Documentation
Powerful tools for quantitative compositional mapping and analysis
Official documentation for XMapTools, a free and versatile software solution for quantitative compositional mapping and analysis. This site is built with VitePress and deployed via GitHub Pages.
Live site: https://xmaptools.github.io/Documentation/
Contributions to improve or extend the documentation are welcome. Please follow the workflow described below.
git clone https://github.com/<your-username>/Documentation.git
+cd Documentationnpm installnpm run devhttp://localhost:5173/Documentation/. Changes to .md files are reflected live in the browser..
+├── index.md # Home page
+├── getting-started.md # Getting started guide
+├── installation_guide.md # Installation instructions
+├── doc-epma.md # EPMA documentation
+├── doc-laicpms.md # LA-ICP-MS documentation
+├── bingo-antidote.md # Bingo-Antidote documentation
+├── tutorials.md # Tutorials
+├── videos.md # Video resources
+├── .vitepress/ # VitePress configuration and theme
+│ ├── config.js # Site configuration (nav, sidebar, ...)
+│ └── theme/ # Custom theme overrides
+└── package.json.md files) at the root of the repository.<img> tag:<img src="https://raw.githubusercontent.com/xmaptools/Documentation_Images/main/Figures/my-image.png" alt="description" style="max-width: 100%; height: auto; display: block; margin: 0 auto;">git checkout -b my-contributionnpm run dev.git add .
+git commit -m "docs: describe your changes"git push origin my-contributionmain branch of this repository.##, ###) to structure content logically.installation_guide.md).npm run buildThe static site is generated in .vitepress/dist/ and automatically deployed to GitHub Pages via the CI workflow on push to main.
This project is licensed under the GNU General Public License v3.0.
`,24)])])}const u=i(n,[["render",l]]);export{g as __pageData,u as default}; diff --git a/.vitepress/dist/assets/README.md.BF1pv8wB.lean.js b/.vitepress/dist/assets/README.md.BF1pv8wB.lean.js new file mode 100644 index 0000000..1bf2c6e --- /dev/null +++ b/.vitepress/dist/assets/README.md.BF1pv8wB.lean.js @@ -0,0 +1 @@ +import{_ as i,o as a,c as e,ae as t}from"./chunks/framework.DIXg7aqb.js";const g=JSON.parse('{"title":"XMapTools Documentation","description":"","frontmatter":{},"headers":[],"relativePath":"README.md","filePath":"README.md"}'),n={name:"README.md"};function l(o,s,r,h,p,d){return a(),e("div",null,[...s[0]||(s[0]=[t("",24)])])}const u=i(n,[["render",l]]);export{g as __pageData,u as default}; diff --git a/.vitepress/dist/assets/app.CLHRpw3L.js b/.vitepress/dist/assets/app.CLHRpw3L.js new file mode 100644 index 0000000..e45fe67 --- /dev/null +++ b/.vitepress/dist/assets/app.CLHRpw3L.js @@ -0,0 +1 @@ +import{R as p}from"./chunks/theme.BZmAIviK.js";import{R as s,a0 as i,a1 as u,a2 as c,a3 as l,a4 as f,a5 as d,a6 as m,a7 as h,a8 as g,a9 as A,d as v,u as R,v as w,s as y,aa as C,ab as P,ac as b,ad as E}from"./chunks/framework.DIXg7aqb.js";function r(e){if(e.extends){const a=r(e.extends);return{...a,...e,async enhanceApp(t){a.enhanceApp&&await a.enhanceApp(t),e.enhanceApp&&await e.enhanceApp(t)}}}return e}const n=r(p),S=v({name:"VitePressApp",setup(){const{site:e,lang:a,dir:t}=R();return w(()=>{y(()=>{document.documentElement.lang=a.value,document.documentElement.dir=t.value})}),e.value.router.prefetchLinks&&C(),P(),b(),n.setup&&n.setup(),()=>E(n.Layout)}});async function T(){globalThis.__VITEPRESS__=!0;const e=_(),a=D();a.provide(u,e);const t=c(e.route);return a.provide(l,t),a.component("Content",f),a.component("ClientOnly",d),Object.defineProperties(a.config.globalProperties,{$frontmatter:{get(){return t.frontmatter.value}},$params:{get(){return t.page.value.params}}}),n.enhanceApp&&await n.enhanceApp({app:a,router:e,siteData:m}),{app:a,router:e,data:t}}function D(){return A(S)}function _(){let e=s;return h(a=>{let t=g(a),o=null;return t&&(e&&(t=t.replace(/\.js$/,".lean.js")),o=import(t)),s&&(e=!1),o},n.NotFound)}s&&T().then(({app:e,router:a,data:t})=>{a.go().then(()=>{i(a.route,t.site),e.mount("#app")})});export{T as createApp}; diff --git a/.vitepress/dist/assets/bingo-antidote.md.CS1jHsQP.js b/.vitepress/dist/assets/bingo-antidote.md.CS1jHsQP.js new file mode 100644 index 0000000..76a3036 --- /dev/null +++ b/.vitepress/dist/assets/bingo-antidote.md.CS1jHsQP.js @@ -0,0 +1 @@ +import{_ as t,o as a,c as o,ae as i}from"./chunks/framework.DIXg7aqb.js";const p=JSON.parse('{"title":"Bingo-Antidote","description":"","frontmatter":{},"headers":[],"relativePath":"bingo-antidote.md","filePath":"bingo-antidote.md"}'),n={name:"bingo-antidote.md"};function r(l,e,s,d,h,m){return a(),o("div",null,[...e[0]||(e[0]=[i('
Bingo-Antidote is a petrological software originally developed by Pierre Lanari and Erik Duesterhoeft that offers an alternative modelling strategy based on iterative thermodynamic models integrated with quantitative compositional mapping. This second version, distributed as an XMapTools add-on, comes with a redesigned graphical user interface and improved features.
