+ }
+
+ incProgress(1 / length(unique_samples))
+ }
+ })
+
+ # Filter annotations to only those with extracted images
+ annotations <- annotations[annotations$file_name %in% extracted_files, ]
+
+ if (nrow(annotations) == 0) {
+ showNotification("No images could be extracted. Check ROI file paths.",
+ type = "error")
+ return()
+ }
+
+ # Build classifications data frame (compatible with gallery)
+ classifications <- data.frame(
+ file_name = annotations$file_name,
+ class_name = annotations$class_name,
+ score = NA_real_,
+ width = NA_real_,
+ height = NA_real_,
+ roi_area = NA_real_,
+ stringsAsFactors = FALSE
+ )
+
+ # Set class review state
+ rv$class_review_mode <- TRUE
+ rv$class_review_class <- class_name
+ rv$class_review_samples <- setdiff(unique_samples, missing_samples)
+ rv$current_sample <- NULL
+ rv$classifications <- classifications
+ rv$original_classifications <- classifications
+ rv$class_review_original <- classifications
+ rv$is_annotation_mode <- TRUE
+ rv$has_both_modes <- FALSE
+ rv$selected_images <- character()
+ rv$current_page <- 1
+ rv$changes_log <- create_empty_changes_log()
+
+ # Update class filter dropdown
+ available_classes <- sort(unique(classifications$class_name))
+ updateSelectInput(session, "class_filter",
+ choices = c("All" = "all", setNames(available_classes, available_classes)),
+ selected = "all")
+
+ # Notify
+ n_extracted <- nrow(classifications)
+ n_samples <- length(rv$class_review_samples)
+ msg <- sprintf("Loaded %d %s images from %d samples", n_extracted, class_name, n_samples)
+ if (length(missing_samples) > 0) {
+ msg <- paste0(msg, sprintf(" (%d samples skipped - ROI not found)", length(missing_samples)))
+ }
+ showNotification(msg, type = "message", duration = 8)
+ })
+
+ # Save class review changes
+ observeEvent(input$save_class_review_btn, {
+ req(rv$class_review_mode)
+ req(rv$classifications)
+ req(rv$class_review_original)
+
+ rv$is_loading <- TRUE
+ on.exit({ rv$is_loading <- FALSE })
+
+ # Find changed rows
+ current <- rv$classifications
+ original <- rv$class_review_original
+ changed_mask <- current$class_name != original$class_name
+ n_changed <- sum(changed_mask)
+
+ if (n_changed == 0) {
+ showNotification("No changes to save", type = "warning")
+ return()
+ }
+
+ # Parse sample_name and roi_number from file_name
+ changed_files <- current$file_name[changed_mask]
+ changes_df <- data.frame(
+ sample_name = sub("_(\\d{5})\\.png$", "", changed_files),
+ roi_number = as.integer(sub(".*_(\\d{5})\\.png$", "\\1", changed_files)),
+ new_class_name = current$class_name[changed_mask],
+ stringsAsFactors = FALSE
+ )
+
+ db_path <- get_db_path(config$db_folder)
+ annotator <- if (!is.null(input$annotator_name) && nzchar(input$annotator_name)) {
+ input$annotator_name
+ } else {
+ "Unknown"
+ }
+
+ tryCatch({
+ withProgress(message = "Saving class review changes...", {
+ updated <- save_class_review_changes_db(db_path, changes_df, annotator)
+ })
+
+ # Update original to reflect saved state
+ rv$class_review_original <- rv$classifications
+
+ showNotification(
+ sprintf("Saved %d changes across %d samples",
+ updated, length(unique(changes_df$sample_name))),
+ type = "message"
+ )
+ }, error = function(e) {
+ showNotification(paste("Error saving:", e$message), type = "error")
+ })
+ })
+
# ============================================================================
# Manual Save
# ============================================================================
@@ -2270,14 +3702,44 @@ server <- function(input, output, session) {
}
tryCatch({
+ is_dashboard <- identical(config$data_source, "dashboard")
+
roi_path <- roi_path_map()[[rv$current_sample]]
adc_folder <- if (!is.null(roi_path)) dirname(roi_path) else NULL
- if (is.null(adc_folder)) {
+
+ # In dashboard mode, adc_folder may be NULL since there are no local ROI files
+ if (is.null(adc_folder) && is_dashboard) {
+ # Try to get ADC from dashboard cache for MAT saving
+ cache_dir <- get_dashboard_cache_dir()
+ parsed <- parse_dashboard_url(config$dashboard_url)
+ adc_path <- download_dashboard_adc(parsed$base_url, rv$current_sample, cache_dir,
+ parallel_downloads = config$dashboard_parallel_downloads,
+ sleep_time = config$dashboard_sleep_time,
+ multi_timeout = config$dashboard_multi_timeout,
+ max_retries = config$dashboard_max_retries)
+ adc_folder <- if (!is.null(adc_path)) dirname(adc_path) else NULL
+ }
+
+ save_fmt <- config$save_format
+
+ # In dashboard mode, if MAT save is requested but no ADC available, fall back to SQLite
+ if (is_dashboard && is.null(adc_folder) && save_fmt %in% c("mat", "both")) {
+ if (save_fmt == "mat") {
+ showNotification("MAT saving requires ADC data (not available from dashboard). Saving to SQLite instead.",
+ type = "warning", duration = 8)
+ save_fmt <- "sqlite"
+ } else {
+ showNotification("ADC data not available from dashboard. Skipping MAT save, saving to SQLite only.",
+ type = "warning", duration = 8)
+ save_fmt <- "sqlite"
+ }
+ }
+
+ if (is.null(adc_folder) && !is_dashboard) {
showNotification("Cannot find ROI data folder for this sample", type = "error")
return()
}
- save_fmt <- config$save_format
progress_msg <- switch(save_fmt,
sqlite = "Saving to database...",
mat = "Saving MAT file...",
diff --git a/inst/app/ui.R b/inst/app/ui.R
index 2f1cd4a..87a8acd 100644
--- a/inst/app/ui.R
+++ b/inst/app/ui.R
@@ -284,6 +284,9 @@ ui <- page_sidebar(
.navbar-mode-validation {
background-color: #28a745 !important; /* Green for validation */
}
+ .navbar-mode-class-review {
+ background-color: #6f42c1 !important; /* Purple for class review */
+ }
/* Override navbar title color for visibility */
.navbar .navbar-brand, .navbar .navbar-brand span {
@@ -307,6 +310,11 @@ ui <- page_sidebar(
background-color: #fff3cd !important;
}
+ /* Taller dropdown for class review select */
+ #class_review_select + .selectize-control .selectize-dropdown-content {
+ max-height: 350px !important;
+ }
+
/* Loading overlay styles */
.loading-overlay {
position: fixed;
@@ -373,7 +381,7 @@ ui <- page_sidebar(
),
sidebar = sidebar(
- width = 320,
+ width = 360,
# Annotator and settings at top
div(
@@ -390,70 +398,124 @@ ui <- page_sidebar(
hr(),
- # Sample selection
- h4("Sample Selection"),
+ # Mode toggle: Sample Mode vs Class Review
+ radioButtons("app_mode", NULL,
+ choices = c("Sample Mode" = "sample",
+ "Class Review" = "class_review"),
+ selected = "sample", inline = TRUE),
- # Year and month filters in a row
- div(
- style = "display: flex; gap: 10px;",
- div(style = "flex: 1;",
- selectInput("year_select", "Year", choices = NULL, width = "100%")),
- div(style = "flex: 1;",
- selectInput("month_select", "Month", choices = c("All" = "all"), width = "100%"))
- ),
+ # ── Sample Mode panel ──
+ conditionalPanel(
+ condition = "input.app_mode == 'sample'",
- selectInput("sample_status_filter", "Show",
- choices = c("All samples" = "all",
- "Auto-classified (validation)" = "classified",
- "Manually annotated" = "annotated",
- "Unannotated" = "unclassified")),
-
- # Sample dropdown with CSS to prevent text wrapping and reduce spacing
- tags$style("
- .sample-dropdown .selectize-input { white-space: nowrap; overflow: hidden; text-overflow: ellipsis; }
- .sample-dropdown .form-group { margin-bottom: 2px; }
- "),
- div(class = "sample-dropdown",
- selectizeInput("sample_select", "Sample", choices = NULL, width = "100%",
- options = list(
- placeholder = "Select sample..."
- ))),
-
- # Legend for sample status symbols (compact, single line)
- div(
- style = "font-size: 12px; color: #666; margin-bottom: 8px; white-space: nowrap;",
- tags$span(style = "margin-right: 8px;", "\u270E Manual"),
- tags$span(style = "margin-right: 8px;", "\u2713 Classified"),
- tags$span("* Unannotated")
- ),
+ h4("Sample Selection"),
- # Navigation buttons
- div(
- style = "display: flex; gap: 5px; margin-bottom: 5px;",
- actionButton("load_sample", "Load",
- class = "btn-primary", style = "flex: 1;"),
- actionButton("prev_sample", label = icon("arrow-left"),
- class = "btn-outline-primary", style = "flex: 0;",
- title = "Previous sample"),
- actionButton("next_sample", label = icon("arrow-right"),
- class = "btn-outline-primary", style = "flex: 0;",
- title = "Next sample"),
- actionButton("random_sample", label = icon("random"),
- class = "btn-outline-secondary", style = "flex: 0;",
- title = "Random sample"),
- actionButton("rescan_folders", label = icon("sync"),
- class = "btn-outline-secondary", style = "flex: 0;",
- title = "Sync folders (refresh file index)")
+ # Year, month, and instrument filters in a row
+ div(
+ style = "display: flex; gap: 10px;",
+ div(style = "flex: 1;",
+ selectInput("year_select", "Year", choices = NULL, width = "100%")),
+ div(style = "flex: 1;",
+ selectInput("month_select", "Month", choices = c("All" = "all"), width = "100%")),
+ div(style = "flex: 1;",
+ selectInput("instrument_select", "IFCB", choices = c("All" = "all"), width = "100%"))
+ ),
+
+ selectInput("sample_status_filter", "Show",
+ choices = c("All samples" = "all",
+ "Auto-classified (validation)" = "classified",
+ "Manually annotated" = "annotated",
+ "Unannotated" = "unclassified")),
+
+ # Sample dropdown with CSS to prevent text wrapping and reduce spacing
+ tags$style("
+ .sample-dropdown .selectize-input { white-space: nowrap; overflow: hidden; text-overflow: ellipsis; }
+ .sample-dropdown .form-group { margin-bottom: 2px; }
+ "),
+ div(class = "sample-dropdown",
+ selectizeInput("sample_select", "Sample", choices = NULL, width = "100%",
+ options = list(
+ placeholder = "Select sample..."
+ ))),
+
+ # Legend for sample status symbols (compact, single line)
+ div(
+ style = "font-size: 12px; color: #666; margin-bottom: 8px; white-space: nowrap;",
+ tags$span(style = "margin-right: 8px;", "\u270E Manual"),
+ tags$span(style = "margin-right: 8px;", "\u2713 Classified"),
+ tags$span("* Unannotated")
+ ),
+
+ # Navigation buttons
+ div(
+ style = "display: flex; gap: 5px; margin-bottom: 5px;",
+ actionButton("load_sample", "Load",
+ class = "btn-primary", style = "flex: 1;"),
+ actionButton("prev_sample", label = icon("arrow-left"),
+ class = "btn-outline-primary", style = "flex: 0;",
+ title = "Previous sample"),
+ actionButton("next_sample", label = icon("arrow-right"),
+ class = "btn-outline-primary", style = "flex: 0;",
+ title = "Next sample"),
+ actionButton("random_sample", label = icon("random"),
+ class = "btn-outline-secondary", style = "flex: 0;",
+ title = "Random sample"),
+ actionButton("rescan_folders", label = icon("sync"),
+ class = "btn-outline-secondary", style = "flex: 0;",
+ title = "Sync folders (refresh file index)")
+ ),
+
+ # Cache age indicator
+ uiOutput("cache_age_text")
),
- # Cache age indicator
- uiOutput("cache_age_text"),
+ # ── Class Review panel ──
+ conditionalPanel(
+ condition = "input.app_mode == 'class_review'",
+
+ h4("Class Review"),
+
+ div(
+ style = "display: flex; gap: 10px; margin-bottom: 10px;",
+ div(style = "flex: 1;",
+ selectInput("cr_year_select", "Year",
+ choices = c("All" = "all"), width = "100%")),
+ div(style = "flex: 1;",
+ selectInput("cr_month_select", "Month",
+ choices = c("All" = "all"), width = "100%")),
+ div(style = "flex: 1;",
+ selectInput("cr_instrument_select", "IFCB",
+ choices = c("All" = "all"), width = "100%"))
+ ),
+
+ selectInput("cr_annotator_select", "Annotator",
+ choices = c("All" = "all"), width = "100%"),
+
+ selectizeInput("class_review_select", "Select Class",
+ choices = NULL, width = "100%",
+ options = list(maxOptions = 1000)),
+
+ uiOutput("class_review_info"),
+
+ div(
+ style = "display: flex; gap: 5px; margin-bottom: 5px;",
+ actionButton("load_class_review", "Load",
+ class = "btn-primary", style = "flex: 1;"),
+ actionButton("save_class_review_btn", "Save Changes",
+ class = "btn-success", style = "flex: 1;")
+ )
+ ),
hr(),
- # Save button (prominent)
- actionButton("save_btn", "Save Annotations",
- class = "btn-success", width = "100%"),
+ # Save and Predict buttons — sample mode only
+ conditionalPanel(
+ condition = "input.app_mode == 'sample'",
+ actionButton("save_btn", "Save Annotations",
+ class = "btn-success", width = "100%"),
+ div(style = "margin-top: 5px;",
+ uiOutput("predict_btn_ui"))
+ ),
uiOutput("python_warning"),
diff --git a/man/download_dashboard_adc.Rd b/man/download_dashboard_adc.Rd
new file mode 100644
index 0000000..20952c8
--- /dev/null
+++ b/man/download_dashboard_adc.Rd
@@ -0,0 +1,37 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{download_dashboard_adc}
+\alias{download_dashboard_adc}
+\title{Download ADC file from the Dashboard}
+\usage{
+download_dashboard_adc(
+ base_url,
+ sample_name,
+ cache_dir = get_dashboard_cache_dir(),
+ parallel_downloads = 5,
+ sleep_time = 2,
+ multi_timeout = 120,
+ max_retries = 3
+)
+}
+\arguments{
+\item{base_url}{Character. Dashboard base URL.}
+
+\item{sample_name}{Character. Sample name.}
+
+\item{cache_dir}{Character. Cache directory.}
+
+\item{parallel_downloads}{Integer. Number of parallel downloads.}
+
+\item{sleep_time}{Numeric. Seconds to sleep between download batches.}
+
+\item{multi_timeout}{Numeric. Timeout in seconds for multi-file downloads.}
+
+\item{max_retries}{Integer. Maximum number of retry attempts.}
+}
+\value{
+Path to the downloaded ADC file, or NULL on failure.
+}
+\description{
+Downloads the ADC file for a sample from the Dashboard on demand.
+}
diff --git a/man/download_dashboard_autoclass.Rd b/man/download_dashboard_autoclass.Rd
new file mode 100644
index 0000000..ac48bc5
--- /dev/null
+++ b/man/download_dashboard_autoclass.Rd
@@ -0,0 +1,39 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{download_dashboard_autoclass}
+\alias{download_dashboard_autoclass}
+\title{Download and parse autoclass scores from the Dashboard}
+\usage{
+download_dashboard_autoclass(
+ base_url,
+ sample_name,
+ cache_dir = get_dashboard_cache_dir(),
+ parallel_downloads = 5,
+ sleep_time = 2,
+ multi_timeout = 120,
+ max_retries = 3
+)
+}
+\arguments{
+\item{base_url}{Character. Dashboard base URL.}
+
+\item{sample_name}{Character. Sample name.}
+
+\item{cache_dir}{Character. Cache directory.}
+
+\item{parallel_downloads}{Integer. Number of parallel downloads.}
+
+\item{sleep_time}{Numeric. Seconds to sleep between download batches.}
+
+\item{multi_timeout}{Numeric. Timeout in seconds for multi-file downloads.}
+
+\item{max_retries}{Integer. Maximum number of retry attempts.}
+}
+\value{
+Data frame with columns \code{file_name}, \code{class_name},
+ \code{score}, or NULL on failure.
+}
+\description{
+Downloads \code{_class_scores.csv} for a sample and extracts the winning
+class (column with max score) per ROI.
+}
diff --git a/man/download_dashboard_image_single.Rd b/man/download_dashboard_image_single.Rd
new file mode 100644
index 0000000..76d5074
--- /dev/null
+++ b/man/download_dashboard_image_single.Rd
@@ -0,0 +1,35 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{download_dashboard_image_single}
+\alias{download_dashboard_image_single}
+\title{Download a single PNG image from the Dashboard}
+\usage{
+download_dashboard_image_single(
+ base_url,
+ sample_name,
+ roi_number,
+ dest_dir,
+ max_retries = 3,
+ timeout = 15
+)
+}
+\arguments{
+\item{base_url}{Character. Dashboard base URL.}
+
+\item{sample_name}{Character. Sample name (bin PID).}
+
+\item{roi_number}{Integer. ROI number to download.}
+
+\item{dest_dir}{Character. Destination directory.}
+
+\item{max_retries}{Integer. Maximum number of retry attempts.}
+
+\item{timeout}{Numeric. Request timeout in seconds.}
+}
+\value{
+File path to the downloaded PNG, or NULL on failure.
+}
+\description{
+Downloads one PNG from the Dashboard's \code{/data/} endpoint.
+The image is saved to \code{dest_dir/sample_name/file_name}.
+}
diff --git a/man/download_dashboard_images.Rd b/man/download_dashboard_images.Rd
new file mode 100644
index 0000000..a7b5650
--- /dev/null
+++ b/man/download_dashboard_images.Rd
@@ -0,0 +1,39 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{download_dashboard_images}
+\alias{download_dashboard_images}
+\title{Download and extract PNG images from the Dashboard}
+\usage{
+download_dashboard_images(
+ base_url,
+ sample_name,
+ cache_dir = get_dashboard_cache_dir(),
+ parallel_downloads = 5,
+ sleep_time = 2,
+ multi_timeout = 120,
+ max_retries = 3
+)
+}
+\arguments{
+\item{base_url}{Character. Dashboard base URL.}
+
+\item{sample_name}{Character. Sample name (bin PID).}
+
+\item{cache_dir}{Character. Cache directory. Defaults to
+\code{\link{get_dashboard_cache_dir}()}.}
+
+\item{parallel_downloads}{Integer. Number of parallel downloads.}
+
+\item{sleep_time}{Numeric. Seconds to sleep between download batches.}
+
+\item{multi_timeout}{Numeric. Timeout in seconds for multi-file downloads.}
+
+\item{max_retries}{Integer. Maximum number of retry attempts.}
+}
+\value{
+Path to the folder containing extracted PNGs, or NULL on failure.
+}
+\description{
+Downloads a zip file of PNG images for a sample from the Dashboard.
+Extracts into the cache directory. Skips re-download if PNGs already exist.
+}
diff --git a/man/download_dashboard_images_bulk.Rd b/man/download_dashboard_images_bulk.Rd
new file mode 100644
index 0000000..8e794ae
--- /dev/null
+++ b/man/download_dashboard_images_bulk.Rd
@@ -0,0 +1,41 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{download_dashboard_images_bulk}
+\alias{download_dashboard_images_bulk}
+\title{Bulk download zip archives for multiple samples from the Dashboard}
+\usage{
+download_dashboard_images_bulk(
+ base_url,
+ sample_names,
+ cache_dir = get_dashboard_cache_dir(),
+ parallel_downloads = 5,
+ sleep_time = 2,
+ multi_timeout = 120,
+ max_retries = 3
+)
+}
+\arguments{
+\item{base_url}{Character. Dashboard base URL.}
+
+\item{sample_names}{Character vector. Sample names to download.}
+
+\item{cache_dir}{Character. Cache directory.}
+
+\item{parallel_downloads}{Integer. Number of parallel downloads.}
+
+\item{sleep_time}{Numeric. Seconds to sleep between download batches.}
+
+\item{multi_timeout}{Numeric. Timeout in seconds for multi-file downloads.}
+
+\item{max_retries}{Integer. Maximum number of retry attempts.}
+}
+\value{
+Character vector of sample names that were successfully downloaded
+ or already cached.
+}
+\description{
+Downloads zip files for all specified samples in a single batched call
+to \code{\link[iRfcb]{ifcb_download_dashboard_data}}, leveraging its
+built-in parallel download support. Samples already cached are skipped.
+After download, zips are extracted and cleaned up.
+}
diff --git a/man/download_dashboard_images_individual.Rd b/man/download_dashboard_images_individual.Rd
new file mode 100644
index 0000000..59bbf06
--- /dev/null
+++ b/man/download_dashboard_images_individual.Rd
@@ -0,0 +1,39 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{download_dashboard_images_individual}
+\alias{download_dashboard_images_individual}
+\title{Download individual PNG images from the Dashboard}
+\usage{
+download_dashboard_images_individual(
+ base_url,
+ file_names,
+ dest_dir,
+ max_retries = 3,
+ sample_fail_threshold = 2
+)
+}
+\arguments{
+\item{base_url}{Character. Dashboard base URL.}
+
+\item{file_names}{Character vector. PNG file names
+(e.g., \code{"D20240716T000431_IFCB134_00108.png"}).}
+
+\item{dest_dir}{Character. Destination directory.}
+
+\item{max_retries}{Integer. Maximum number of retry attempts per image.}
+
+\item{sample_fail_threshold}{Integer. After this many consecutive failures
+from the same sample, skip all remaining images from that sample.}
+}
+\value{
+Character vector of successfully downloaded file names.
+}
+\description{
+Downloads specific PNG files from the Dashboard's \code{/data/} endpoint,
+one at a time. This is much faster than downloading entire zip archives
+when only a subset of ROIs are needed (e.g., class review mode).
+}
+\details{
+Samples that fail repeatedly are automatically skipped to avoid long
+waits when annotations reference samples not available on the dashboard.
+}
diff --git a/man/figures/class-editor.png b/man/figures/class-editor.png
index 2c2c5cd..a4b904c 100644
Binary files a/man/figures/class-editor.png and b/man/figures/class-editor.png differ
diff --git a/man/figures/class-review.png b/man/figures/class-review.png
new file mode 100644
index 0000000..69ce36e
Binary files /dev/null and b/man/figures/class-review.png differ
diff --git a/man/figures/interface-overview.png b/man/figures/interface-overview.png
index f2568a0..ce9732b 100644
Binary files a/man/figures/interface-overview.png and b/man/figures/interface-overview.png differ
diff --git a/man/figures/sample-browser.png b/man/figures/sample-browser.png
index 9e0def3..9ba1b9d 100644
Binary files a/man/figures/sample-browser.png and b/man/figures/sample-browser.png differ
diff --git a/man/figures/settings-dialog.png b/man/figures/settings-dialog.png
index 464830b..e9290c6 100644
Binary files a/man/figures/settings-dialog.png and b/man/figures/settings-dialog.png differ
diff --git a/man/get_dashboard_cache_dir.Rd b/man/get_dashboard_cache_dir.Rd
new file mode 100644
index 0000000..006957e
--- /dev/null
+++ b/man/get_dashboard_cache_dir.Rd
@@ -0,0 +1,19 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{get_dashboard_cache_dir}
+\alias{get_dashboard_cache_dir}
+\title{Get persistent cache directory for dashboard downloads}
+\usage{
+get_dashboard_cache_dir()
+}
+\value{
+Path to the dashboard cache directory
+}
+\description{
+Returns the path to the dashboard cache directory. During R CMD check,
+uses a temporary directory.
+}
+\examples{
+cache_dir <- get_dashboard_cache_dir()
+print(cache_dir)
+}
diff --git a/man/import_png_folder_to_db.Rd b/man/import_png_folder_to_db.Rd
new file mode 100644
index 0000000..2471e96
--- /dev/null
+++ b/man/import_png_folder_to_db.Rd
@@ -0,0 +1,49 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/database.R
+\name{import_png_folder_to_db}
+\alias{import_png_folder_to_db}
+\title{Import annotations from a PNG class folder into the SQLite database}
+\usage{
+import_png_folder_to_db(
+ png_folder,
+ db_path,
+ class2use,
+ class_mapping = NULL,
+ annotator = "imported"
+)
+}
+\arguments{
+\item{png_folder}{Path to the top-level folder containing class subfolders}
+
+\item{db_path}{Path to the SQLite database file}
+
+\item{class2use}{Character vector of class names (preserves index order for
+.mat export)}
+
+\item{class_mapping}{Optional named character vector mapping scanned class
+names to target class names. Names are the source classes, values are the
+target classes. Classes not in the mapping are kept as-is.}
+
+\item{annotator}{Annotator name (defaults to \code{"imported"})}
+}
+\value{
+Named list with counts: \code{success}, \code{failed}
+}
+\description{
+Scans a folder of PNG images organized in class-name subfolders (via
+\code{\link{scan_png_class_folder}}) and imports the annotations into the
+database. An optional \code{class_mapping} named vector remaps class names
+before saving.
+}
+\examples{
+\dontrun{
+db_path <- get_db_path("/data/manual")
+class2use <- c("Diatom", "Dinoflagellate", "Ciliate")
+result <- import_png_folder_to_db(
+ "/data/png_export", db_path, class2use,
+ class_mapping = c("OldName" = "NewName"),
+ annotator = "Jane"
+)
+cat(result$success, "imported,", result$failed, "failed\n")
+}
+}
diff --git a/man/list_annotation_metadata_db.Rd b/man/list_annotation_metadata_db.Rd
new file mode 100644
index 0000000..3830707
--- /dev/null
+++ b/man/list_annotation_metadata_db.Rd
@@ -0,0 +1,31 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/database.R
+\name{list_annotation_metadata_db}
+\alias{list_annotation_metadata_db}
+\title{List distinct years, months, and instruments from annotations}
+\usage{
+list_annotation_metadata_db(db_path)
+}
+\arguments{
+\item{db_path}{Path to the SQLite database file}
+}
+\value{
+A list with character vectors: \code{years}, \code{months},
+ \code{instruments}, and \code{annotators}. Returns empty vectors if the
+ database does not exist or has no annotations.
+}
+\description{
+Extracts metadata from sample names in the annotations table for use as
+filter options. Sample names follow the IFCB naming convention
+\code{DYYYYMMDDTHHMMSS_INSTRUMENT}.
+}
+\examples{
+\dontrun{
+db_path <- get_db_path("/data/manual")
+meta <- list_annotation_metadata_db(db_path)
+meta$years # e.g. c("2022", "2023")
+meta$months # e.g. c("01", "06", "12")
+meta$instruments # e.g. c("IFCB134", "IFCB135")
+meta$annotators # e.g. c("Jane", "imported")
+}
+}
diff --git a/man/list_classes_db.Rd b/man/list_classes_db.Rd
new file mode 100644
index 0000000..36a5ab7
--- /dev/null
+++ b/man/list_classes_db.Rd
@@ -0,0 +1,47 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/database.R
+\name{list_classes_db}
+\alias{list_classes_db}
+\title{List all classes with counts in the annotations database}
+\usage{
+list_classes_db(
+ db_path,
+ year = NULL,
+ month = NULL,
+ instrument = NULL,
+ annotator = NULL
+)
+}
+\arguments{
+\item{db_path}{Path to the SQLite database file}
+
+\item{year}{Optional year filter (e.g. \code{"2023"}). When not \code{"all"}
+or \code{NULL}, restricts to sample names starting with \code{DYYYY}.}
+
+\item{month}{Optional month filter (e.g. \code{"03"}). When not \code{"all"}
+or \code{NULL}, restricts to sample names with that month at positions 6-7.}
+
+\item{instrument}{Optional instrument filter (e.g. \code{"IFCB134"}). When
+not \code{"all"} or \code{NULL}, restricts to sample names ending with
+\code{_INSTRUMENT}.}
+
+\item{annotator}{Optional annotator name filter (e.g. \code{"Jane"}). When
+not \code{"all"} or \code{NULL}, restricts to annotations by that annotator.}
+}
+\value{
+Data frame with columns \code{class_name} and \code{count}, ordered
+ alphabetically by class name. Returns an empty data frame if the database
+ does not exist or has no annotations.
+}
+\description{
+Queries the database for distinct class names and their annotation counts.
+Useful for populating class review mode dropdowns. Optional filters restrict
+results to annotations matching a given year, month, or instrument.
+}
+\examples{
+\dontrun{
+db_path <- get_db_path("/data/manual")
+classes <- list_classes_db(db_path)
+classes_2023 <- list_classes_db(db_path, year = "2023")
+}
+}
diff --git a/man/list_dashboard_bins.Rd b/man/list_dashboard_bins.Rd
new file mode 100644
index 0000000..a000907
--- /dev/null
+++ b/man/list_dashboard_bins.Rd
@@ -0,0 +1,26 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{list_dashboard_bins}
+\alias{list_dashboard_bins}
+\title{List bins from an IFCB Dashboard}
+\usage{
+list_dashboard_bins(base_url, dataset_name = NULL)
+}
+\arguments{
+\item{base_url}{Character. Base URL (e.g. \code{"https://habon-ifcb.whoi.edu"}).}
+
+\item{dataset_name}{Optional character. Dataset slug (e.g. \code{"tangosund"}).}
+}
+\value{
+Character vector of bin (sample) names.
+}
+\description{
+Fetches the bin list from the Dashboard API. This is a vendored copy of
+\code{iRfcb::ifcb_list_dashboard_bins()} from the development version that
+supports the \code{dataset_name} parameter.
+}
+\examples{
+\donttest{
+ bins <- list_dashboard_bins("https://ifcb-data.whoi.edu", "mvco")
+}
+}
diff --git a/man/load_class_annotations_db.Rd b/man/load_class_annotations_db.Rd
new file mode 100644
index 0000000..9b420cc
--- /dev/null
+++ b/man/load_class_annotations_db.Rd
@@ -0,0 +1,45 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/database.R
+\name{load_class_annotations_db}
+\alias{load_class_annotations_db}
+\title{Load all annotations for a specific class from the database}
+\usage{
+load_class_annotations_db(
+ db_path,
+ class_name,
+ year = NULL,
+ month = NULL,
+ instrument = NULL,
+ annotator = NULL
+)
+}
+\arguments{
+\item{db_path}{Path to the SQLite database file}
+
+\item{class_name}{Class name to load}
+
+\item{year}{Optional year filter (e.g. \code{"2023"})}
+
+\item{month}{Optional month filter (e.g. \code{"03"})}
+
+\item{instrument}{Optional instrument filter (e.g. \code{"IFCB134"})}
+
+\item{annotator}{Optional annotator name filter (e.g. \code{"Jane"})}
+}
+\value{
+Data frame with columns \code{sample_name}, \code{roi_number},
+ \code{class_name}, and \code{file_name}. Returns \code{NULL} if no
+ annotations match.
+}
+\description{
+Returns every annotation matching \code{class_name}, with a computed
+\code{file_name} column for gallery display. Optional filters restrict
+results by year, month, or instrument.
+}
+\examples{
+\dontrun{
+db_path <- get_db_path("/data/manual")
+diatoms <- load_class_annotations_db(db_path, "Diatom")
+diatoms_2023 <- load_class_annotations_db(db_path, "Diatom", year = "2023")
+}
+}
diff --git a/man/load_from_csv.Rd b/man/load_from_csv.Rd
index 774dadd..96475f6 100644
--- a/man/load_from_csv.Rd
+++ b/man/load_from_csv.Rd
@@ -4,10 +4,14 @@
\alias{load_from_csv}
\title{Load classifications from CSV file (validation mode)}
\usage{
-load_from_csv(csv_path)
+load_from_csv(csv_path, use_threshold = TRUE)
}
\arguments{
\item{csv_path}{Path to classification CSV file}
+
+\item{use_threshold}{Logical, whether to use the threshold-filtered
+\code{class_name} column (default \code{TRUE}) or the raw
+\code{class_name_auto} column when available.}
}
\value{
Data frame with classifications. Expected columns: `file_name`,
@@ -25,9 +29,12 @@ The CSV file must contain the following columns:
\item{class_name}{Predicted class name (e.g., `Diatom`).}
}
-An optional column may also be included:
+Optional columns may also be included:
\describe{
\item{score}{Classification confidence value between 0 and 1.}
+ \item{class_name_auto}{Raw (unthresholded) class prediction. When
+ \code{use_threshold = FALSE} and this column exists, its values are
+ used as \code{class_name}.}
}
The CSV file must be named after the sample it describes
diff --git a/man/load_from_h5.Rd b/man/load_from_h5.Rd
new file mode 100644
index 0000000..23a0751
--- /dev/null
+++ b/man/load_from_h5.Rd
@@ -0,0 +1,39 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/sample_loading.R
+\name{load_from_h5}
+\alias{load_from_h5}
+\title{Load classifications from HDF5 classifier output file}
+\usage{
+load_from_h5(h5_path, sample_name, roi_dimensions, use_threshold = TRUE)
+}
+\arguments{
+\item{h5_path}{Path to classifier H5 file (matching pattern *_class*.h5)}
+
+\item{sample_name}{Sample name (e.g., "D20220522T000439_IFCB134")}
+
+\item{roi_dimensions}{Data frame from \code{\link{read_roi_dimensions}}}
+
+\item{use_threshold}{Logical, whether to use the threshold-filtered
+\code{class_name} dataset (default \code{TRUE}) or the raw
+\code{class_name_auto} dataset.}
+}
+\value{
+Data frame with columns: file_name, class_name, score, width, height,
+ roi_area
+}
+\description{
+Reads an HDF5 classifier output file (from iRfcb 0.8.0+) and extracts
+class predictions. Requires the \pkg{hdf5r} package.
+}
+\examples{
+\dontrun{
+dims <- read_roi_dimensions("/data/raw/2022/D20220522/D20220522T000439_IFCB134.adc")
+classifications <- load_from_h5(
+ h5_path = "/data/classified/D20220522T000439_IFCB134_class.h5",
+ sample_name = "D20220522T000439_IFCB134",
+ roi_dimensions = dims,
+ use_threshold = TRUE
+)
+head(classifications)
+}
+}
diff --git a/man/parse_dashboard_url.Rd b/man/parse_dashboard_url.Rd
new file mode 100644
index 0000000..bdcdfed
--- /dev/null
+++ b/man/parse_dashboard_url.Rd
@@ -0,0 +1,24 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/dashboard.R
+\name{parse_dashboard_url}
+\alias{parse_dashboard_url}
+\title{Parse an IFCB Dashboard URL}
+\usage{
+parse_dashboard_url(url)
+}
+\arguments{
+\item{url}{Character. A Dashboard URL, e.g.
+\code{"https://habon-ifcb.whoi.edu/"} or
+\code{"https://habon-ifcb.whoi.edu/timeline?dataset=tangosund"}.}
+}
+\value{
+A list with \code{base_url} (without trailing slash) and
+ \code{dataset_name} (character or NULL).
+}
+\description{
+Extracts the base URL and optional dataset name from a Dashboard URL.