Bingo-Antidote proposes a hybrid strategy combining the strengths of Gibbs energy minimisation (GEM) and inverse thermobarometry models, based on the comparison between modelled and observed mineral assemblages, modes and compositions. The overall technique relies on quantitative compositional maps obtained by electron probe microanalysis to provide a mutually consistent set of observed data such as bulk rock and mineral compositions.
Bingo-Antidote uses the Gibbs energy minimiser Theriak developed by Christian de Capitani. The latest version of Theriak-Domino, provided by Doug Tinkham, is required to use Bingo-Antidote.
Download Theriak-Domino from Github

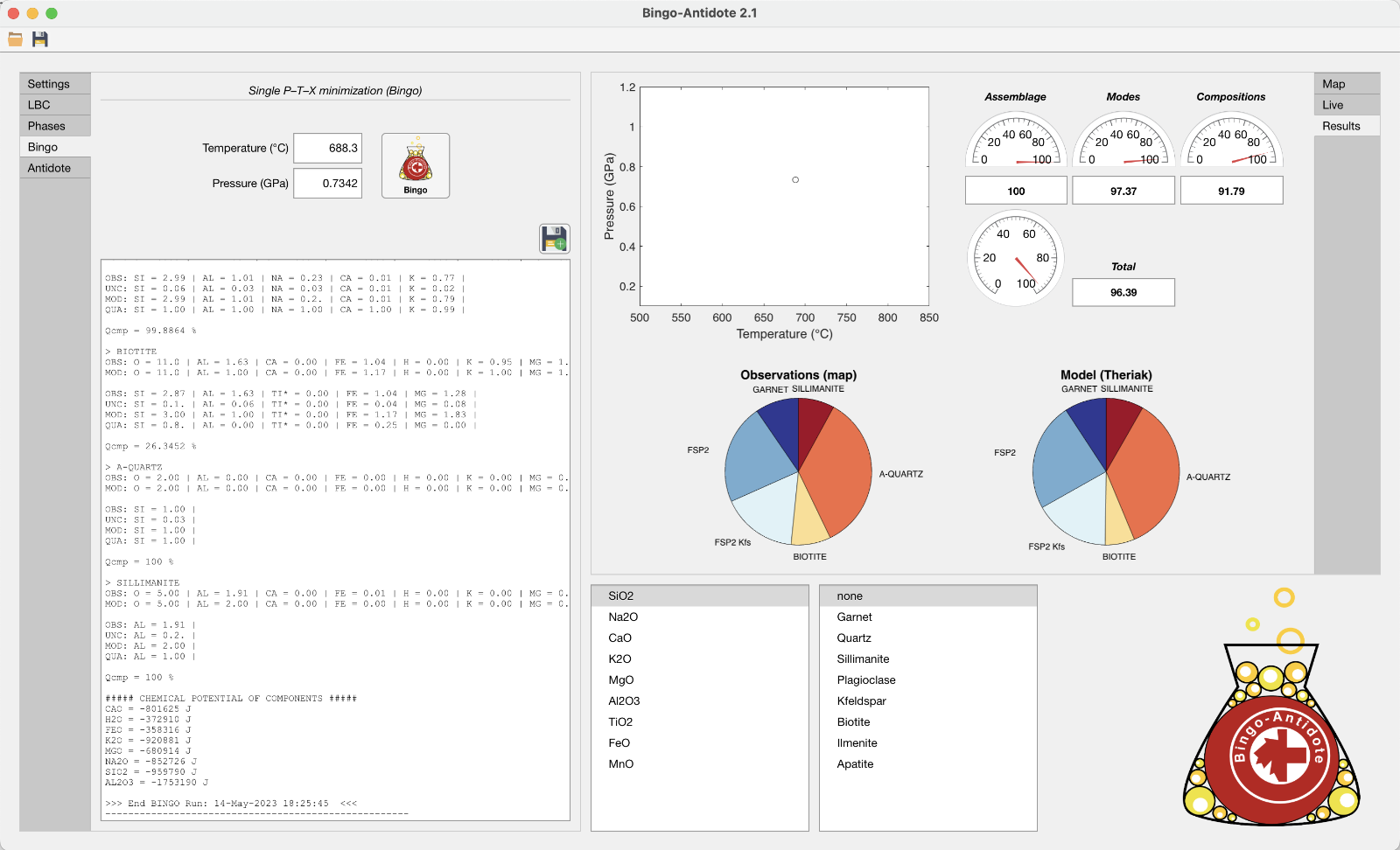
There are two technical papers, the first by Duesterhoeft & Lanari (2020) and the second by Lanari & Hermann (2021), which includes a proof of concept and an application example.
The philosophy of iterative thermodynamic modelling behind Bingo-Antidote is also described in older publications: Lanari & Engi (2017) and Lanari & Duesterhoeft (2019). Note that the Lanari & Duesterhoeft (2019) paper gives a general overview of thermodynamic modelling and also provides an example of an application of Bingo-Antidote.
This short tutorial on Bingo-Antidote is from the Goldschmidt Workshop, which took place in Lyon on 8–9 July 2023 and was entitled 'Modern Applications of Phase Equilibrium Modelling'.
Based on XMapTools 4.5 embedded documentation – Help file version 09.06.2026
This page describes the tools available in the Add-ons section. Contact Pierre Lanari if you want to develop an add-on for XMapTools.
This module allows XMapTools data to be exported. It is available for both Map Data and Spot Data.
The export format is defined by the user and can be used to export data for a specific software (e.g. MinPlot). Select the Data source, a maskfile and the type of data to be exported (e.g. Median, Mean or random selection, see Fig. 1).
The BRC correction is a correction from XMapTools 4.3 (Lanari et al. 2019) that eliminates mixing pixels using the maskfile data.
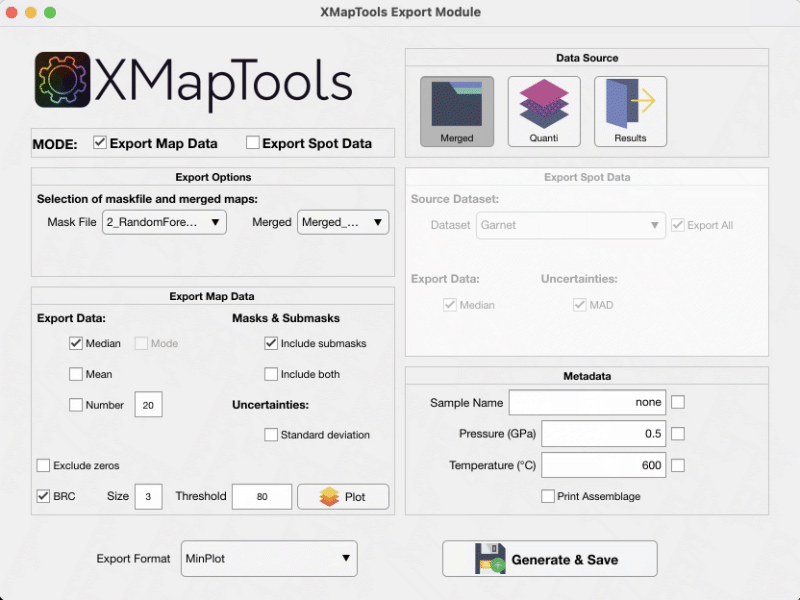
Figure 1: Export module for Map Data in XMapTools 4.5.
Spot data can be exported using the Export module available in the Add-on section. Open the module and select the Export Spot Data mode. Select your data source (e.g. Quanti in Fig. 2) and press the Generate & Save button.
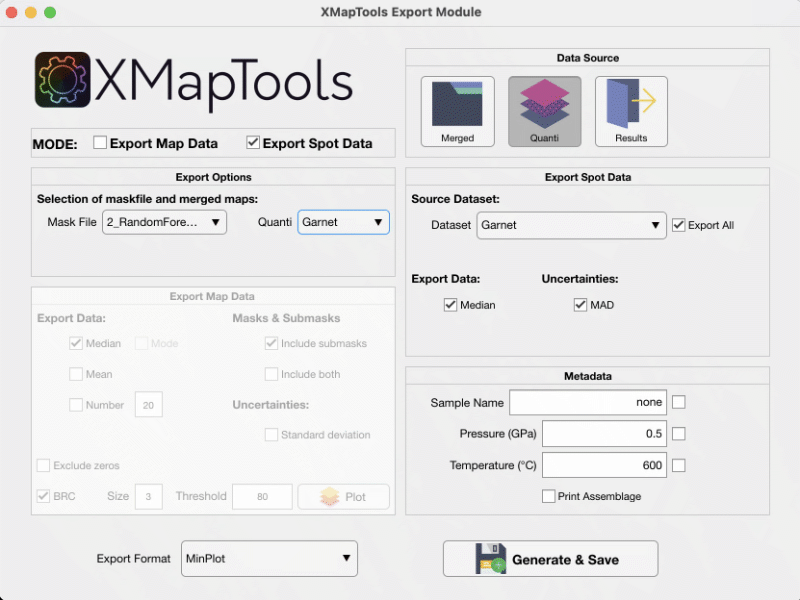
Figure 2: Export module for Spot Data in XMapTools 4.5.