+}
+\examples{
+parse_dashboard_url("https://habon-ifcb.whoi.edu/")
+parse_dashboard_url("https://habon-ifcb.whoi.edu/timeline?dataset=tangosund")
+}
diff --git a/man/rescan_file_index.Rd b/man/rescan_file_index.Rd
index 30876d0..fb54b64 100644
--- a/man/rescan_file_index.Rd
+++ b/man/rescan_file_index.Rd
@@ -9,13 +9,15 @@ rescan_file_index(
csv_folder = NULL,
output_folder = NULL,
verbose = TRUE,
- db_folder = NULL
+ db_folder = NULL,
+ data_source = "local",
+ dashboard_url = NULL
)
}
\arguments{
\item{roi_folder}{Path to ROI data folder. If NULL, read from saved settings.}
-\item{csv_folder}{Path to classification folder (CSV/MAT). If NULL, read from saved settings.}
+\item{csv_folder}{Path to classification folder (CSV/H5/MAT). If NULL, read from saved settings.}
\item{output_folder}{Path to output folder for MAT annotations. If NULL, read from saved settings.}
@@ -24,6 +26,12 @@ rescan_file_index(
\item{db_folder}{Path to the database folder for SQLite annotations. If NULL,
read from saved settings; if not found in settings, defaults to
\code{\link{get_default_db_dir}()}.}
+
+\item{data_source}{Either \code{"local"} (default) for local folder scanning,
+or \code{"dashboard"} to fetch the sample list from a remote IFCB Dashboard.}
+
+\item{dashboard_url}{When \code{data_source = "dashboard"}, the full Dashboard
+URL (e.g. \code{"https://habon-ifcb.whoi.edu/timeline?dataset=tangosund"}).}
}
\value{
Invisibly returns the file index list, or NULL if roi_folder is invalid.
@@ -49,6 +57,10 @@ rescan_file_index(
output_folder = "/data/ifcb/manual"
)
+# Scan from a remote Dashboard
+rescan_file_index(data_source = "dashboard",
+ dashboard_url = "https://habon-ifcb.whoi.edu/timeline?dataset=tangosund")
+
# Use in a cron job:
# Rscript -e 'ClassiPyR::rescan_file_index()'
}
diff --git a/man/save_class_review_changes_db.Rd b/man/save_class_review_changes_db.Rd
new file mode 100644
index 0000000..4f75b97
--- /dev/null
+++ b/man/save_class_review_changes_db.Rd
@@ -0,0 +1,35 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/database.R
+\name{save_class_review_changes_db}
+\alias{save_class_review_changes_db}
+\title{Save class review changes to the database}
+\usage{
+save_class_review_changes_db(db_path, changes_df, annotator)
+}
+\arguments{
+\item{db_path}{Path to the SQLite database file}
+
+\item{changes_df}{Data frame with columns \code{sample_name},
+\code{roi_number}, and \code{new_class_name}}
+
+\item{annotator}{Annotator name}
+}
+\value{
+Integer count of rows updated
+}
+\description{
+Performs row-level UPDATEs for reclassified images identified during class
+review mode. Only the changed rows are updated; other annotations for the
+same samples are left untouched.
+}
+\examples{
+\dontrun{
+db_path <- get_db_path("/data/manual")
+changes <- data.frame(
+ sample_name = "D20230101T120000_IFCB134",
+ roi_number = 5L,
+ new_class_name = "Ciliate"
+)
+save_class_review_changes_db(db_path, changes, "Jane")
+}
+}
diff --git a/man/scan_png_class_folder.Rd b/man/scan_png_class_folder.Rd
new file mode 100644
index 0000000..e942fa9
--- /dev/null
+++ b/man/scan_png_class_folder.Rd
@@ -0,0 +1,34 @@
+% Generated by roxygen2: do not edit by hand
+% Please edit documentation in R/sample_loading.R
+\name{scan_png_class_folder}
+\alias{scan_png_class_folder}
+\title{Scan a PNG folder with class subfolders}
+\usage{
+scan_png_class_folder(png_folder)
+}
+\arguments{
+\item{png_folder}{Path to the top-level folder containing class subfolders}
+}
+\value{
+A list with components:
+ \describe{
+ \item{annotations}{Data frame with columns \code{sample_name},
+ \code{roi_number}, \code{file_name}, and \code{class_name}}
+ \item{classes_found}{Character vector of unique class names found}
+ \item{sample_names}{Character vector of unique sample names found}
+ }
+}
+\description{
+Scans a directory containing PNG images organized into class-name
+subfolders (e.g. as exported by \code{\link{export_db_to_png}} or other
+tools). Folder names follow the iRfcb convention where a trailing 3-digit
+suffix is stripped (e.g. \code{Diatom_001} becomes \code{Diatom}).
+}
+\examples{
+\dontrun{
+result <- scan_png_class_folder("/data/png_export")
+head(result$annotations)
+result$classes_found
+result$sample_names
+}
+}
diff --git a/tests/testthat/setup.R b/tests/testthat/setup.R
index 45e1beb..2a133fa 100644
--- a/tests/testthat/setup.R
+++ b/tests/testthat/setup.R
@@ -8,20 +8,26 @@ on_cran <- !identical(Sys.getenv("NOT_CRAN"), "true") &&
if (!on_cran) {
# Try to initialize Python for tests
- # First check if Python is already available
if (!reticulate::py_available(initialize = FALSE)) {
- # Try to discover and use system Python
- python_config <- tryCatch(
- reticulate::py_discover_config(),
- error = function(e) NULL
- )
-
- if (!is.null(python_config) && !is.null(python_config$python)) {
- tryCatch({
- reticulate::use_python(python_config$python, required = FALSE)
- }, error = function(e) {
- message("Could not configure Python: ", e$message)
- })
+ # Prefer the r-reticulate virtualenv (where py_install puts packages)
+ venv_path <- "~/.virtualenvs/r-reticulate"
+ if (dir.exists(path.expand(venv_path))) {
+ tryCatch(
+ reticulate::use_virtualenv(venv_path, required = FALSE),
+ error = function(e) message("Could not use virtualenv: ", e$message)
+ )
+ } else {
+ # Fall back to system Python discovery
+ python_config <- tryCatch(
+ reticulate::py_discover_config(),
+ error = function(e) NULL
+ )
+ if (!is.null(python_config) && !is.null(python_config$python)) {
+ tryCatch(
+ reticulate::use_python(python_config$python, required = FALSE),
+ error = function(e) message("Could not configure Python: ", e$message)
+ )
+ }
}
}
diff --git a/tests/testthat/test-dashboard.R b/tests/testthat/test-dashboard.R
new file mode 100644
index 0000000..eff9cd0
--- /dev/null
+++ b/tests/testthat/test-dashboard.R
@@ -0,0 +1,700 @@
+# Tests for dashboard helper functions
+
+# ---------------------------------------------------------------------------
+# Unit tests (always run)
+# ---------------------------------------------------------------------------
+
+test_that("parse_dashboard_url extracts base URL from simple URL", {
+ result <- parse_dashboard_url("https://habon-ifcb.whoi.edu/")
+ expect_equal(result$base_url, "https://habon-ifcb.whoi.edu")
+ expect_null(result$dataset_name)
+})
+
+test_that("parse_dashboard_url extracts base URL without trailing slash", {
+ result <- parse_dashboard_url("https://habon-ifcb.whoi.edu")
+ expect_equal(result$base_url, "https://habon-ifcb.whoi.edu")
+ expect_null(result$dataset_name)
+})
+
+test_that("parse_dashboard_url extracts dataset from query parameter", {
+ result <- parse_dashboard_url("https://habon-ifcb.whoi.edu/timeline?dataset=tangosund")
+ expect_equal(result$base_url, "https://habon-ifcb.whoi.edu")
+ expect_equal(result$dataset_name, "tangosund")
+})
+
+test_that("parse_dashboard_url handles dataset with trailing slash", {
+ result <- parse_dashboard_url("https://habon-ifcb.whoi.edu/timeline/?dataset=tangosund")
+ expect_equal(result$base_url, "https://habon-ifcb.whoi.edu")
+ expect_equal(result$dataset_name, "tangosund")
+})
+
+test_that("parse_dashboard_url handles URL without dataset parameter", {
+ result <- parse_dashboard_url("https://ifcb-data.whoi.edu/timeline")
+ expect_equal(result$base_url, "https://ifcb-data.whoi.edu")
+ expect_null(result$dataset_name)
+})
+
+test_that("parse_dashboard_url handles URL with multiple query params", {
+ result <- parse_dashboard_url("https://habon-ifcb.whoi.edu/timeline?foo=bar&dataset=mydata&baz=1")
+ expect_equal(result$base_url, "https://habon-ifcb.whoi.edu")
+ expect_equal(result$dataset_name, "mydata")
+})
+
+test_that("parse_dashboard_url rejects NULL input", {
+ expect_error(parse_dashboard_url(NULL), "non-empty character string")
+})
+
+test_that("parse_dashboard_url rejects empty string", {
+ expect_error(parse_dashboard_url(""), "non-empty character string")
+})
+
+test_that("parse_dashboard_url rejects non-HTTP URLs", {
+ expect_error(parse_dashboard_url("file:///etc/passwd"), "http:// or https://")
+ expect_error(parse_dashboard_url("ftp://example.com"), "http:// or https://")
+ expect_error(parse_dashboard_url("javascript:alert(1)"), "http:// or https://")
+})
+
+test_that("get_dashboard_cache_dir returns a path", {
+ cache_dir <- get_dashboard_cache_dir()
+ expect_type(cache_dir, "character")
+ expect_true(nzchar(cache_dir))
+ expect_true(grepl("dashboard", cache_dir))
+})
+
+# ---------------------------------------------------------------------------
+# list_dashboard_bins — error handling (offline-safe, no network needed)
+# ---------------------------------------------------------------------------
+
+test_that("list_dashboard_bins errors on unreachable host", {
+ expect_error(
+ list_dashboard_bins("https://this-host-does-not-exist.invalid"),
+ "Failed to connect"
+ )
+})
+
+test_that("list_dashboard_bins builds correct API URL without dataset", {
+ # Trailing slashes should be stripped before appending /api/list_bins
+ # We cannot hit the network here, but we can verify the error message
+ # contains the expected URL fragment
+
+ err <- tryCatch(
+ list_dashboard_bins("https://this-host-does-not-exist.invalid/"),
+ error = function(e) e$message
+ )
+ expect_true(grepl("this-host-does-not-exist", err))
+})
+
+test_that("list_dashboard_bins builds correct API URL with dataset", {
+ err <- tryCatch(
+ list_dashboard_bins("https://this-host-does-not-exist.invalid", "mydata"),
+ error = function(e) e$message
+ )
+ expect_true(grepl("this-host-does-not-exist", err))
+})
+
+# ---------------------------------------------------------------------------
+# download_dashboard_images — cache-hit path (offline-safe)
+# ---------------------------------------------------------------------------
+
+test_that("download_dashboard_images returns cached folder when PNGs exist", {
+ tmp <- file.path(tempdir(), "dashboard_test_cache_hit")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ sample_name <- "D20160810T104734_IFCB110"
+ png_subfolder <- file.path(tmp, sample_name, sample_name)
+ dir.create(png_subfolder, recursive = TRUE)
+ file.create(file.path(png_subfolder, paste0(sample_name, "_00001.png")))
+
+ result <- download_dashboard_images(
+ base_url = "https://unused.example.com",
+ sample_name = sample_name,
+ cache_dir = tmp
+ )
+ expect_equal(result, file.path(tmp, sample_name))
+})
+
+test_that("download_dashboard_images skips download when cache has PNGs", {
+ tmp <- file.path(tempdir(), "dashboard_test_cache_skip")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ sample_name <- "D20160810T104734_IFCB110"
+ png_subfolder <- file.path(tmp, sample_name, sample_name)
+ dir.create(png_subfolder, recursive = TRUE)
+ file.create(file.path(png_subfolder, "img1.png"))
+ file.create(file.path(png_subfolder, "img2.png"))
+
+ # Should return immediately without hitting the network
+ result <- download_dashboard_images("https://unused.example.com", sample_name, tmp)
+ expect_equal(result, file.path(tmp, sample_name))
+ expect_equal(length(list.files(png_subfolder, pattern = "\\.png$")), 2)
+})
+
+# ---------------------------------------------------------------------------
+# download_dashboard_adc — cache-hit path (offline-safe)
+# ---------------------------------------------------------------------------
+
+test_that("download_dashboard_adc returns cached ADC file when it exists", {
+ tmp <- file.path(tempdir(), "dashboard_test_adc_cache")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ sample_name <- "D20160810T104734_IFCB110"
+ date_part <- substr(sample_name, 1, 9)
+ adc_dir <- file.path(tmp, date_part)
+ dir.create(adc_dir, recursive = TRUE)
+ adc_path <- file.path(adc_dir, paste0(sample_name, ".adc"))
+ writeLines("1,2,3", adc_path)
+
+ result <- download_dashboard_adc("https://unused.example.com", sample_name, tmp)
+ expect_equal(result, adc_path)
+})
+
+# ---------------------------------------------------------------------------
+# download_dashboard_images — error handling (offline-safe)
+# ---------------------------------------------------------------------------
+
+test_that("download_dashboard_images returns NULL on download failure", {
+ tmp <- file.path(tempdir(), "dashboard_test_dl_fail")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- suppressWarnings(
+ download_dashboard_images(
+ "https://this-host-does-not-exist.invalid",
+ "D20160810T104734_IFCB110",
+ tmp
+ )
+ )
+ expect_null(result)
+})
+
+# ---------------------------------------------------------------------------
+# download_dashboard_adc — error handling (offline-safe)
+# ---------------------------------------------------------------------------
+
+test_that("download_dashboard_adc returns NULL on download failure", {
+ tmp <- file.path(tempdir(), "dashboard_test_adc_fail")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- suppressWarnings(
+ download_dashboard_adc(
+ "https://this-host-does-not-exist.invalid",
+ "D20160810T104734_IFCB110",
+ tmp
+ )
+ )
+ expect_null(result)
+})
+
+# ---------------------------------------------------------------------------
+# download_dashboard_autoclass — error handling (offline-safe)
+# ---------------------------------------------------------------------------
+
+test_that("download_dashboard_autoclass returns NULL on download failure", {
+ tmp <- file.path(tempdir(), "dashboard_test_autoclass_fail")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- suppressWarnings(
+ download_dashboard_autoclass(
+ "https://this-host-does-not-exist.invalid",
+ "D20160810T104734_IFCB110",
+ tmp
+ )
+ )
+ expect_null(result)
+})
+
+# ---------------------------------------------------------------------------
+# download_dashboard_images_bulk — cache logic (offline-safe)
+# ---------------------------------------------------------------------------
+
+test_that("download_dashboard_images_bulk skips already-cached samples", {
+ tmp <- file.path(tempdir(), "dashboard_test_bulk_cache")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ # Pre-populate cache for one sample
+ s1 <- "D20160810T104734_IFCB110"
+ s1_sub <- file.path(tmp, s1, s1)
+ dir.create(s1_sub, recursive = TRUE)
+ file.create(file.path(s1_sub, paste0(s1, "_00001.png")))
+
+ # The second sample is not cached and the host doesn't exist,
+ # so it will fail to download — but the cached sample should still be returned
+ s2 <- "D20160810T112000_IFCB110"
+
+ result <- suppressWarnings(
+ download_dashboard_images_bulk(
+ "https://this-host-does-not-exist.invalid",
+ c(s1, s2),
+ tmp
+ )
+ )
+ expect_true(s1 %in% result)
+ expect_false(s2 %in% result)
+})
+
+test_that("download_dashboard_images_bulk returns empty when all fail", {
+ tmp <- file.path(tempdir(), "dashboard_test_bulk_allfail")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- suppressWarnings(
+ download_dashboard_images_bulk(
+ "https://this-host-does-not-exist.invalid",
+ c("D20160810T104734_IFCB110", "D20160810T112000_IFCB110"),
+ tmp
+ )
+ )
+ expect_length(result, 0)
+})
+
+test_that("download_dashboard_images_bulk returns all when all cached", {
+ tmp <- file.path(tempdir(), "dashboard_test_bulk_allcached")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ samples <- c("D20160810T104734_IFCB110", "D20160810T112000_IFCB110")
+ for (sn in samples) {