The image converter module allows images to be converted to numeric data. Press the Load Image button to load an image. The image can be cropped by using the pink ROI and clicking the Crop Image button. The min and max values can be set manually (e.g. 0 and 100 in Fig. 3). Press the Save Map button to save the converted data into a txt file.
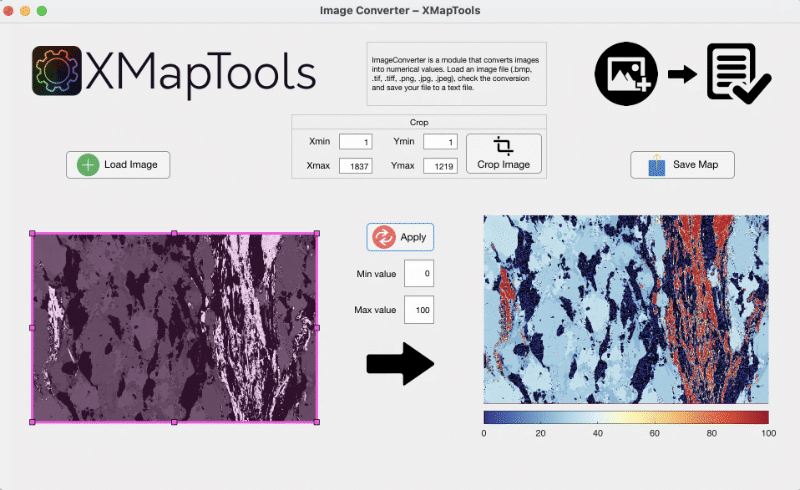
Figure 3: Image Converter module.
Bingo-Antidote is a petrological software originally developed by Pierre Lanari and Erik Duesterhoeft that offers an alternative modelling strategy based on iterative thermodynamic models integrated with quantitative compositional mapping. Bingo-Antidote 2.0 is distributed as an add-on to XMapTools. It comes with a redesigned graphical user interface and improved functionality.
Visit https://xmaptools.ch/bingo-antidote/ to find out more. See also the dedicated Bingo-Antidote documentation page.
',23)])])}const g=t(i,[["render",r]]);export{h as __pageData,g as default}; diff --git a/.vitepress/dist/assets/doc-addons.md.BJFpqniu.lean.js b/.vitepress/dist/assets/doc-addons.md.BJFpqniu.lean.js new file mode 100644 index 0000000..408cfa0 --- /dev/null +++ b/.vitepress/dist/assets/doc-addons.md.BJFpqniu.lean.js @@ -0,0 +1 @@ +import{_ as t,o as a,c as o,ae as n}from"./chunks/framework.DIXg7aqb.js";const h=JSON.parse('{"title":"Add-ons","description":"","frontmatter":{},"headers":[],"relativePath":"doc-addons.md","filePath":"doc-addons.md"}'),i={name:"doc-addons.md"};function r(d,e,s,p,l,m){return a(),o("div",null,[...e[0]||(e[0]=[n("",23)])])}const g=t(i,[["render",r]]);export{h as __pageData,g as default}; diff --git a/.vitepress/dist/assets/doc-epma.md.DFglyrsL.js b/.vitepress/dist/assets/doc-epma.md.DFglyrsL.js new file mode 100644 index 0000000..4431520 --- /dev/null +++ b/.vitepress/dist/assets/doc-epma.md.DFglyrsL.js @@ -0,0 +1,166 @@ +import{_ as a,o as e,c as n,ae as s}from"./chunks/framework.DIXg7aqb.js";const h=JSON.parse('{"title":"XMapTools documentation for EPMA","description":"","frontmatter":{},"headers":[],"relativePath":"doc-epma.md","filePath":"doc-epma.md"}'),i={name:"doc-epma.md"};function o(l,t,p,r,d,c){return e(),n("div",null,[...t[0]||(t[0]=[s(`EPMA & SEM:
Because of the file format used by CAMECA, XMapTools is reading the information of the following lines in the header:
Therefore the minimum number of header lines is:
Start : X = 15702 Y = -24236 Z = 240
+Step Number : 416
+Line Number : 440
+
+Y Unit : cts
+Image : Ca Ka
+
+[Data table should start here... use tabulation, no coma]XMapTools can import two types of map data file that are generated by CAMECA microprobes.
FileName : Bustamante 18-1-2013.impDat (Current dataset : 4)
+Signal(s) Used : Vs1 BSE Z, Na Ka, Ca Ka, Fe Ka, Mg Ka, Al Ka, Si Ka Eds, S Ka Eds, Cl Ka Eds, K Ka Eds, Ti Ka Eds, Mn Ka Eds, Zr La Eds, Ba La Eds
+Spectromers Conditions : Sp1 LTAP, Sp2 LPET, Sp3 LLIF, Sp4 LTAP, Sp5 LTAP, Eds 1, Eds 2, Eds 3, Eds 4, Eds 5, Eds 6, Eds 7, Eds 8
+Full Spectromers Conditions : Sp1 LTAP(2d= 25.745,K= 0.00218), Sp2 LPET(2d= 8.75,K= 0.000144), Sp3 LLIF(2d= 4.0267,K= 0.000058), Sp4 LTAP(2d= 25.745,K= 0.00218), Sp5 LTAP(2d= 25.745,K= 0.00218), Eds 1, Eds 2, Eds 3, Eds 4, Eds 5, Eds 6, Eds 7, Eds 8
+Column Conditions : Cond 1 : 15.1keV 302.5757nA
+Date : 19-Jan-2013
+User Name : sx
+Setup Name : Bustamante.impSet
+DataSet Comment :
+Comment :
+Analysis Date : 1/18/2013 12:42:21 PM
+Project Name : Caetano
+Sample Name : 2013-01-18
+Pha Parameters :
+ Bias Gain Dtime Blin Wind Mode
+ Sp1(Na Ka) 1300 2635 3 560 Inte
+ Sp2(Ca Ka) 1289 1100 3 560 Inte
+ Sp3(Fe Ka) 1824 550 3 560 Inte
+ Sp4(Mg Ka) 1293 2964 3 560 Inte
+ Sp5(Al Ka) 1280 2943 3 560 Inte
+ EDS: Si Ka
+ EDS: S Ka
+ EDS: Cl Ka
+ EDS: K Ka
+ EDS: Ti Ka
+ EDS: Mn Ka
+ EDS: Zr La
+ EDS: Ba La
+Peak Position : Sp1 46363, Sp2 38388, Sp3 48084, Sp4 38500, Sp5 32460
+Start : X = 7386 Y = -17430 Z = 49
+Stop :
+Dwell Time : 0.03 Sec
+Acquisition type : Stage Grid
+Step Number : 1001
+Line Number : 614
+Beam Size : 3 µm
+Step Size : 4.
+
+Y Unit :
+Image : BSE Z
+
+83.0000 83.0000 88.0000 82.0000 82.0000 81.0000 82.0000 83.0000 83.0000 84.0000 89.0000 92.0000 83.0000 87.0000 83.0000 85.0000 86.0000 86.0000 85.0000 86.0000 88.0000 85.0000 82.0000 87.0000 85.0000 84.0000 87.0000 87.0000 85.0000 85.0000 85.0000 87.0000 86.0000 86.0000 85.0000 86.0000 90.0000 87.0000 87.0000 87.0000 86.0000 96.0000 107.0000 89.0000 88.0000 90.0000 101.0000 113.0000 92.0000 101.0000 81.0000 96.0000 79.0000 81.0000 91.0000 82.0000 128.0000 135.0000 126.0000 120.0000 94.0000 112.0000 84.0000 80.0000 82.0000 83.0000 88.0000 97.0000 82.0000 81.0000 80.0000 81.0000 82.0000 81.0000 131.0000 97.0000 82.0000 83.0000 84.0000 84.0000 80.0000 80.0000 81.0000 87.0000 96.0000 83.0000 84.0000 85.0000 79.0000 82.0000 81.0000 80.0000 81.0000 80.0000 80.0000 79.0000 80.0000 83.0000 82.0000 82.0000 82.0000 82.0000 80.0000 80.0000 86.0000 82.0000 86.0000 85.0000 81.0000 84.0000 83.0000 84.0000 85.0000 83.0000 82.0000 79.0000 81.0000 80.0000 83.0000 84.0000 84.0000 83.0000 84.0000 84.0000 80.0000 80.0000 80.0000 81.0000 85.0000 85.0000 85.0000 83.0000 84.0000 82.0000 83.0000 87.0000 87.0000 84.0000 80.0000 81.0000 85.0000 85.0000 84.0000 83.0000 85.0000 92.0000 81.0000 83.0000 85.0000 85.0000 84.0000 84.0000 84.0000 85.0000 87.0000 87.0000 85.0000 84.0000 84.0000 81.0000 85.0000 80.0000 84.0000 83.0000 84.0000 83.0000 89.0000 91.0000 88.0000 85.0000 89.0000 91.0000 95.0000 91.0000 96.0000 82.0000 84.0000 83.0000 82.0000 86.0000 82.0000 79.0000 86.0000 84.0000 83.0000 83.0000 83.0000 84.0000 83.0000 83.0000 84.0000 84.0000 80.0000 86.0000 86.0000 83.0000 85.0000 83.0000 83.0000 82.0000 91.0000 79.0000 80.0000 76.0000 82.0000 81.0000 81.0000 81.0000 81.0000 81.0000 80.0000 81.0000 83.0000 85.0000 85.0000 86.0000 87.0000 81.0000 83.0000 83.0000 79.0000 80.0000 81.0000 78.0000 84.0000 