+ sub <- file.path(tmp, sn, sn)
+ dir.create(sub, recursive = TRUE)
+ file.create(file.path(sub, paste0(sn, "_00001.png")))
+ }
+
+ result <- download_dashboard_images_bulk("https://unused.example.com", samples, tmp)
+ expect_equal(sort(result), sort(samples))
+})
+
+# ---------------------------------------------------------------------------
+# download_dashboard_image_single — cache-hit path (offline-safe)
+# ---------------------------------------------------------------------------
+
+test_that("download_dashboard_image_single returns cached file when it exists", {
+ tmp <- file.path(tempdir(), "dashboard_test_single_cache")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ sample_name <- "D20160810T104734_IFCB110"
+ roi_number <- 42
+ file_name <- sprintf("%s_%05d.png", sample_name, roi_number)
+ dest_folder <- file.path(tmp, sample_name)
+ dir.create(dest_folder, recursive = TRUE)
+ dest_path <- file.path(dest_folder, file_name)
+ writeBin(charToRaw("fake png"), dest_path)
+
+ result <- download_dashboard_image_single(
+ base_url = "https://unused.example.com",
+ sample_name = sample_name,
+ roi_number = roi_number,
+ dest_dir = tmp
+ )
+ expect_equal(result, dest_path)
+})
+
+test_that("download_dashboard_image_single returns NULL on download failure", {
+ tmp <- file.path(tempdir(), "dashboard_test_single_fail")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- download_dashboard_image_single(
+ base_url = "https://this-host-does-not-exist.invalid",
+ sample_name = "D20160810T104734_IFCB110",
+ roi_number = 1,
+ dest_dir = tmp,
+ max_retries = 1
+ )
+ expect_null(result)
+})
+
+test_that("download_dashboard_image_single creates dest directory", {
+ tmp <- file.path(tempdir(), "dashboard_test_single_mkdir")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ # Will fail to download but should still create the directory
+ result <- download_dashboard_image_single(
+ base_url = "https://this-host-does-not-exist.invalid",
+ sample_name = "D20160810T104734_IFCB110",
+ roi_number = 1,
+ dest_dir = tmp,
+ max_retries = 1
+ )
+ expect_null(result)
+ expect_true(dir.exists(file.path(tmp, "D20160810T104734_IFCB110")))
+})
+
+test_that("download_dashboard_image_single formats ROI number with 5 digits", {
+ tmp <- file.path(tempdir(), "dashboard_test_single_format")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ sample_name <- "D20160810T104734_IFCB110"
+ # Pre-create the file with the expected 5-digit format
+ dest_folder <- file.path(tmp, sample_name)
+ dir.create(dest_folder, recursive = TRUE)
+ expected_file <- file.path(dest_folder, paste0(sample_name, "_00007.png"))
+ writeBin(charToRaw("fake png"), expected_file)
+
+ result <- download_dashboard_image_single(
+ base_url = "https://unused.example.com",
+ sample_name = sample_name,
+ roi_number = 7,
+ dest_dir = tmp
+ )
+ expect_equal(result, expected_file)
+})
+
+# ---------------------------------------------------------------------------
+# download_dashboard_images_individual — offline-safe
+# ---------------------------------------------------------------------------
+
+test_that("download_dashboard_images_individual returns cached files", {
+ tmp <- file.path(tempdir(), "dashboard_test_indiv_cache")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ sample_name <- "D20160810T104734_IFCB110"
+ dest_folder <- file.path(tmp, sample_name)
+ dir.create(dest_folder, recursive = TRUE)
+
+ # Pre-create two files
+ f1 <- paste0(sample_name, "_00001.png")
+ f2 <- paste0(sample_name, "_00042.png")
+ writeBin(charToRaw("fake png 1"), file.path(dest_folder, f1))
+ writeBin(charToRaw("fake png 2"), file.path(dest_folder, f2))
+
+ result <- download_dashboard_images_individual(
+ base_url = "https://unused.example.com",
+ file_names = c(f1, f2),
+ dest_dir = tmp
+ )
+ expect_equal(sort(result), sort(c(f1, f2)))
+})
+
+test_that("download_dashboard_images_individual returns empty on all failures", {
+ tmp <- file.path(tempdir(), "dashboard_test_indiv_fail")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- suppressWarnings(
+ download_dashboard_images_individual(
+ base_url = "https://this-host-does-not-exist.invalid",
+ file_names = c("D20160810T104734_IFCB110_00001.png",
+ "D20160810T104734_IFCB110_00002.png"),
+ dest_dir = tmp,
+ max_retries = 1
+ )
+ )
+ expect_length(result, 0)
+})
+
+test_that("download_dashboard_images_individual handles mixed cached and uncached", {
+ tmp <- file.path(tempdir(), "dashboard_test_indiv_mixed")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ sample_name <- "D20160810T104734_IFCB110"
+ dest_folder <- file.path(tmp, sample_name)
+ dir.create(dest_folder, recursive = TRUE)
+
+ # Pre-create only the first file
+ f1 <- paste0(sample_name, "_00001.png")
+ f2 <- paste0(sample_name, "_00099.png")
+ writeBin(charToRaw("fake png"), file.path(dest_folder, f1))
+
+ result <- download_dashboard_images_individual(
+ base_url = "https://this-host-does-not-exist.invalid",
+ file_names = c(f1, f2),
+ dest_dir = tmp,
+ max_retries = 1
+ )
+ expect_true(f1 %in% result)
+ expect_false(f2 %in% result)
+})
+
+test_that("download_dashboard_images_individual skips malformed file names", {
+ tmp <- file.path(tempdir(), "dashboard_test_indiv_malformed")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- download_dashboard_images_individual(
+ base_url = "https://unused.example.com",
+ file_names = c("not_a_valid_filename.txt", "also-bad"),
+ dest_dir = tmp,
+ max_retries = 1
+ )
+ expect_length(result, 0)
+})
+
+test_that("download_dashboard_images_individual skips unavailable samples after threshold", {
+ tmp <- file.path(tempdir(), "dashboard_test_indiv_skip")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ # 5 images from a non-existent host — after 2 failures the rest should be skipped
+ file_names <- sprintf("D20160810T104734_IFCB999_%05d.png", 1:5)
+
+ expect_warning(
+ result <- download_dashboard_images_individual(
+ base_url = "https://this-host-does-not-exist.invalid",
+ file_names = file_names,
+ dest_dir = tmp,
+ max_retries = 1,
+ sample_fail_threshold = 2
+ ),
+ "Skipping remaining images"
+ )
+ expect_length(result, 0)
+})
+
+test_that("download_dashboard_images_individual handles files from multiple samples", {
+ tmp <- file.path(tempdir(), "dashboard_test_indiv_multi")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ s1 <- "D20160810T104734_IFCB110"
+ s2 <- "D20160811T090000_IFCB110"
+ dir.create(file.path(tmp, s1), recursive = TRUE)
+ dir.create(file.path(tmp, s2), recursive = TRUE)
+
+ f1 <- paste0(s1, "_00001.png")
+ f2 <- paste0(s2, "_00005.png")
+ writeBin(charToRaw("fake"), file.path(tmp, s1, f1))
+ writeBin(charToRaw("fake"), file.path(tmp, s2, f2))
+
+ result <- download_dashboard_images_individual(
+ base_url = "https://unused.example.com",
+ file_names = c(f1, f2),
+ dest_dir = tmp
+ )
+ expect_equal(sort(result), sort(c(f1, f2)))
+})
+
+# ---------------------------------------------------------------------------
+# rescan_file_index — dashboard mode, offline-safe
+# ---------------------------------------------------------------------------
+
+test_that("rescan_file_index returns NULL when dashboard URL is empty", {
+ result <- rescan_file_index(data_source = "dashboard", dashboard_url = "")
+ expect_null(result)
+})
+
+test_that("rescan_file_index returns NULL when dashboard URL is NULL", {
+ result <- rescan_file_index(data_source = "dashboard", dashboard_url = NULL)
+ expect_null(result)
+})
+
+# ---------------------------------------------------------------------------
+# Integration tests — require network access to habon-ifcb.whoi.edu
+# ---------------------------------------------------------------------------
+
+# Helper: skip unless the dashboard is actually reachable
+skip_if_dashboard_unavailable <- function() {
+ skip_on_cran()
+ skip_if_offline()
+ avail <- tryCatch({
+ resp <- curl::curl_fetch_memory(
+ "https://habon-ifcb.whoi.edu/api/list_bins?dataset=tangosund",
+ handle = curl::new_handle(timeout = 15)
+ )
+ resp$status_code == 200
+ }, error = function(e) FALSE)
+ skip_if_not(avail, "IFCB Dashboard at habon-ifcb.whoi.edu is not reachable")
+}
+
+# A small dataset with a known sample
+DASHBOARD_BASE <- "https://habon-ifcb.whoi.edu"
+DASHBOARD_DATASET <- "tangosund"
+# Pick a specific small sample to keep downloads fast
+DASHBOARD_SAMPLE <- "D20160810T104734_IFCB110"
+
+# Pick a sample with class_scores
+DASHBOARD_CLASS_DATASET <- "mvco"
+DASHBOARD_CLASS_SAMPLE <- "D20190402T200352_IFCB010"
+
+test_that("list_dashboard_bins returns character vector from real API", {
+ skip_if_dashboard_unavailable()
+
+ bins <- list_dashboard_bins(DASHBOARD_BASE, DASHBOARD_DATASET)
+ expect_type(bins, "character")
+ expect_true(length(bins) > 0)
+ # All bins should look like IFCB sample names
+ expect_true(all(grepl("^D\\d{8}T\\d{6}_IFCB\\d+", bins)))
+})
+
+test_that("list_dashboard_bins without dataset returns bins", {
+ skip_if_dashboard_unavailable()
+
+ bins <- list_dashboard_bins(DASHBOARD_BASE)
+ expect_type(bins, "character")
+ expect_true(length(bins) > 0)
+})
+
+test_that("list_dashboard_bins includes known sample", {
+ skip_if_dashboard_unavailable()
+
+ bins <- list_dashboard_bins(DASHBOARD_BASE, DASHBOARD_DATASET)
+ expect_true(DASHBOARD_SAMPLE %in% bins)
+})
+
+test_that("download_dashboard_images downloads and extracts PNGs", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_images")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- download_dashboard_images(DASHBOARD_BASE, DASHBOARD_SAMPLE, tmp)
+
+ expect_false(is.null(result))
+ expect_true(dir.exists(result))
+ pngs <- list.files(file.path(result, DASHBOARD_SAMPLE), pattern = "\\.png$")
+ expect_true(length(pngs) > 0)
+})
+
+test_that("download_dashboard_images uses cache on second call", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_cache")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ # First call downloads
+ r1 <- download_dashboard_images(DASHBOARD_BASE, DASHBOARD_SAMPLE, tmp)
+ expect_false(is.null(r1))
+
+ # Second call should return immediately from cache (same result)
+ r2 <- download_dashboard_images(DASHBOARD_BASE, DASHBOARD_SAMPLE, tmp)
+ expect_equal(r1, r2)
+})
+
+test_that("download_dashboard_adc downloads ADC file", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_adc")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- download_dashboard_adc(DASHBOARD_BASE, DASHBOARD_SAMPLE, tmp)
+
+ expect_false(is.null(result))
+ expect_true(file.exists(result))
+ expect_true(grepl("\\.adc$", result))
+ expect_true(file.size(result) > 0)
+})
+
+test_that("download_dashboard_adc uses cache on second call", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_adc_cache")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ r1 <- download_dashboard_adc(DASHBOARD_BASE, DASHBOARD_SAMPLE, tmp)
+ expect_false(is.null(r1))
+
+ r2 <- download_dashboard_adc(DASHBOARD_BASE, DASHBOARD_SAMPLE, tmp)
+ expect_equal(r1, r2)
+})
+
+test_that("download_dashboard_autoclass returns data frame with expected columns", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_autoclass")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- download_dashboard_autoclass(paste0(DASHBOARD_BASE, "/", DASHBOARD_CLASS_DATASET), DASHBOARD_CLASS_SAMPLE, tmp)
+
+ # autoclass may not be available for every sample — skip if NULL
+ skip_if(is.null(result), "No autoclass data available for test sample")
+
+ expect_s3_class(result, "data.frame")
+ expect_true(all(c("file_name", "class_name", "score") %in% names(result)))
+ expect_true(nrow(result) > 0)
+ # file_name should match the sample
+ expect_true(all(grepl(DASHBOARD_CLASS_SAMPLE, result$file_name)))
+ # scores should be numeric between 0 and 1
+ expect_true(all(result$score >= 0 & result$score <= 1))
+})
+
+test_that("download_dashboard_images_bulk downloads multiple samples", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_bulk")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ # Use a single sample to keep it fast
+ result <- download_dashboard_images_bulk(DASHBOARD_BASE, DASHBOARD_SAMPLE, tmp)
+
+ expect_true(DASHBOARD_SAMPLE %in% result)
+ pngs <- list.files(
+ file.path(tmp, DASHBOARD_SAMPLE, DASHBOARD_SAMPLE),
+ pattern = "\\.png$"
+ )
+ expect_true(length(pngs) > 0)
+})
+
+test_that("rescan_file_index works in dashboard mode with real API", {
+ skip_if_dashboard_unavailable()
+
+ # Use a temp db folder so we don't pollute real data
+ tmp_db <- file.path(tempdir(), "dashboard_integ_rescan_db")
+ on.exit(unlink(tmp_db, recursive = TRUE), add = TRUE)
+ dir.create(tmp_db, recursive = TRUE, showWarnings = FALSE)
+
+ result <- rescan_file_index(
+ data_source = "dashboard",
+ dashboard_url = paste0(
+ "https://habon-ifcb.whoi.edu/timeline?dataset=", DASHBOARD_DATASET
+ ),
+ db_folder = tmp_db,
+ verbose = FALSE
+ )
+
+ expect_type(result, "list")
+ expect_equal(result$data_source, "dashboard")
+ expect_equal(result$dashboard_base_url, DASHBOARD_BASE)
+ expect_equal(result$dashboard_dataset, DASHBOARD_DATASET)
+ expect_true(length(result$sample_names) > 0)
+ expect_true(DASHBOARD_SAMPLE %in% result$sample_names)
+})
+
+test_that("download_dashboard_image_single downloads a real PNG", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_single")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ result <- download_dashboard_image_single(
+ base_url = DASHBOARD_BASE,
+ sample_name = DASHBOARD_SAMPLE,
+ roi_number = 1,
+ dest_dir = tmp
+ )
+
+ expect_false(is.null(result))
+ expect_true(file.exists(result))
+ expect_true(file.info(result)$size > 0)
+ expect_true(grepl("\\.png$", result))
+})
+
+test_that("download_dashboard_image_single uses cache on second call", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_single_cache")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ r1 <- download_dashboard_image_single(DASHBOARD_BASE, DASHBOARD_SAMPLE, 1, tmp)
+ expect_false(is.null(r1))
+
+ r2 <- download_dashboard_image_single(DASHBOARD_BASE, DASHBOARD_SAMPLE, 1, tmp)
+ expect_equal(r1, r2)
+})
+
+test_that("download_dashboard_images_individual downloads real PNGs", {
+ skip_if_dashboard_unavailable()
+
+ tmp <- file.path(tempdir(), "dashboard_integ_individual")
+ on.exit(unlink(tmp, recursive = TRUE), add = TRUE)
+
+ file_names <- c(
+ paste0(DASHBOARD_SAMPLE, "_00001.png"),
+ paste0(DASHBOARD_SAMPLE, "_00002.png")
+ )
+
+ result <- download_dashboard_images_individual(
+ base_url = DASHBOARD_BASE,
+ file_names = file_names,
+ dest_dir = tmp
+ )
+
+ expect_true(length(result) > 0)
+ # At least the first ROI should exist
+ expect_true(file_names[1] %in% result)
+ # Verify the files actually exist on disk
+ for (f in result) {
+ sample_name <- sub("_\\d+\\.png$", "", f)
+ expect_true(file.exists(file.path(tmp, sample_name, f)))
+ }
+})
diff --git a/tests/testthat/test-database.R b/tests/testthat/test-database.R
index 0ba03d0..c9a18b4 100644
--- a/tests/testthat/test-database.R
+++ b/tests/testthat/test-database.R
@@ -501,6 +501,114 @@ test_that("export_db_to_mat returns FALSE for non-existent database", {
expect_false(result)
})
+test_that("export_db_to_mat returns FALSE when class list is missing", {
+ # Create a DB with annotations but manually remove the class_lists entries
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ sample_name <- "D20230101T120000_IFCB134"
+ class2use <- c("unclassified", "Diatom")
+