83.0000 83.0000 76.0000 78.0000 81.0000 80.0000 80.0000 81.0000 80.0000 81.0000 75.0000 81.0000 80.0000 86.0000 115.0000 78.0000 80.0000 83.0000 79.0000 104.0000 102.0000 94.0000 90.0000 84.0000 84.0000 82.0000 84.0000 82.0000 81.0000 81.0000 82.0000 82.0000 83.0000 83.0000 83.0000 83.0000 80.0000 87.0000 98.0000 85.0000 79.0000 82.0000 82.0000 82.0000 83.0000 79.0000 100.0000 115.0000 80.0000 80.0000 80.0000 81.0000 90.0000 75.0000 82.0000 80.0000 79.0000 78.0000 84.0000 81.0000 81.0000 111.0000 107.0000 83.0000 85.0000 83.0000 82.0000 111.0000 84.0000 80.0000 83.0000 70.0000 80.0000 82.0000 83.0000 82.0000 82.0000 81.0000 92.0000 83.0000 86.0000 85.0000 102.0000 101.0000 79.0000 87.0000 84.0000 84.0000 82.0000 86.0000 95.0000 124.0000 84.0000 82.0000 80.0000 85.0000 85.0000 80.0000 81.0000 88.0000 92.0000 88.0000 86.0000 76.0000 83.0000 86.0000 82.0000 82.0000 83.0000 80.0000 80.0000 79.0000 80.0000 80.0000 78.0000 82.0000 88.0000 89.0000 91.0000 88.0000 86.0000 92.0000 97.0000 81.0000 78.0000 79.0000 79.0000 82.0000 82.0000 80.0000 77.0000 80.0000 79.0000 79.0000 80.0000 87.0000 85.0000 85.0000 85.0000 84.0000 83.0000 80.0000 80.0000 79.0000 79.0000 78.0000 84.0000 82.0000 81.0000 80.0000 78.0000 79.0000 80.0000 83.0000 79.0000 81.0000 80.0000 79.0000 77.0000 77.0000 78.0000 85.0000 85.0000 82.0000 85.0000 79.0000 80.0000 80.0000 79.0000 79.0000 79.0000 79.0000 80.0000 85.0000 84.0000 82.0000 80.0000 80.0000 79.0000 79.0000 80.0000 79.0000 80.0000 80.0000 79.0000 80.0000 80.0000 80.0000 82.0000 85.0000 85.0000 85.0000 85.0000 87.0000 80.0000 80.0000 80.0000 84.0000 81.0000 81.0000 94.0000 81.0000 81.0000 81.0000 89.0000 81.0000 80.0000 79.0000 80.0000 85.0000 85.0000 83.0000 84.0000 85.0000 80.0000 80.0000 83.0000 85.0000 88.0000 86.0000 80.0000 80.0000 80.0000 80.0000 80.0000 80.0000 80.0000 81.0000 82.0000 80.0000 83.0000 84.0000 122.0000 99.0000 76.0000 80.0000 82.0000 83.0000 82.0000 86.0000 80.0000 75.0000 91.0000 87.0000 61.0000 74.0000 59.0000 82.0000 94.0000 85.0000 105.0000 87.0000 52.0000 82.0000 88.0000 84.0000 79.0000 80.0000 80.0000 79.0000 80.0000 79.0000 79.0000 81.0000 85.0000 84.0000 84.0000 85.0000 86.0000 85.0000 85.0000 79.0000 80.0000 79.0000 83.0000 80.0000 79.0000 79.0000 79.0000 89.0000 78.0000 80.0000 80.0000 79.0000 79.0000 80.0000 80.0000 81.0000 85.0000 79.0000 78.0000 85.0000 82.0000 78.0000 79.0000 79.0000 78.0000 80.0000 78.0000 78.0000 78.0000 78.0000 79.0000 78.0000 78.0000 78.0000 78.0000 79.0000 79.0000 78.0000 84.0000 85.0000 84.0000 79.0000 79.0000 79.0000 88.0000 83.0000 79.0000 86.0000 98.0000 85.0000 81.0000 80.0000 79.0000 83.0000 79.0000 79.0000 83.0000 85.0000 84.0000 77.0000 77.0000 80.0000 81.0000 79.0000 81.0000 78.0000 78.0000 78.0000 79.0000 78.0000 78.0000 79.0000 79.0000 79.0000 79.0000 86.0000 84.0000 80.0000 79.0000 79.0000 82.0000 80.0000 79.0000 79.0000 78.0000 78.0000 79.0000 79.0000 78.0000 78.0000 79.0000 80.0000 78.0000 85.0000 77.0000 76.0000 78.0000 78.0000 77.0000 76.0000 79.0000 78.0000 79.0000 78.0000 79.0000 84.0000 85.0000 84.0000 85.0000 84.0000 81.0000 85.0000 84.0000 78.0000 78.0000 79.0000 79.0000 78.0000 79.0000 80.0000 80.0000 112.0000 86.0000 88.0000 86.0000 86.0000 84.0000 83.0000 83.0000 