+ save_annotations_db(db_path, sample_name,
+ data.frame(file_name = paste0(sample_name, "_00001.png"),
+ class_name = "Diatom",
+ stringsAsFactors = FALSE),
+ class2use, "test")
+
+ # Remove the class_lists entries so the function hits the "no class list" path
+
+ con <- DBI::dbConnect(RSQLite::SQLite(), db_path)
+ DBI::dbExecute(con, "DELETE FROM class_lists WHERE sample_name = ?",
+ params = list(sample_name))
+ DBI::dbDisconnect(con)
+
+ expect_warning(
+ result <- export_db_to_mat(db_path, sample_name, tempdir()),
+ "No class list found"
+ )
+ expect_false(result)
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("import_all_mat_to_db returns zero counts for empty folder", {
+ mat_dir <- tempfile("mat_empty_")
+ dir.create(mat_dir)
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ result <- import_all_mat_to_db(mat_dir, db_path)
+ expect_equal(result$success, 0L)
+ expect_equal(result$failed, 0L)
+ expect_equal(result$skipped, 0L)
+
+ unlink(c(mat_dir, db_dir), recursive = TRUE)
+})
+
+test_that("import_all_mat_to_db excludes _class and class2use files", {
+ mat_dir <- tempfile("mat_filter_")
+ dir.create(mat_dir)
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ # Create files that should be excluded
+ file.create(file.path(mat_dir, "sample_A_class_v1.mat"))
+ file.create(file.path(mat_dir, "class2use_manual.mat"))
+
+ result <- import_all_mat_to_db(mat_dir, db_path)
+ # All files are filtered out, so nothing to import
+ expect_equal(result$success, 0L)
+ expect_equal(result$failed, 0L)
+ expect_equal(result$skipped, 0L)
+
+ unlink(c(mat_dir, db_dir), recursive = TRUE)
+})
+
+test_that("export_all_db_to_mat returns zero counts for empty database", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ # Create empty database (no annotations)
+ con <- DBI::dbConnect(RSQLite::SQLite(), db_path)
+ DBI::dbDisconnect(con)
+
+ result <- export_all_db_to_mat(db_path, tempdir())
+ expect_equal(result$success, 0L)
+ expect_equal(result$failed, 0L)
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("export_all_db_to_png returns zero counts for empty database", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ # Create empty database
+ con <- DBI::dbConnect(RSQLite::SQLite(), db_path)
+ DBI::dbDisconnect(con)
+
+ result <- export_all_db_to_png(db_path, tempdir(), list())
+ expect_equal(result$success, 0L)
+ expect_equal(result$failed, 0L)
+ expect_equal(result$skipped, 0L)
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("export_db_to_png returns FALSE for missing database", {
+ expect_warning(
+ result <- export_db_to_png("/nonexistent/db.sqlite", "sample",
+ "/nonexistent/file.roi", tempdir()),
+ "Database not found"
+ )
+ expect_false(result)
+})
+
test_that("import_all_mat_to_db imports multiple files and returns correct counts", {
skip_if_not_installed("iRfcb")
skip_if_not(reticulate::py_available(), "Python not available")
@@ -733,9 +841,10 @@ test_that("export_db_to_png skip_class with all ROIs skipped returns TRUE", {
)
save_annotations_db(db_path, sample_name, classifications, class2use, "test")
- roi_path <- testthat::test_path("test_data", "raw", "2022", "D20220522",
- "D20220522T000439_IFCB134.roi")
- skip_if_not(file.exists(roi_path), "Test ROI file not found")
+ # Function returns TRUE at skip_class filter before using the ROI file,
+ # so we only need a file that exists
+ roi_path <- tempfile(fileext = ".roi")
+ file.create(roi_path)
png_dir <- tempfile("png_")
dir.create(png_dir)
@@ -747,7 +856,7 @@ test_that("export_db_to_png skip_class with all ROIs skipped returns TRUE", {
# No class subfolders should be created
expect_equal(length(list.dirs(png_dir, recursive = FALSE)), 0)
- unlink(c(db_dir, png_dir), recursive = TRUE)
+ unlink(c(db_dir, png_dir, roi_path), recursive = TRUE)
})
test_that("export_db_to_png returns FALSE for missing sample", {
@@ -762,9 +871,9 @@ test_that("export_db_to_png returns FALSE for missing sample", {
stringsAsFactors = FALSE),
c("unclassified", "Diatom"), "test")
- roi_path <- testthat::test_path("test_data", "raw", "2022", "D20220522",
- "D20220522T000439_IFCB134.roi")
- skip_if_not(file.exists(roi_path), "Test ROI file not found")
+ # Need a real file for roi_path since the function checks existence first
+ roi_path <- tempfile(fileext = ".roi")
+ file.create(roi_path)
expect_warning(
result <- export_db_to_png(db_path, "nonexistent_sample", roi_path, tempdir()),
@@ -772,7 +881,7 @@ test_that("export_db_to_png returns FALSE for missing sample", {
)
expect_false(result)
- unlink(db_dir, recursive = TRUE)
+ unlink(c(db_dir, roi_path), recursive = TRUE)
})
test_that("export_db_to_png returns FALSE for missing ROI file", {
@@ -1112,3 +1221,298 @@ test_that("full roundtrip: .mat -> SQLite -> .mat preserves NaN and class list",
unlink(c(mat_dir, db_dir, export_dir), recursive = TRUE)
})
+
+# ============================================================================
+# Class Review Mode database functions
+# ============================================================================
+
+test_that("list_classes_db returns correct class counts", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ # Non-existent database returns empty data frame
+ result <- list_classes_db(db_path)
+ expect_s3_class(result, "data.frame")
+ expect_equal(nrow(result), 0)
+ expect_true(all(c("class_name", "count") %in% names(result)))
+
+ # Add annotations across two samples
+ class2use <- c("unclassified", "Diatom", "Ciliate")
+
+ save_annotations_db(db_path, "sample_A",
+ data.frame(file_name = sprintf("sample_A_%05d.png", 1:3),
+ class_name = c("Diatom", "Diatom", "Ciliate"),
+ stringsAsFactors = FALSE),
+ class2use, "test")
+ save_annotations_db(db_path, "sample_B",
+ data.frame(file_name = sprintf("sample_B_%05d.png", 1:2),
+ class_name = c("Ciliate", "Diatom"),
+ stringsAsFactors = FALSE),
+ class2use, "test")
+
+ result <- list_classes_db(db_path)
+ expect_equal(nrow(result), 2) # Ciliate and Diatom
+ expect_equal(result$class_name, c("Ciliate", "Diatom")) # alphabetical
+ expect_equal(result$count, c(2L, 3L))
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("load_class_annotations_db returns correct file_names", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ class2use <- c("unclassified", "Diatom", "Ciliate")
+
+ save_annotations_db(db_path, "sample_A",
+ data.frame(file_name = sprintf("sample_A_%05d.png", 1:3),
+ class_name = c("Diatom", "Ciliate", "Diatom"),
+ stringsAsFactors = FALSE),
+ class2use, "test")
+ save_annotations_db(db_path, "sample_B",
+ data.frame(file_name = sprintf("sample_B_%05d.png", 1:2),
+ class_name = c("Diatom", "Ciliate"),
+ stringsAsFactors = FALSE),
+ class2use, "test")
+
+ # Load all Diatom annotations
+ result <- load_class_annotations_db(db_path, "Diatom")
+ expect_s3_class(result, "data.frame")
+ expect_equal(nrow(result), 3)
+ expect_true(all(c("sample_name", "roi_number", "class_name", "file_name") %in% names(result)))
+
+ # Check file_name format
+ expect_equal(result$file_name,
+ c("sample_A_00001.png", "sample_A_00003.png", "sample_B_00001.png"))
+ expect_equal(result$sample_name, c("sample_A", "sample_A", "sample_B"))
+
+ # Non-existent class returns NULL
+ result2 <- load_class_annotations_db(db_path, "Nonexistent")
+ expect_null(result2)
+
+ # Non-existent database returns NULL
+ result3 <- load_class_annotations_db("/nonexistent/db.sqlite", "Diatom")
+ expect_null(result3)
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("save_class_review_changes_db updates only targeted rows", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ class2use <- c("unclassified", "Diatom", "Ciliate", "Dinoflagellate")
+
+ # Create two samples
+ save_annotations_db(db_path, "sample_A",
+ data.frame(file_name = sprintf("sample_A_%05d.png", 1:3),
+ class_name = c("Diatom", "Diatom", "Ciliate"),
+ stringsAsFactors = FALSE),
+ class2use, "OrigUser")
+ save_annotations_db(db_path, "sample_B",
+ data.frame(file_name = sprintf("sample_B_%05d.png", 1:2),
+ class_name = c("Diatom", "Ciliate"),
+ stringsAsFactors = FALSE),
+ class2use, "OrigUser")
+
+ # Reclassify: sample_A ROI 1 from Diatom to Dinoflagellate,
+ # sample_B ROI 2 from Ciliate to Diatom
+ changes <- data.frame(
+ sample_name = c("sample_A", "sample_B"),
+ roi_number = c(1L, 2L),
+ new_class_name = c("Dinoflagellate", "Diatom"),
+ stringsAsFactors = FALSE
+ )
+
+ updated <- save_class_review_changes_db(db_path, changes, "Reviewer")
+ expect_equal(updated, 2L)
+
+ # Verify only changed rows were updated
+ con <- DBI::dbConnect(RSQLite::SQLite(), db_path)
+ on.exit(DBI::dbDisconnect(con))
+
+ # sample_A ROI 1 should be Dinoflagellate with Reviewer annotator
+ row_a1 <- DBI::dbGetQuery(con,
+ "SELECT class_name, annotator, is_manual FROM annotations WHERE sample_name = 'sample_A' AND roi_number = 1")
+ expect_equal(row_a1$class_name, "Dinoflagellate")
+ expect_equal(row_a1$annotator, "Reviewer")
+ expect_equal(row_a1$is_manual, 1L)
+
+ # sample_A ROI 2 should be unchanged (still Diatom, OrigUser)
+ row_a2 <- DBI::dbGetQuery(con,
+ "SELECT class_name, annotator FROM annotations WHERE sample_name = 'sample_A' AND roi_number = 2")
+ expect_equal(row_a2$class_name, "Diatom")
+ expect_equal(row_a2$annotator, "OrigUser")
+
+ # sample_B ROI 2 should be Diatom with Reviewer annotator
+ row_b2 <- DBI::dbGetQuery(con,
+ "SELECT class_name, annotator FROM annotations WHERE sample_name = 'sample_B' AND roi_number = 2")
+ expect_equal(row_b2$class_name, "Diatom")
+ expect_equal(row_b2$annotator, "Reviewer")
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("list_classes_db filters by year, month, instrument", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ class2use <- c("Diatom", "Ciliate")
+
+ # Two samples: different year/month/instrument
+ save_annotations_db(db_path, "D20230615T120000_IFCB134",
+ data.frame(file_name = sprintf("D20230615T120000_IFCB134_%05d.png", 1:2),
+ class_name = c("Diatom", "Ciliate"),
+ stringsAsFactors = FALSE),
+ class2use, "test")
+ save_annotations_db(db_path, "D20240815T120000_IFCB135",
+ data.frame(file_name = sprintf("D20240815T120000_IFCB135_%05d.png", 1:3),
+ class_name = c("Diatom", "Diatom", "Ciliate"),
+ stringsAsFactors = FALSE),
+ class2use, "test")
+
+ # No filter — all 5 annotations
+ all <- list_classes_db(db_path)
+ expect_equal(sum(all$count), 5L)
+
+ # Filter by year
+ y2023 <- list_classes_db(db_path, year = "2023")
+ expect_equal(sum(y2023$count), 2L)
+
+ y2024 <- list_classes_db(db_path, year = "2024")
+ expect_equal(sum(y2024$count), 3L)
+
+ # Filter by month
+ m06 <- list_classes_db(db_path, month = "06")
+ expect_equal(sum(m06$count), 2L)
+
+ # Filter by instrument
+ i134 <- list_classes_db(db_path, instrument = "IFCB134")
+ expect_equal(sum(i134$count), 2L)
+
+ i135 <- list_classes_db(db_path, instrument = "IFCB135")
+ expect_equal(sum(i135$count), 3L)
+
+ # Combined filter
+ combo <- list_classes_db(db_path, year = "2024", instrument = "IFCB135")
+ expect_equal(sum(combo$count), 3L)
+
+ # Filter with no matches
+ empty <- list_classes_db(db_path, year = "2025")
+ expect_equal(nrow(empty), 0)
+
+ # "all" values are treated as no filter
+ all2 <- list_classes_db(db_path, year = "all", month = "all", instrument = "all")
+ expect_equal(sum(all2$count), 5L)
+
+ # Filter by annotator
+ by_test <- list_classes_db(db_path, annotator = "test")
+ expect_equal(sum(by_test$count), 5L)
+
+ by_nobody <- list_classes_db(db_path, annotator = "nobody")
+ expect_equal(nrow(by_nobody), 0)
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("load_class_annotations_db filters by year, month, instrument", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ class2use <- c("Diatom", "Ciliate")
+
+ save_annotations_db(db_path, "D20230615T120000_IFCB134",
+ data.frame(file_name = sprintf("D20230615T120000_IFCB134_%05d.png", 1:2),
+ class_name = c("Diatom", "Diatom"),
+ stringsAsFactors = FALSE),
+ class2use, "alice")
+ save_annotations_db(db_path, "D20240815T120000_IFCB135",
+ data.frame(file_name = sprintf("D20240815T120000_IFCB135_%05d.png", 1:3),
+ class_name = c("Diatom", "Diatom", "Diatom"),
+ stringsAsFactors = FALSE),
+ class2use, "bob")
+
+ # No filter — all 5 Diatom
+ all <- load_class_annotations_db(db_path, "Diatom")
+ expect_equal(nrow(all), 5)
+
+ # Filter by year
+ y2023 <- load_class_annotations_db(db_path, "Diatom", year = "2023")
+ expect_equal(nrow(y2023), 2)
+ expect_true(all(grepl("D2023", y2023$sample_name)))
+
+ # Filter by instrument
+ i135 <- load_class_annotations_db(db_path, "Diatom", instrument = "IFCB135")
+ expect_equal(nrow(i135), 3)
+
+ # Filter by annotator
+ by_alice <- load_class_annotations_db(db_path, "Diatom", annotator = "alice")
+ expect_equal(nrow(by_alice), 2)
+
+ by_bob <- load_class_annotations_db(db_path, "Diatom", annotator = "bob")
+ expect_equal(nrow(by_bob), 3)
+
+ by_nobody <- load_class_annotations_db(db_path, "Diatom", annotator = "nobody")
+ expect_null(by_nobody)
+
+ # Filter with no matches
+ none <- load_class_annotations_db(db_path, "Diatom", year = "2025")
+ expect_null(none)
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("list_annotation_metadata_db returns correct metadata", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ # Non-existent database
+ meta <- list_annotation_metadata_db(db_path)
+ expect_equal(meta$years, character())
+ expect_equal(meta$months, character())
+ expect_equal(meta$instruments, character())
+ expect_equal(meta$annotators, character())
+
+ class2use <- c("Diatom", "Ciliate")
+
+ save_annotations_db(db_path, "D20230615T120000_IFCB134",
+ data.frame(file_name = "D20230615T120000_IFCB134_00001.png",
+ class_name = "Diatom",
+ stringsAsFactors = FALSE),
+ class2use, "alice")
+ save_annotations_db(db_path, "D20240815T120000_IFCB135",
+ data.frame(file_name = "D20240815T120000_IFCB135_00001.png",
+ class_name = "Ciliate",
+ stringsAsFactors = FALSE),
+ class2use, "bob")
+
+ meta <- list_annotation_metadata_db(db_path)
+ expect_equal(meta$years, c("2023", "2024"))
+ expect_equal(meta$months, c("06", "08"))
+ expect_equal(meta$instruments, c("IFCB134", "IFCB135"))
+ expect_equal(meta$annotators, c("alice", "bob"))
+
+ unlink(db_dir, recursive = TRUE)
+})
+
+test_that("save_class_review_changes_db handles empty input", {
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+
+ # NULL input
+ expect_equal(save_class_review_changes_db(db_path, NULL, "test"), 0L)
+
+ # Empty data frame
+ empty_df <- data.frame(sample_name = character(), roi_number = integer(),
+ new_class_name = character(), stringsAsFactors = FALSE)
+ expect_equal(save_class_review_changes_db(db_path, empty_df, "test"), 0L)
+
+ unlink(db_dir, recursive = TRUE)
+})
diff --git a/tests/testthat/test-png-import.R b/tests/testthat/test-png-import.R
new file mode 100644
index 0000000..9d351a0
--- /dev/null
+++ b/tests/testthat/test-png-import.R
@@ -0,0 +1,300 @@
+# Tests for PNG folder import functionality
+
+library(testthat)
+
+# Helper: create a temporary PNG class folder structure
+create_test_png_folder <- function() {
+ base <- tempfile("png_import_")
+ dir.create(base)
+
+ # Create class subfolders with _NNN suffixes (iRfcb convention)
+ diatom_dir <- file.path(base, "Diatom_001")
+ ciliate_dir <- file.path(base, "Ciliate_002")
+ dir.create(diatom_dir)
+ dir.create(ciliate_dir)
+
+ # Create dummy PNG files (1x1 pixel PNGs)
+ png_bytes <- as.raw(c(