84.0000 83.0000 84.0000 76.0000 79.0000 79.0000 78.0000 78.0000 79.0000 79.0000 79.0000 79.0000 79.0000 79.0000 82.0000 89.0000 88.0000 85.0000 85.0000 86.0000 85.0000 84.0000 85.0000 83.0000 85.0000 85.0000 83.0000 89.0000 86.0000 84.0000 86.0000 87.0000 85.0000 86.0000 85.0000 84.0000 87.0000 88.0000 88.0000 85.0000 80.0000 80.0000 80.0000 79.0000 78.0000 79.0000 79.0000 79.0000 80.0000 79.0000 81.0000 86.0000 85.0000 86.0000 86.0000 84.0000 85.0000 84.0000 84.0000 84.0000 91.0000 85.0000 84.0000 83.0000 83.0000 86.0000 88.0000 89.0000 88.0000 86.0000 85.0000 83.0000 84.0000 84.0000 84.0000 84.0000 83.0000 89.0000 79.0000 78.0000 84.0000 81.0000 84.0000 84.0000 83.0000 80.0000 86.0000 83.0000 81.0000 81.0000 80.0000 78.0000 78.0000 81.0000 78.0000 79.0000 123.0000 107.0000 82.0000 83.0000 78.0000 76.0000 78.0000 77.0000 78.0000 78.0000 81.0000 80.0000 75.0000 77.0000 78.0000 78.0000 84.0000 78.0000 79.0000 79.0000 85.0000 85.0000 84.0000 82.0000 84.0000 85.0000 81.0000 85.0000 83.0000 84.0000 84.0000 84.0000 84.0000 84.0000 85.0000 84.0000 85.0000 84.0000 84.0000 84.0000 85.0000 85.0000 90.0000 86.0000 92.0000 86.0000 85.0000 85.0000 85.0000 86.0000 106.0000 84.0000 83.0000 85.0000 85.0000 84.0000 87.0000 108.0000 84.0000 83.0000 84.0000 84.0000 84.0000 88.0000 83.0000 83.0000 83.0000 84.0000 83.0000 85.0000 94.0000 84.0000 85.0000 85.0000 79.0000 78.0000 78.0000 78.0000 78.0000 79.0000 79.0000 79.0000 79.0000 81.0000 86.0000 117.0000 85.0000 84.0000 85.0000 85.0000 83.0000 105.0000 86.0000 86.0000 89.0000 89.0000 88.0000 89.0000 90.0000 84.0000 84.0000 83.0000 77.0000 76.0000 80.0000 82.0000 79.0000 82.0000 88.0000 82.0000 84.0000 83.0000 92.0000 88.0000 78.0000 78.0000 77.0000 78.0000 78.0000 77.0000 80.0000 80.0000 77.0000 77.0000 77.0000 77.0000 77.0000 82.0000 96.0000 83.0000 83.0000 77.0000 77.0000 77.0000 78.0000 77.0000 78.0000 78.0000 78.0000 84.0000 83.0000 84.0000 84.0000 84.0000 84.0000 83.0000 84.0000 84.0000 83.0000 83.0000 84.0000 82.0000 82.0000 83.0000 83.0000 87.0000 82.0000 77.0000 85.0000 76.0000 77.0000 77.0000 78.0000 78.0000 77.0000 77.0000 78.0000 83.0000 84.0000 84.0000 85.0000 82.0000 82.0000 83.0000 82.0000 82.0000 82.0000 83.0000 86.0000 81.0000 82.0000 84.0000 83.0000 83.0000 83.0000 83.0000 77.0000 77.0000 78.0000 77.0000 76.0000 77.0000 79.0000 80.0000 90.0000 83.0000 82.0000 82.0000 82.0000 82.0000 83.0000 85.0000 87.0000 84.0000 82.0000 90.0000 83.0000 89.0000 82.0000 81.0000 83.0000 83.0000 83.0000 83.0000 83.0000 82.0000 80.0000 77.0000 83.0000 82.0000 82.0000 82.0000 83.0000 82.0000 82.0000 82.0000 78.0000 77.0000 77.0000 77.0000 77.0000 77.0000 77.0000 78.0000 78.0000 88.0000 82.0000 82.0000 83.0000 83.0000 84.0000 83.0000 77.0000 77.0000 77.0000 77.0000 77.0000 77.0000 77.0000 82.0000 76.0000 77.0000 79.0000 76.0000 77.0000 78.0000 76.0000 76.0000 77.0000 76.0000 77.0000 77.0000 74.0000 74.0000 77.0000 77.0000 77.0000 77.0000 76.0000 76.0000 77.0000 77.0000 77.0000 76.0000 77.0000 77.0000 76.0000 77.0000 78.0000