+ 0x89, 0x50, 0x4E, 0x47, 0x0D, 0x0A, 0x1A, 0x0A,
+ 0x00, 0x00, 0x00, 0x0D, 0x49, 0x48, 0x44, 0x52,
+ 0x00, 0x00, 0x00, 0x01, 0x00, 0x00, 0x00, 0x01,
+ 0x08, 0x02, 0x00, 0x00, 0x00, 0x90, 0x77, 0x53,
+ 0xDE, 0x00, 0x00, 0x00, 0x0C, 0x49, 0x44, 0x41,
+ 0x54, 0x08, 0xD7, 0x63, 0xF8, 0xCF, 0xC0, 0x00,
+ 0x00, 0x00, 0x02, 0x00, 0x01, 0xE2, 0x21, 0xBC,
+ 0x33, 0x00, 0x00, 0x00, 0x00, 0x49, 0x45, 0x4E,
+ 0x44, 0xAE, 0x42, 0x60, 0x82
+ ))
+
+ # Sample 1: two ROIs in Diatom, one in Ciliate
+ writeBin(png_bytes, file.path(diatom_dir, "D20230101T120000_IFCB134_00001.png"))
+ writeBin(png_bytes, file.path(diatom_dir, "D20230101T120000_IFCB134_00003.png"))
+ writeBin(png_bytes, file.path(ciliate_dir, "D20230101T120000_IFCB134_00002.png"))
+
+ # Sample 2: one ROI in Diatom
+ writeBin(png_bytes, file.path(diatom_dir, "D20230202T080000_IFCB134_00001.png"))
+
+ base
+}
+
+# ===========================================================================
+# scan_png_class_folder tests
+# ===========================================================================
+
+test_that("scan_png_class_folder parses folder structure correctly", {
+ png_folder <- create_test_png_folder()
+ on.exit(unlink(png_folder, recursive = TRUE))
+
+ result <- scan_png_class_folder(png_folder)
+
+ expect_type(result, "list")
+ expect_s3_class(result$annotations, "data.frame")
+ expect_equal(nrow(result$annotations), 4)
+
+ expect_true(all(c("sample_name", "roi_number", "file_name", "class_name")
+ %in% names(result$annotations)))
+
+ # Check class names have _NNN stripped
+ expect_equal(sort(result$classes_found), c("Ciliate", "Diatom"))
+
+ # Check sample names
+ expect_equal(sort(result$sample_names),
+ c("D20230101T120000_IFCB134", "D20230202T080000_IFCB134"))
+})
+
+test_that("scan_png_class_folder strips trailing _NNN from folder names", {
+ base <- tempfile("png_strip_")
+ dir.create(base)
+ on.exit(unlink(base, recursive = TRUE))
+
+ # Create folders with various suffix patterns
+ dir.create(file.path(base, "Mesodinium_rubrum_005"))
+ dir.create(file.path(base, "Strombidium-like_008"))
+ dir.create(file.path(base, "NoSuffix"))
+
+ png_bytes <- as.raw(c(
+ 0x89, 0x50, 0x4E, 0x47, 0x0D, 0x0A, 0x1A, 0x0A,
+ 0x00, 0x00, 0x00, 0x0D, 0x49, 0x48, 0x44, 0x52,
+ 0x00, 0x00, 0x00, 0x01, 0x00, 0x00, 0x00, 0x01,
+ 0x08, 0x02, 0x00, 0x00, 0x00, 0x90, 0x77, 0x53,
+ 0xDE, 0x00, 0x00, 0x00, 0x0C, 0x49, 0x44, 0x41,
+ 0x54, 0x08, 0xD7, 0x63, 0xF8, 0xCF, 0xC0, 0x00,
+ 0x00, 0x00, 0x02, 0x00, 0x01, 0xE2, 0x21, 0xBC,
+ 0x33, 0x00, 0x00, 0x00, 0x00, 0x49, 0x45, 0x4E,
+ 0x44, 0xAE, 0x42, 0x60, 0x82
+ ))
+
+ writeBin(png_bytes, file.path(base, "Mesodinium_rubrum_005", "D20230101T120000_IFCB134_00001.png"))
+ writeBin(png_bytes, file.path(base, "Strombidium-like_008", "D20230101T120000_IFCB134_00002.png"))
+ writeBin(png_bytes, file.path(base, "NoSuffix", "D20230101T120000_IFCB134_00003.png"))
+
+ result <- scan_png_class_folder(base)
+ expect_equal(sort(result$classes_found),
+ c("Mesodinium_rubrum", "NoSuffix", "Strombidium-like"))
+})
+
+test_that("scan_png_class_folder handles empty folder", {
+ empty <- tempfile("png_empty_")
+ dir.create(empty)
+ on.exit(unlink(empty, recursive = TRUE))
+
+ result <- scan_png_class_folder(empty)
+ expect_equal(nrow(result$annotations), 0)
+ expect_equal(length(result$classes_found), 0)
+ expect_equal(length(result$sample_names), 0)
+})
+
+test_that("scan_png_class_folder errors on nonexistent folder", {
+ expect_error(scan_png_class_folder("/nonexistent/path"),
+ "does not exist")
+})
+
+test_that("scan_png_class_folder warns about invalid filenames", {
+ base <- tempfile("png_bad_")
+ dir.create(base)
+ dir.create(file.path(base, "Diatom_001"))
+ on.exit(unlink(base, recursive = TRUE))
+
+ png_bytes <- as.raw(c(
+ 0x89, 0x50, 0x4E, 0x47, 0x0D, 0x0A, 0x1A, 0x0A,
+ 0x00, 0x00, 0x00, 0x0D, 0x49, 0x48, 0x44, 0x52,
+ 0x00, 0x00, 0x00, 0x01, 0x00, 0x00, 0x00, 0x01,
+ 0x08, 0x02, 0x00, 0x00, 0x00, 0x90, 0x77, 0x53,
+ 0xDE, 0x00, 0x00, 0x00, 0x0C, 0x49, 0x44, 0x41,
+ 0x54, 0x08, 0xD7, 0x63, 0xF8, 0xCF, 0xC0, 0x00,
+ 0x00, 0x00, 0x02, 0x00, 0x01, 0xE2, 0x21, 0xBC,
+ 0x33, 0x00, 0x00, 0x00, 0x00, 0x49, 0x45, 0x4E,
+ 0x44, 0xAE, 0x42, 0x60, 0x82
+ ))
+
+ writeBin(png_bytes, file.path(base, "Diatom_001", "badname.png"))
+ writeBin(png_bytes, file.path(base, "Diatom_001", "D20230101T120000_IFCB134_00001.png"))
+
+ expect_warning(
+ result <- scan_png_class_folder(base),
+ "unexpected name format"
+ )
+ expect_equal(nrow(result$annotations), 1)
+})
+
+test_that("scan_png_class_folder warns about duplicate ROIs", {
+ base <- tempfile("png_dup_")
+ dir.create(base)
+ dir.create(file.path(base, "Diatom_001"))
+ dir.create(file.path(base, "Ciliate_002"))
+ on.exit(unlink(base, recursive = TRUE))
+
+ png_bytes <- as.raw(c(
+ 0x89, 0x50, 0x4E, 0x47, 0x0D, 0x0A, 0x1A, 0x0A,
+ 0x00, 0x00, 0x00, 0x0D, 0x49, 0x48, 0x44, 0x52,
+ 0x00, 0x00, 0x00, 0x01, 0x00, 0x00, 0x00, 0x01,
+ 0x08, 0x02, 0x00, 0x00, 0x00, 0x90, 0x77, 0x53,
+ 0xDE, 0x00, 0x00, 0x00, 0x0C, 0x49, 0x44, 0x41,
+ 0x54, 0x08, 0xD7, 0x63, 0xF8, 0xCF, 0xC0, 0x00,
+ 0x00, 0x00, 0x02, 0x00, 0x01, 0xE2, 0x21, 0xBC,
+ 0x33, 0x00, 0x00, 0x00, 0x00, 0x49, 0x45, 0x4E,
+ 0x44, 0xAE, 0x42, 0x60, 0x82
+ ))
+
+ # Same file in two class folders
+ writeBin(png_bytes, file.path(base, "Diatom_001", "D20230101T120000_IFCB134_00001.png"))
+ writeBin(png_bytes, file.path(base, "Ciliate_002", "D20230101T120000_IFCB134_00001.png"))
+
+ expect_warning(
+ result <- scan_png_class_folder(base),
+ "Duplicate ROI"
+ )
+ # Only the first occurrence should be kept
+ expect_equal(nrow(result$annotations), 1)
+})
+
+# ===========================================================================
+# import_png_folder_to_db tests
+# ===========================================================================
+
+test_that("import_png_folder_to_db writes annotations to database", {
+ png_folder <- create_test_png_folder()
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+ on.exit({
+ unlink(png_folder, recursive = TRUE)
+ unlink(db_dir, recursive = TRUE)
+ })
+
+ class2use <- c("unclassified", "Diatom", "Ciliate")
+
+ result <- import_png_folder_to_db(png_folder, db_path, class2use,
+ annotator = "TestUser")
+
+ expect_equal(result$success, 2L) # 2 samples
+ expect_equal(result$failed, 0L)
+
+ # Verify database contents
+ samples <- list_annotated_samples_db(db_path)
+ expect_equal(sort(samples),
+ c("D20230101T120000_IFCB134", "D20230202T080000_IFCB134"))
+})
+
+test_that("import_png_folder_to_db applies class mapping", {
+ png_folder <- create_test_png_folder()
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+ on.exit({
+ unlink(png_folder, recursive = TRUE)
+ unlink(db_dir, recursive = TRUE)
+ })
+
+ class2use <- c("unclassified", "Renamed_Diatom", "Ciliate")
+
+ # Map "Diatom" -> "Renamed_Diatom"
+ result <- import_png_folder_to_db(
+ png_folder, db_path, class2use,
+ class_mapping = c("Diatom" = "Renamed_Diatom"),
+ annotator = "TestUser"
+ )
+
+ expect_equal(result$success, 2L)
+
+ # Verify mapped class names in database
+ con <- DBI::dbConnect(RSQLite::SQLite(), db_path)
+ on.exit(DBI::dbDisconnect(con), add = TRUE)
+
+ rows <- DBI::dbGetQuery(con,
+ "SELECT DISTINCT class_name FROM annotations ORDER BY class_name")
+ expect_true("Renamed_Diatom" %in% rows$class_name)
+ expect_false("Diatom" %in% rows$class_name)
+ expect_true("Ciliate" %in% rows$class_name)
+})
+
+test_that("import_png_folder_to_db overwrites existing samples", {
+ png_folder <- create_test_png_folder()
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+ on.exit({
+ unlink(png_folder, recursive = TRUE)
+ unlink(db_dir, recursive = TRUE)
+ })
+
+ class2use <- c("unclassified", "Diatom", "Ciliate")
+
+ # First import
+ result1 <- import_png_folder_to_db(png_folder, db_path, class2use,
+ annotator = "User1")
+ expect_equal(result1$success, 2L)
+
+ # Second import (should overwrite)
+ result2 <- import_png_folder_to_db(png_folder, db_path, class2use,
+ annotator = "User2")
+ expect_equal(result2$success, 2L)
+
+ # Verify annotator was updated (overwritten)
+ con <- DBI::dbConnect(RSQLite::SQLite(), db_path)
+ on.exit(DBI::dbDisconnect(con), add = TRUE)
+
+ rows <- DBI::dbGetQuery(con, "SELECT DISTINCT annotator FROM annotations")
+ expect_equal(rows$annotator, "User2")
+})
+
+test_that("import_png_folder_to_db handles empty folder", {
+ empty <- tempfile("png_empty_")
+ dir.create(empty)
+ db_dir <- tempfile("db_")
+ dir.create(db_dir)
+ db_path <- get_db_path(db_dir)
+ on.exit({
+ unlink(empty, recursive = TRUE)
+ unlink(db_dir, recursive = TRUE)
+ })
+
+ result <- import_png_folder_to_db(empty, db_path, c("unclassified"))
+ expect_equal(result$success, 0L)
+ expect_equal(result$failed, 0L)
+})
+
+# ===========================================================================
+# scan_png_class_folder with example_data
+# ===========================================================================
+
+test_that("scan_png_class_folder parses example_data/png correctly", {
+
+ example_png <- file.path(
+ testthat::test_path(), "test_data", "example_png"
+ )
+
+ skip_if_not(dir.exists(example_png), "example_data/png not available")
+
+ result <- scan_png_class_folder(example_png)
+
+ expect_true(nrow(result$annotations) > 0)
+ expect_true(length(result$classes_found) > 0)
+ expect_true(length(result$sample_names) > 0)
+
+ # Verify _NNN suffixes are stripped
+ expect_false(any(grepl("_\\d{3}$", result$classes_found)))
+
+ # Known classes from example data (without _NNN suffix)
+ expect_true("Mesodinium_rubrum" %in% result$classes_found)
+})
diff --git a/tests/testthat/test-sample_loading.R b/tests/testthat/test-sample_loading.R
index 4198e49..8945e09 100644
--- a/tests/testthat/test-sample_loading.R
+++ b/tests/testthat/test-sample_loading.R
@@ -95,6 +95,136 @@ test_that("filter_to_extracted returns all if folder doesn't exist", {
expect_equal(nrow(filtered), 2)
})
+test_that("load_from_csv with use_threshold=FALSE uses class_name_auto", {
+ temp_csv <- tempfile(fileext = ".csv")
+ mock_data <- data.frame(
+ file_name = c("D20230101_00001.png", "D20230101_00002.png"),
+ class_name = c("unclassified", "Diatom"),
+ class_name_auto = c("Ciliate", "Diatom"),
+ score = c(0.45, 0.95),
+ stringsAsFactors = FALSE
+ )
+ write.csv(mock_data, temp_csv, row.names = FALSE)
+
+ # With threshold (default) — uses class_name
+ with_threshold <- load_from_csv(temp_csv, use_threshold = TRUE)
+ expect_equal(with_threshold$class_name, c("unclassified", "Diatom"))
+
+ # Without threshold — uses class_name_auto
+ without_threshold <- load_from_csv(temp_csv, use_threshold = FALSE)
+ expect_equal(without_threshold$class_name, c("Ciliate", "Diatom"))
+
+ unlink(temp_csv)
+})
+
+test_that("load_from_csv with use_threshold=FALSE falls back when class_name_auto missing", {
+ temp_csv <- tempfile(fileext = ".csv")
+ mock_data <- data.frame(
+ file_name = c("D20230101_00001.png"),
+ class_name = c("Diatom"),
+ score = c(0.95),
+ stringsAsFactors = FALSE
+ )
+ write.csv(mock_data, temp_csv, row.names = FALSE)
+
+ # Without threshold but no class_name_auto column — uses class_name
+ result <- load_from_csv(temp_csv, use_threshold = FALSE)
+ expect_equal(result$class_name, "Diatom")
+
+ unlink(temp_csv)
+})
+
+test_that("load_from_h5 reads H5 classification file correctly", {
+ skip_if_not_installed("hdf5r")
+
+ h5_path <- testthat::test_path("test_data", "D20220522T000439_IFCB134_class.h5")
+ skip_if_not(file.exists(h5_path), "Test H5 file not found")
+
+ sample_name <- "D20220522T000439_IFCB134"
+
+ # Create mock roi_dimensions matching the H5 file's ROI numbers
+ h5 <- hdf5r::H5File$new(h5_path, "r")
+ roi_numbers <- h5[["roi_numbers"]]$read()
+ h5$close_all()
+
+ roi_dimensions <- data.frame(
+ roi_number = roi_numbers,
+ width = rep(100, length(roi_numbers)),
+ height = rep(100, length(roi_numbers)),
+ area = seq(10000, 10000 + length(roi_numbers) - 1)
+ )
+
+ classifications <- load_from_h5(
+ h5_path = h5_path,
+ sample_name = sample_name,
+ roi_dimensions = roi_dimensions,
+ use_threshold = TRUE
+ )
+
+ expect_s3_class(classifications, "data.frame")
+ expect_true(nrow(classifications) > 0)
+ expect_named(classifications, c("file_name", "class_name", "score", "width", "height", "roi_area"))
+ expect_type(classifications$class_name, "character")
+ expect_type(classifications$score, "double")
+ # All scores should be between 0 and 1
+ expect_true(all(classifications$score >= 0 & classifications$score <= 1))
+ # Should be sorted by area descending
+ expect_equal(classifications$roi_area, sort(classifications$roi_area, decreasing = TRUE))
+})
+
+test_that("load_from_h5 with use_threshold=FALSE uses class_name_auto", {
+ skip_if_not_installed("hdf5r")
+
+ h5_path <- testthat::test_path("test_data", "D20220522T000439_IFCB134_class.h5")
+ skip_if_not(file.exists(h5_path), "Test H5 file not found")
+
+ sample_name <- "D20220522T000439_IFCB134"
+
+ h5 <- hdf5r::H5File$new(h5_path, "r")
+ roi_numbers <- h5[["roi_numbers"]]$read()
+ h5$close_all()
+
+ roi_dimensions <- data.frame(
+ roi_number = roi_numbers,
+ width = rep(100, length(roi_numbers)),
+ height = rep(100, length(roi_numbers)),
+ area = seq(10000, 10000 + length(roi_numbers) - 1)
+ )
+
+ with_threshold <- load_from_h5(h5_path, sample_name, roi_dimensions, use_threshold = TRUE)
+ without_threshold <- load_from_h5(h5_path, sample_name, roi_dimensions, use_threshold = FALSE)
+
+ expect_s3_class(without_threshold, "data.frame")
+ expect_equal(nrow(with_threshold), nrow(without_threshold))
+ # Without threshold should have no "unclassified" (all have raw predictions)
+ expect_type(without_threshold$class_name, "character")
+})
+
+test_that("load_from_h5 errors without hdf5r package", {
+ # We can't truly unload hdf5r, but we can check the function exists
+ expect_true(is.function(load_from_h5))
+ expect_equal(
+ names(formals(load_from_h5)),
+ c("h5_path", "sample_name", "roi_dimensions", "use_threshold")
+ )
+})
+
+test_that("load_from_csv reads real CSV with class_name_auto", {
+ csv_path <- testthat::test_path("test_data", "D20220522T000439_IFCB134.csv")
+ skip_if_not(file.exists(csv_path), "Test CSV file not found")
+
+ # With threshold
+ with_threshold <- load_from_csv(csv_path, use_threshold = TRUE)
+ expect_s3_class(with_threshold, "data.frame")
+ expect_true(nrow(with_threshold) > 0)
+ expect_true("file_name" %in% names(with_threshold))
+ expect_true("class_name" %in% names(with_threshold))
+
+ # Without threshold
+ without_threshold <- load_from_csv(csv_path, use_threshold = FALSE)
+ expect_equal(nrow(with_threshold), nrow(without_threshold))
+})
+
test_that("load_from_classifier_mat handles class names correctly", {
skip_if_not_installed("iRfcb")
@@ -222,6 +352,39 @@ test_that("load_from_classifier_mat works with use_threshold=FALSE", {
expect_type(classifications$class_name, "character")
})
+test_that("rescan_file_index discovers H5 classifier files", {
+ # Create temp folder structure
+ roi_dir <- file.path(tempdir(), "test_h5_scan", "roi", "2022", "D20220522")
+ csv_dir <- file.path(tempdir(), "test_h5_scan", "classified")
+ output_dir <- file.path(tempdir(), "test_h5_scan", "output")
+ dir.create(roi_dir, recursive = TRUE, showWarnings = FALSE)
+ dir.create(csv_dir, recursive = TRUE, showWarnings = FALSE)
+ dir.create(output_dir, recursive = TRUE, showWarnings = FALSE)
+
+ # Create dummy ROI file
+ file.create(file.path(roi_dir, "D20220522T000439_IFCB134.roi"))
+
+ # Create dummy H5 and MAT classifier files
+ file.create(file.path(csv_dir, "D20220522T000439_IFCB134_class.h5"))
+ file.create(file.path(csv_dir, "D20220522T000439_IFCB134_class_v1.mat"))
+
+ result <- rescan_file_index(
+ roi_folder = file.path(tempdir(), "test_h5_scan", "roi"),
+ csv_folder = csv_dir,
+ output_folder = output_dir,
+ verbose = FALSE,
+ db_folder = tempdir()
+ )
+
+ expect_type(result, "list")
+ expect_true("D20220522T000439_IFCB134" %in% result$classified_samples)
+ expect_true("D20220522T000439_IFCB134" %in% names(result$classifier_h5_files))
+ expect_true("D20220522T000439_IFCB134" %in% names(result$classifier_mat_files))
+
+ # Cleanup
+ unlink(file.path(tempdir(), "test_h5_scan"), recursive = TRUE)
+})
+
test_that("read_roi_dimensions reads real ADC file correctly", {
adc_path <- testthat::test_path("test_data", "raw", "2022", "D20220522", "D20220522T000439_IFCB134.adc")
skip_if_not(file.exists(adc_path), "Test ADC file not found")
diff --git a/tests/testthat/test-utils.R b/tests/testthat/test-utils.R
index da8f9b8..3761701 100644
--- a/tests/testthat/test-utils.R
+++ b/tests/testthat/test-utils.R
@@ -616,6 +616,106 @@ test_that("rescan_file_index works with non-standard folder structure", {
unlink(temp_root, recursive = TRUE)
})
+test_that("rescan_file_index detects H5 and MAT classifier files", {
+ temp_root <- tempfile("h5_test_")
+ roi_folder <- file.path(temp_root, "raw")
+ csv_folder <- file.path(temp_root, "classified")
+ output_folder <- file.path(temp_root, "manual")
+ dir.create(roi_folder, recursive = TRUE)
+ dir.create(csv_folder, recursive = TRUE)
+ dir.create(output_folder, recursive = TRUE)
+
+ # Create ROI files
+ file.create(file.path(roi_folder, "D20230101T120000_IFCB134.roi"))
+ file.create(file.path(roi_folder, "D20230102T120000_IFCB134.roi"))
+ file.create(file.path(roi_folder, "D20230103T120000_IFCB134.roi"))
+
+ # Create H5 classifier file
+ file.create(file.path(csv_folder, "D20230101T120000_IFCB134_class_v1.h5"))
+
+ # Create MAT classifier file
+ file.create(file.path(csv_folder, "D20230102T120000_IFCB134_class_v1.mat"))
+
+ # Create CSV classifier file
+ writeLines("file_name,class_name",
+ file.path(csv_folder, "D20230103T120000_IFCB134.csv"))
+
+ result <- rescan_file_index(
+ roi_folder = roi_folder,
+ csv_folder = csv_folder,
+ output_folder = output_folder,
+ verbose = FALSE
+ )
+
+ expect_type(result, "list")
+ expect_length(result$sample_names, 3)
+
+ # H5 classifier detected
+ expect_true("D20230101T120000_IFCB134" %in%
+ names(result$classifier_h5_files))
+
+ # MAT classifier detected
+ expect_true("D20230102T120000_IFCB134" %in%
+ names(result$classifier_mat_files))
+
+ # All three should be classified
+ expect_length(result$classified_samples, 3)
+
+ unlink(temp_root, recursive = TRUE)
+})
+
+test_that("rescan_file_index uses db_folder for annotated sample lookup", {
+ temp_root <- tempfile("dbfolder_test_")
+ roi_folder <- file.path(temp_root, "raw")
+ output_folder <- file.path(temp_root, "manual")
+ db_folder <- file.path(temp_root, "db")
+ dir.create(roi_folder, recursive = TRUE)
+ dir.create(output_folder, recursive = TRUE)
+ dir.create(db_folder, recursive = TRUE)
+
+ # Create ROI file
+ file.create(file.path(roi_folder, "D20230101T120000_IFCB134.roi"))
+
+ # Save an annotation in the SQLite DB in db_folder
+ db_path <- get_db_path(db_folder)
+ save_annotations_db(
+ db_path, "D20230101T120000_IFCB134",
+ data.frame(file_name = "D20230101T120000_IFCB134_00001.png",
+ class_name = "Diatom", stringsAsFactors = FALSE),
+ c("unclassified", "Diatom"), "test"
+ )
+
+ result <- rescan_file_index(
+ roi_folder = roi_folder,
+ csv_folder = roi_folder, # no CSV files here
+ output_folder = output_folder,
+ db_folder = db_folder,
+ verbose = FALSE
+ )
+
+ expect_type(result, "list")
+ expect_true("D20230101T120000_IFCB134" %in% result$annotated_samples)
+
+ unlink(temp_root, recursive = TRUE)
+})
+
+test_that("rescan_file_index returns NULL when no ROI files found", {
+ temp_root <- tempfile("noroi_test_")
+ roi_folder <- file.path(temp_root, "empty_raw")
+ dir.create(roi_folder, recursive = TRUE)
+
+ result <- rescan_file_index(
+ roi_folder = roi_folder,
+ csv_folder = roi_folder,
+ output_folder = roi_folder,
+ verbose = FALSE
+ )
+
+ expect_null(result)
+
+ unlink(temp_root, recursive = TRUE)
+})
+
test_that("rescan_file_index reads folder paths from saved settings", {
# This test verifies that rescan_file_index falls back to saved settings
# We can't easily mock get_settings_path, so we test the fallback path:
diff --git a/tests/testthat/test_data/D20220522T000439_IFCB134.csv b/tests/testthat/test_data/D20220522T000439_IFCB134.csv
new file mode 100644
index 0000000..9ca8a0a
--- /dev/null
+++ b/tests/testthat/test_data/D20220522T000439_IFCB134.csv
@@ -0,0 +1,8 @@
+"file_name","class_name","class_name_auto","score"
+"D20220522T000439_IFCB134_00002.png","Mesodinium_rubrum","Mesodinium_rubrum",0.900653839111328
+"D20220522T000439_IFCB134_00003.png","Strombidium-like","Strombidium-like",0.997846364974976
+"D20220522T000439_IFCB134_00004.png","Mesodinium_rubrum","Mesodinium_rubrum",0.514867901802063
+"D20220522T000439_IFCB134_00005.png","Ciliophora","Ciliophora",0.852679908275604
+"D20220522T000439_IFCB134_00006.png","Mesodinium_rubrum","Mesodinium_rubrum",0.991099894046783
+"D20220522T000439_IFCB134_00007.png","Mesodinium_rubrum","Mesodinium_rubrum",0.909600496292114
+"D20220522T000439_IFCB134_00008.png","Mesodinium_rubrum","Mesodinium_rubrum",0.993842244148254
diff --git a/tests/testthat/test_data/D20220522T000439_IFCB134_class.h5 b/tests/testthat/test_data/D20220522T000439_IFCB134_class.h5
new file mode 100644
index 0000000..1ce5ef9
Binary files /dev/null and b/tests/testthat/test_data/D20220522T000439_IFCB134_class.h5 differ
diff --git a/tests/testthat/test_data/example_png/Mesodinium_rubrum/D20251013T113302_IFCB134_00651.png b/tests/testthat/test_data/example_png/Mesodinium_rubrum/D20251013T113302_IFCB134_00651.png
new file mode 100644
index 0000000..9200346
Binary files /dev/null and b/tests/testthat/test_data/example_png/Mesodinium_rubrum/D20251013T113302_IFCB134_00651.png differ
diff --git a/vignettes/class-management.Rmd b/vignettes/class-management.Rmd
index c111e40..1762956 100644
--- a/vignettes/class-management.Rmd
+++ b/vignettes/class-management.Rmd
@@ -46,9 +46,9 @@ When you save an annotation as a .mat file for ifcb-analysis, the file stores:
You can create a new class list directly in the app without uploading a file:
-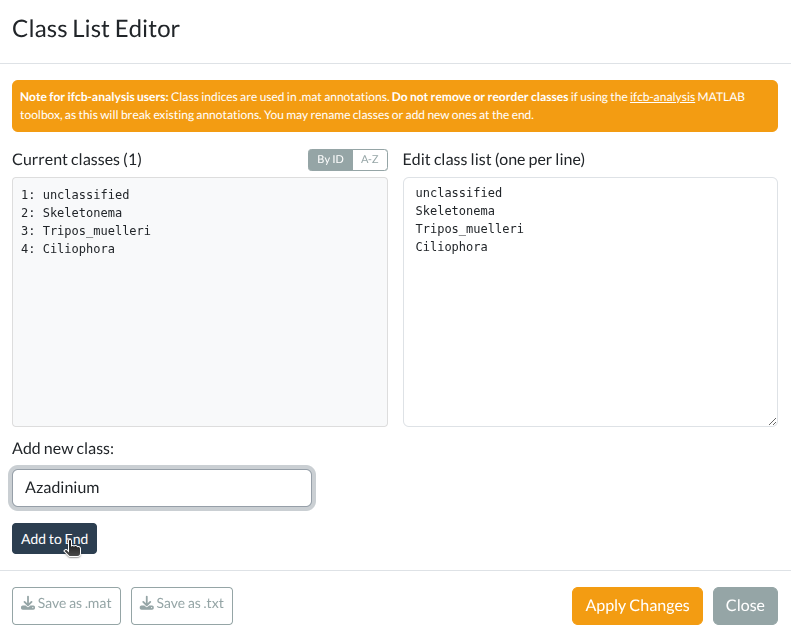 +
+ -
-Class list editor for adding, removing, and reordering classes. Click to enlarge.
+Class list editor showing classes with annotation counts, editing area, and export options. Click to enlarge.
1. Open **Settings** → **Edit Class List**
2. The editor opens with an empty class list
@@ -99,7 +99,7 @@ Diatom
## Viewing Classes
1. Open Settings → **Edit Class List**
-2. The left panel shows all classes with indices
+2. The left panel shows all classes with their indices and the number of annotated images per class (queried from the database)
3. Toggle **By ID** / **A-Z** to sort the view
**Note**: Sorting is for viewing only - it doesn't change actual indices.
diff --git a/vignettes/faq.Rmd b/vignettes/faq.Rmd
index 546de09..7b676c7 100644
--- a/vignettes/faq.Rmd
+++ b/vignettes/faq.Rmd
@@ -36,6 +36,18 @@ A: The app is specifically designed for IFCB data format. For other imaging syst
A: No. The app only reads your original files. All output is written to separate folders.
+**Q: How can I review all images of a specific class?**
+
+A: Use **Class Review mode**. Switch to "Class Review" using the mode toggle in the sidebar, select a class from the dropdown, and click Load. This loads all images annotated as that class from every sample in the database. You can then reclassify any mistakes using the normal relabeling tools.
+
+**Q: Can I reclassify images across multiple samples at once?**
+
+A: Yes. Class Review mode loads images from all samples and saves changes as row-level updates to the database. This means only the images you reclassify are updated — other annotations in those samples remain untouched.
+
+**Q: Can I classify images without pre-computed classifier files?**
+
+A: Yes. Configure a Gradio API URL and model in Settings > Live Prediction, then click the **Predict** button in the sidebar after loading a sample. This sends images to a remote CNN model and applies the predictions directly. See the [User Guide](user-guide.html#live-prediction) for details.
+
---
## Installation Issues
@@ -59,7 +71,7 @@ A: No. The default storage format is SQLite, which works out of the box with no
**Q: Where is the Python virtual environment created?**
-A: By default, `ifcb_py_install()` creates a `venv` folder in your home directory. You can specify a different location:
+A: By default, `ifcb_py_install()` creates a virtual environment at `~/.virtualenvs/iRfcb`. You can specify a different location:
```{r, eval = FALSE}
ifcb_py_install("/path/to/your/venv")
@@ -95,7 +107,7 @@ remotes::install_github("EuropeanIFCBGroup/ClassiPyR")
A: Try reinstalling the package:
```{r, eval = FALSE}
-install.packages("iRfcb")
+remotes::install_github("EuropeanIFCBGroup/ClassiPyR")
```
**Q: iRfcb won't install**
@@ -132,15 +144,19 @@ A: The app scans the ROI Data Folder recursively, so any subfolder layout works
A: For CSV files:
- Must have columns named `file_name` and `class_name` (exact names required)
-- Optionally include a `score` column (confidence value between 0 and 1)
+- Optionally include `score` and `class_name_auto` columns
- The CSV file must be named after the sample (e.g., `D20230101T120000_IFCB134.csv`)
- File should be in the Classification Folder (indexed via file cache; click Sync to refresh)
+For H5 files:
+
+- Must match pattern `*_class*.h5`
+- Requires the `hdf5r` package (`install.packages("hdf5r")`)
+
For MAT files:
- Must match pattern `*_class*.mat`
- Must contain `roinum` and `TBclass` variables
-- Must contain `roinum` and `TBclass` variables
---
@@ -158,15 +174,17 @@ D20230101T120000_IFCB134_00001.png,Diatom
D20230101T120000_IFCB134_00002.png,Ciliate
```
-With optional `score` column (confidence values between 0 and 1):
+With optional `score` and `class_name_auto` columns:
```
-file_name,class_name,score
-D20230101T120000_IFCB134_00001.png,Diatom,0.95
-D20230101T120000_IFCB134_00002.png,Ciliate,0.87
-D20230101T120000_IFCB134_00003.png,Dinoflagellate,0.72
+file_name,class_name,class_name_auto,score
+D20230101T120000_IFCB134_00001.png,unclassified,Diatom,0.45
+D20230101T120000_IFCB134_00002.png,Ciliate,Ciliate,0.87
+D20230101T120000_IFCB134_00003.png,Dinoflagellate,Dinoflagellate,0.72
```
+The `class_name_auto` column contains the raw prediction without threshold. When "Apply classification threshold" is disabled in Settings, ClassiPyR uses `class_name_auto` instead of `class_name`.
+
**Q: My CNN classifier outputs different column names**
A: The column names must be exactly `file_name` and `class_name`. If your classifier uses different names, rename the columns before loading. For example in R:
@@ -313,10 +331,9 @@ Or use the **Export SQLite → .mat** button in Settings.
```{r, eval = FALSE}
library(ClassiPyR)
-class2use <- load_class_list("/shared/network/class2use.mat")
db_path <- get_db_path(get_default_db_dir())
# Import .mat files from the shared folder into the local database
-result <- import_all_mat_to_db("/shared/network/manual", db_path, class2use)
+result <- import_all_mat_to_db("/shared/network/manual", db_path)
cat(result$success, "imported,", result$skipped, "skipped\n")
```
@@ -343,19 +360,17 @@ You can also import programmatically — a single file:
```{r, eval = FALSE}
library(ClassiPyR)
-class2use <- load_class_list("/path/to/class2use.mat")
import_mat_to_db(
mat_path = "/data/manual/D20230101T120000_IFCB134.mat",
db_path = get_db_path(get_default_db_dir()),
- sample_name = "D20230101T120000_IFCB134",
- class2use = class2use
+ sample_name = "D20230101T120000_IFCB134"
)
```
Or bulk-import all `.mat` files in a folder:
```{r, eval = FALSE}
-result <- import_all_mat_to_db("/data/manual", get_db_path(get_default_db_dir()), class2use)
+result <- import_all_mat_to_db("/data/manual", get_db_path(get_default_db_dir()))
cat(result$success, "imported,", result$failed, "failed,", result$skipped, "skipped\n")
```
@@ -374,6 +389,27 @@ result <- export_all_db_to_mat(get_db_path(get_default_db_dir()), "/data/manual"
cat(result$success, "exported,", result$failed, "failed\n")
```
+**Q: Can I import images classified in another tool?**
+
+A: Yes. Organize your PNG images into subfolders named after each class (e.g., `Diatom/`, `Ciliate_002/`). Then use **Import PNG → SQLite** in Settings > Import / Export. The app strips trailing `_NNN` suffixes from folder names (following the iRfcb convention) and maps images to class names based on which subfolder they are in.
+
+If your folder class names don't match the app's current class list, a mapping dialog will appear letting you remap them to existing classes or add them as new classes.
+
+Note: ROI files are needed for viewing images and re-exporting. Without ROI files, annotations are stored in the database but images cannot be displayed in the gallery.
+
+You can also import programmatically:
+
+```{r, eval = FALSE}
+library(ClassiPyR)
+result <- import_png_folder_to_db(
+ "/data/png_export",
+ get_db_path(get_default_db_dir()),
+ class2use = c("Diatom", "Ciliate", "Dinoflagellate"),
+ annotator = "Jane"
+)
+cat(result$success, "imported,", result$failed, "failed\n")
+```
+
**Q: Can I change the annotator name for existing annotations?**
A: Yes. Use `update_annotator()` from the R console:
diff --git a/vignettes/getting-started.Rmd b/vignettes/getting-started.Rmd
index 15e732c..6ae4beb 100644
--- a/vignettes/getting-started.Rmd
+++ b/vignettes/getting-started.Rmd
@@ -21,9 +21,10 @@ This tutorial walks you through your first session with `ClassiPyR`.
Make sure you have:
1. The package installed (see [Installation](https://europeanifcbgroup.github.io/ClassiPyR/))
-2. Your IFCB data files (ROI, ADC, HDR)
+2. Your IFCB data files (ROI, ADC, HDR) — or a remote IFCB Dashboard URL (see [Dashboard Mode](#dashboard-mode) below)
3. Optionally: a class list file (.mat or .txt) - you can also create one from scratch in the app
-4. Optionally: existing classifications (CSV or classifier MAT files, see below)
+4. Optionally: existing classifications (CSV, H5, or classifier MAT files, see below) — or use [Live Prediction](user-guide.html#live-prediction) to classify on the fly
+5. Optionally: existing PNG-organized datasets can be imported via Settings > Import PNG → SQLite
### Python Requirements
@@ -41,7 +42,7 @@ D20230101T120000_IFCB134_00001.png,Diatom
D20230101T120000_IFCB134_00002.png,Ciliate
```
-An optional `score` column (confidence values between 0 and 1) can also be included. See the [User Guide](user-guide.html) for more details.
+An optional `score` column (confidence values between 0 and 1) can also be included. HDF5 (`.h5`) and MATLAB (`.mat`) classifier output files from [iRfcb](https://github.com/EuropeanIFCBGroup/iRfcb) are also supported. See the [User Guide](user-guide.html) for more details.
### Python Setup (optional)
@@ -49,7 +50,7 @@ Only needed if you plan to export `.mat` files. Skip this step if using the defa
```{r, eval = FALSE}
library(iRfcb)
-ifcb_py_install(envname = "./venv") # Creates venv in current working directory
+ifcb_py_install() # Creates venv at ~/.virtualenvs/iRfcb by default
```
---
@@ -69,15 +70,15 @@ run_app(venv_path = "./venv")
Click the **gear icon** next to your username in the sidebar.
-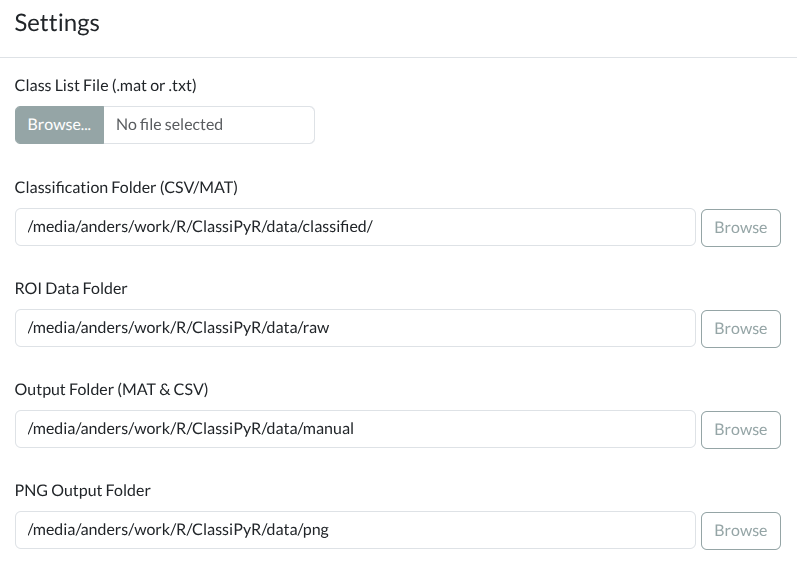 +
+ -
-Settings dialog showing folder configuration options. Click to enlarge.
+Settings dialog showing folder paths, classification threshold toggle, annotation storage, and import/export options. Click to enlarge.
Configure your folders using the built-in folder browser:
| Setting | Description | Example |
|---------|-------------|---------|
-| Classification Folder | Where your CSV/MAT classifications are | `/ifcb/classified/` |
+| Classification Folder | Where your CSV/H5/MAT classifications are | `/ifcb/classified/` |
| ROI Data Folder | Where your IFCB raw files are | `/ifcb/raw/` |
| Output Folder | Where MAT files and statistics go | `/ifcb/manual/` |
| Database Folder | Where the SQLite database is stored (must be local) | auto-detected |
@@ -209,7 +210,7 @@ The images will move to their new class group.
Click **Save Annotations** to save:
-- **SQLite database** (default) - annotations are written to `annotations.sqlite` in your Output Folder. This single file stores annotations for all samples. No Python needed.
+- **SQLite database** (default) - annotations are written to `annotations.sqlite` in your Database Folder. This single file stores annotations for all samples. No Python needed.
- Statistics CSV with accuracy metrics
- PNGs organized by class
@@ -238,8 +239,33 @@ Work is automatically saved when:
4. **Check statistics** - The "Validation Statistics" tab shows your progress
+5. **Use Live Prediction** - For unannotated samples, use the Predict button to get CNN classifications as a starting point, then correct mistakes manually
+
+---
+
+## Dashboard Mode {#dashboard-mode}
+
+If your IFCB data is hosted on a remote Dashboard (e.g. [habon-ifcb.whoi.edu](https://habon-ifcb.whoi.edu/)), you can work directly with the Dashboard without downloading data locally:
+
+1. Open **Settings** (gear icon)
+2. Under **Data Source**, select **IFCB Dashboard**
+3. Enter the Dashboard URL, e.g.:
+ - `https://habon-ifcb.whoi.edu/` (all datasets)
+ - `https://habon-ifcb.whoi.edu/timeline?dataset=tangosund` (specific dataset)
+4. Optionally check **Use dashboard auto-classifications** to load the dashboard's automated classifications for validation
+5. Optionally set a **Classification Folder** to use local CSV/H5/MAT files instead of (or as fallback to) dashboard auto-classifications
+6. Click **Save Settings**
+
+The app will fetch the sample list from the Dashboard API. When you load a sample, PNG images are downloaded on demand and cached locally for fast subsequent access. ADC files (for image dimensions and MAT export) are also downloaded as needed. Download parameters (parallel downloads, timeout, retries) can be tuned via the **Advanced Download Settings** section.
+
+> **Note**: MAT file export in dashboard mode requires downloading the ADC file for each sample. If the download fails, the app falls back to SQLite-only saving.
+
---
+## Reviewing Annotations Across Samples
+
+Once you have annotated several samples, you can use **Class Review mode** to verify annotations across the entire database. Switch to Class Review in the sidebar, use the searchable class dropdown to find a class (each class shows its image count), and load all images annotated as that class. Image labels show the full name (sample + ROI) so you can identify which sample each image belongs to. This makes it easy to spot and fix misclassifications. See the [User Guide](user-guide.html#class-review-mode) for details.
+
## Next Steps
- [User Guide](user-guide.html) - Complete feature documentation
diff --git a/vignettes/user-guide.Rmd b/vignettes/user-guide.Rmd
index a8d20ec..4517431 100644
--- a/vignettes/user-guide.Rmd
+++ b/vignettes/user-guide.Rmd
@@ -32,6 +32,7 @@ Complete documentation for all `ClassiPyR` features.
- No sample loaded: Initial state before selecting a sample
- Validation mode: Shows accuracy percentage
- Annotation mode: Shows progress (X/Y classified)
+ - Class Review mode: Shows selected class and image counts
### Sidebar
@@ -41,6 +42,7 @@ Complete documentation for all `ClassiPyR` features.
- **Navigation**: Load, previous, next, random, sync
- **Cache age**: Shows when folders were last scanned
- **Save button**: Manual save trigger
+- **Predict button**: One-click CNN classification (when configured)
### Main Area (Tabs)
@@ -76,6 +78,63 @@ Some samples may have both manual annotations AND auto-classifications (e.g., yo
---
+## Class Review Mode
+
+Class Review mode lets you view **all annotated images of a specific class** across the entire database. This is useful for verifying that annotations are consistent and correcting misclassified images without loading samples one by one.
+
+
+ +
+
+
+Class Review mode showing all images of a selected class across the database. Click to enlarge.
+
+### How to use
+
+1. Switch to **Class Review** using the mode toggle in the sidebar
+2. The class dropdown is searchable — type to filter. Each class shows its image count in parentheses
+3. Select a class and click **Load** — images are extracted from ROI files across all samples
+4. Review images in the gallery. Each image label shows the full name (sample + ROI) so you can identify which sample it belongs to
+5. Use the same selection and relabeling tools as in sample mode
+6. Click **Save Changes** to write row-level updates to the database
+
+### Key differences from sample mode
+
+- Images come from **multiple samples**, not just one
+- Image labels show the **full image name** (e.g. `D20230101T120000_IFCB134_00001`) instead of just the ROI number
+- Saving performs **surgical row-level UPDATEs** rather than replacing an entire sample's annotations
+- The title bar turns **purple** to indicate class review mode
+- Navigation buttons (prev/next/random sample) are not available
+
+---
+
+## Live Prediction
+
+The **Predict** button in the sidebar lets you classify all images in the loaded sample using a remote CNN model, without needing pre-computed classifier files. This uses `iRfcb::ifcb_classify_images()` to send images to a Gradio-hosted classification model.
+
+### Setup
+
+1. Open **Settings** (gear icon)
+2. Under **Live Prediction**, enter a Gradio API URL (e.g. `https://irfcb-classify.hf.space`)
+3. The **Prediction Model** dropdown is populated automatically from the server
+4. Select a model and click **Save Settings**
+
+The Predict button appears in the sidebar once a Gradio URL and model are configured.
+
+### Using Predict
+
+1. Load a sample in Sample Mode (annotated or unannotated)
+2. Click **Predict** in the sidebar
+3. A progress bar shows per-image classification progress
+4. When complete, the app switches to validation mode with the predicted classes
+
+### Behaviour details
+
+- **Manual labels are preserved**: If you have already reclassified some images, those are skipped during prediction. Only unchanged images are sent to the model.
+- **Threshold setting applies**: The "Apply classification threshold" setting controls whether thresholded or raw predictions are used, just like for CSV/H5/MAT classifications.
+- **New classes are added automatically**: If the model returns class names not in your current class list, they are added so the filter and relabel dropdowns work correctly.
+- **Predictions become the baseline**: After prediction, the predicted classes are treated as the "original" classifications for validation statistics.
+
+---
+
## Working with Images
### Image Cards
@@ -83,7 +142,7 @@ Some samples may have both manual annotations AND auto-classifications (e.g., yo
Each image card displays:
- The plankton image
-- ROI number
+- ROI number (in sample mode) or full image name (in class review mode)
- Classification score (if available)
- Original class (if relabeled)
@@ -152,20 +211,23 @@ The default scale is 3.4 pixels per micrometer (standard for IFCB). To adjust:
## Classification Sources
-`ClassiPyR` supports multiple classification input formats.
+`ClassiPyR` supports multiple classification input formats. When multiple formats exist for the same sample, the priority is: CSV > H5 > MAT. Samples without any pre-computed classifications can be classified on the fly using [Live Prediction](#live-prediction).
+
+All formats (including live predictions) support a **classification threshold** option (configurable in Settings under the Classification Folder). When enabled, predictions below the confidence threshold are shown as "unclassified"; when disabled, the raw (unthresholded) class prediction is used.
### CSV Files
-Standard classification CSV output. The CSV file must be named after the sample it describes (e.g., `D20230101T120000_IFCB134.csv`).
+Standard classification CSV output from [iRfcb](https://github.com/EuropeanIFCBGroup/iRfcb). The CSV file must be named after the sample it describes (e.g., `D20230101T120000_IFCB134.csv`).
Required columns (exact names):
- `file_name`: Image filename including `.png` extension (e.g., `D20230101T120000_IFCB134_00001.png`)
-- `class_name`: Predicted class name
+- `class_name`: Predicted class name (threshold-applied)
Optional columns:
- `score`: Classification confidence (0-1)
+- `class_name_auto`: Raw class prediction without threshold (used when threshold is disabled)
**Minimal example:**
@@ -175,28 +237,38 @@ D20230101T120000_IFCB134_00001.png,Diatom
D20230101T120000_IFCB134_00002.png,Ciliate
```
-**Example with confidence scores:**
+**Example with confidence scores and raw predictions:**
```
-file_name,class_name,score
-D20230101T120000_IFCB134_00001.png,Diatom,0.95
-D20230101T120000_IFCB134_00002.png,Ciliate,0.87
-D20230101T120000_IFCB134_00003.png,Dinoflagellate,0.72
+file_name,class_name,class_name_auto,score
+D20230101T120000_IFCB134_00001.png,unclassified,Diatom,0.45
+D20230101T120000_IFCB134_00002.png,Ciliate,Ciliate,0.87
```
**Different CNN pipelines**: If your classifier produces different column names, rename them to `file_name` and `class_name` before placing the CSV in the Classification Folder.
Files are looked up from the file index cache (see [File Index Cache](#file-index-cache) below).
+### HDF5 Classifier Output
+
+Files matching `*_class*.h5` pattern, produced by [iRfcb](https://github.com/EuropeanIFCBGroup/iRfcb) (>= 0.8.0). Contains:
+
+- `roi_numbers`: ROI identifiers
+- `class_name`: Predicted class with threshold applied
+- `class_name_auto`: Predicted class without threshold
+- `output_scores`: Per-class confidence scores
+- `class_labels`: All possible class names
+- `classifier_name`: Name of the classifier model
+
+Requires the optional [`hdf5r`](https://CRAN.R-project.org/package=hdf5r) package. Install with `install.packages("hdf5r")`.
+
### MATLAB Classifier Output
-Files matching `*_class*.mat` pattern containing:
+Files matching `*_class*.mat` pattern from [ifcb-analysis](https://github.com/hsosik/ifcb-analysis) containing:
- `roinum`: ROI numbers
-- `TBclass_above_threshold`: With threshold
-- `TBclass`: Without threshold
-
-**Threshold option**: Enable in Settings to include unclassified predictions below confidence threshold.
+- `TBclass_above_threshold`: Predicted class with threshold
+- `TBclass`: Predicted class without threshold
### Existing Annotations
@@ -296,12 +368,45 @@ Images organized into class folders for training CNN models or other classifiers
## Settings Reference
+### Data Source
+
+ClassiPyR supports two data source modes, selectable in Settings:
+
+| Mode | Description |
+|------|-------------|
+| **Local Folders** (default) | Read IFCB data from local ROI/ADC/HDR files |
+| **IFCB Dashboard** | Connect to a remote IFCB Dashboard instance |
+
+#### IFCB Dashboard Mode
+
+When "IFCB Dashboard" is selected, enter a Dashboard URL such as:
+
+- `https://habon-ifcb.whoi.edu/` — all datasets on the dashboard
+- `https://habon-ifcb.whoi.edu/timeline?dataset=tangosund` — a specific dataset
+
+The app fetches the sample list from the Dashboard API. When you load a sample, PNG images are downloaded from the dashboard and cached locally at `tools::R_user_dir("ClassiPyR", "cache")/dashboard/`. ADC files are downloaded on demand for image dimensions and MAT export.
+
+| Dashboard Setting | Description |
+|-------------------|-------------|
+| Dashboard URL | The full URL of the IFCB Dashboard (with optional `?dataset=` parameter) |
+| Use dashboard auto-classifications | When checked, downloads the dashboard's `_class_scores.csv` for validation mode |
+| Advanced Download Settings | Parallel downloads, sleep time, timeout, and max retries for dashboard downloads |
+
+The **Classification Folder** setting is available in both local and dashboard mode. In dashboard mode, local classification files (CSV/H5/MAT) take priority over dashboard auto-classifications. The priority order is:
+
+1. Database annotations (manual, existing)
+2. Local classification files (CSV > H5 > MAT) — if Classification Folder is configured
+3. Dashboard auto-classifications (if enabled)
+4. New annotation mode
+
+> **Note**: In dashboard mode, the ROI Data Folder setting is not used. The Output Folder, Database Folder, and PNG Output Folder still apply for saving annotations and exports.
+
### Folder Paths
| Setting | Description |
|---------|-------------|
-| Classification Folder | Source of CSV/MAT classifications |
-| ROI Data Folder | IFCB raw files (ROI/ADC/HDR) |
+| Classification Folder | Source of CSV/H5/MAT classifications (both local and dashboard mode) |
+| ROI Data Folder | IFCB raw files (ROI/ADC/HDR) (local mode only) |
| Output Folder | Where MAT files and statistics go (can be on a network drive) |
| Database Folder | Where the SQLite database is stored (must be a local drive) |
| PNG Output Folder | Where organized images go |
@@ -321,6 +426,9 @@ Below the format selector, two buttons allow bulk conversion between formats:
- **Import .mat → SQLite**: Imports all `.mat` annotation files from the output folder into the SQLite database. Already-imported samples are skipped.
- **Export SQLite → .mat**: Exports all annotated samples from the database to `.mat` files. Requires Python with scipy.
- **Export SQLite → PNG**: Extracts annotated images from ROI files into class-name subfolders in the PNG Output Folder. Useful for building training datasets for CNN classifiers.
+- **Import PNG → SQLite**: Imports annotations from a folder of PNG images organized in class-name subfolders. Class names are extracted from folder names (trailing `_NNN` suffixes are stripped). Useful for re-importing corrected exports or importing external classification datasets.
+
+> **Note**: When importing external datasets without corresponding ROI files, the annotations are stored in the database but images cannot be viewed or re-exported through the app. For full round-trip support, ensure ROI files are available in the configured ROI Data Folder.
### Auto-Sync
@@ -338,9 +446,22 @@ run_app(venv_path = "/path/to/your/venv")
The path is remembered for future sessions. **Priority order**: `run_app(venv_path=)` argument > saved settings > default (`./venv`).
+### Live Prediction
+
+| Setting | Description |
+|---------|-------------|
+| Gradio API URL | URL of a Gradio-hosted CNN classification server (e.g. `https://irfcb-classify.hf.space`) |
+| Prediction Model | CNN model to use for classification. Choices are fetched from the Gradio server. |
+
+When both fields are configured, a **Predict** button appears in the sidebar for Sample Mode. See [Live Prediction](#live-prediction) for usage details.
+
### Classifier Options
-**Apply classification threshold**: When loading MATLAB classifier output, use `TBclass_above_threshold` (checked) or `TBclass` (unchecked).
+**Apply classification threshold**: When loading classifier output (CSV, H5, or MAT) or live predictions, use the threshold-filtered class predictions (checked) or the raw unthresholded predictions (unchecked).
+
+### PNG Export Options
+
+**Skip class from PNG export**: Optionally exclude a class (e.g. "unclassified") from the organized PNG output. Set in Settings under the PNG Output Folder section.
---
@@ -409,4 +530,8 @@ Settings are loaded automatically when you start the app, so your folder paths,
- **[`iRfcb`](https://github.com/EuropeanIFCBGroup/iRfcb)** for IFCB data operations (extracting images, reading ADC metadata, reading/writing `.mat` files, class list handling)
- **[`RSQLite`](https://CRAN.R-project.org/package=RSQLite)** and **[`DBI`](https://CRAN.R-project.org/package=DBI)** for the SQLite annotation database
+Optional dependencies:
+
+- **[`hdf5r`](https://CRAN.R-project.org/package=hdf5r)** for reading HDF5 (`.h5`) classifier output files. Install with `install.packages("hdf5r")`.
+
All R dependencies are installed automatically when you install `ClassiPyR`. Python is only needed for `.mat` file export.
 +
+ -
- +
+ -
- +
+
+
+