+82.0000 83.0000 82.0000 80.0000 79.0000 80.0000 84.0000 81.0000 81.0000 82.0000 85.0000 88.0000 86.0000 86.0000 84.0000 83.0000 83.0000 84.0000 83.0000 84.0000 83.0000 85.0000 85.0000 87.0000 86.0000 84.0000 91.0000 91.0000 85.0000 85.0000 85.0000 86.0000 87.0000 89.0000 91.0000 87.0000 94.0000 85.0000 86.0000 90.0000 128.0000 95.0000 110.0000 115.0000 105.0000 105.0000 93.0000 95.0000 102.0000 78.0000 87.0000 85.0000 80.0000 80.0000 88.0000 80.0000 84.0000 128.0000 112.0000 113.0000 102.0000 126.0000 80.0000 80.0000 80.0000 83.0000 85.0000 82.0000 77.0000 79.0000 79.0000 81.0000 81.0000 81.0000 97.0000 90.0000 81.0000 82.0000 83.0000 82.0000 79.0000 82.0000 82.0000 84.0000 87.0000 82.0000 82.0000 84.0000 77.0000 85.0000 80.0000 80.0000 79.0000 80.0000 79.0000 79.0000 82.0000 82.0000 82.0000 81.0000 81.0000 81.0000 81.0000 81.0000 84.0000 82.0000 82.0000 79.0000 79.0000 81.0000 83.0000 83.0000 84.0000 82.0000 78.0000 78.0000 81.0000 80.0000 84.0000 83.0000 83.0000 82.0000 83.0000 81.0000 79.0000 79.0000 79.0000 79.0000 83.0000 83.0000 83.0000 85.0000 85.0000 80.0000 82.0000 84.0000 84.0000 82.0000 81.0000 83.0000 82.0000 83.0000 83.0000 80.0000 83.0000 82.0000 85.0000 83.0000 79.0000 82.0000 83.0000 83.0000 82.0000 85.0000 86.0000 87.0000 84.0000 84.0000 82.0000 82.0000 83.0000 82.0000 81.0000 83.0000 84.0000 84.0000 89.0000 90.0000 81.0000 87.0000 90.0000 89.0000 90.0000 94.0000 89.0000 81.0000 82.0000 82.0000 80.0000 80.0000 84.0000 79.0000 83.0000 82.0000 82.0000 82.0000 82.0000 83.0000 81.0000 95.0000 84.0000 82.0000 83.0000 86.0000 92.0000 84.0000 81.0000 82.0000 82.0000 81.0000 88.0000 78.0000 80.0000 79.0000 81.0000 80.0000 80.0000 80.0000 81.0000 80.0000 80.0000 82.0000 82.0000 79.0000 82.0000 78.0000 83.0000 80.0000 82.0000 83.0000 78.0000 81.0000 80.0000 83.0000 82.0000 82.0000 84.0000 76.0000 78.0000 79.0000 79.0000 80.0000 81.0000 79.0000 80.0000 80.0000 79.0000 80.0000 82.0000 123.0000 88.0000 78.0000 81.0000 75.0000 89.0000 84.0000 90.0000 87.0000 82.0000 160.0000 105.0000 97.0000 82.0000 79.0000 82.0000 81.0000 82.0000 81.0000 82.0000 81.0000 82.0000 75.0000 89.0000 85.0000 83.0000 79.0000 80.0000 81.0000 80.0000 81.0000 82.0000 105.0000 93.0000 79.0000 80.0000 80.0000 80.0000 106.0000 81.0000 77.0000 79.0000 78.0000 83.0000 82.0000 80.0000 82.0000 125.0000 84.0000 83.0000 81.0000 87.0000 81.0000 73.0000 82.0000 82.0000 83.0000 91.0000 81.0000 80.0000 80.0000 81.0000 81.0000 81.0000 87.0000 82.0000 84.0000 82.0000 92.0000 79.0000 78.0000 83.0000 84.0000 82.0000 81.0000 82.0000 87.0000 92.0000 79.0000 79.0000 81.0000 87.0000 79.0000 83.0000 83.0000 86.0000 89.0000 91.0000 79.0000 78.0000 90.0000 78.0000 77.0000 78.0000 83.0000 80.0000 79.0000 79.0000 79.0000 84.0000 79.0000 80.0000 81.0000 83.0000 87.0000 82.0000 79.0000 80.0000 89.0000 81.0000 77.0000 77.0000 80.0000 79.0000 78.0000 80.0000 80.0000 80.0000 78.0000 76.0000 79.0000 82.0000 84.0000 85.0000 84.0000 83.0000 78.0000 78.0000 79.0000 79.0000 79.0000 80.0000 79.0000 78.0000 80.0000 76.0000 77.0000 78.0000 79.0000 79.0000 91.0000 81.0000 79.0000 81.0000 81.0000 82.0000 82.0000 84.0000 84.0000 81.0000 79.0000 82.0000 77.0000 80.0000 80.0000 82.0000 78.0000 78.0000 78.0000 92.0000 81.0000 85.0000 78.0000 84.0000 77.0000 79.0000 79.0000 78.0000 79.0000 79.0000 78.0000 79.0000 86.0000 79.0000 86.0000 84.0000 84.0000 85.0000 85.0000 90.0000 80.0000 79.0000 79.0000 83.0000 80.0000 80.0000 84.0000 79.0000 80.0000 131.0000 91.0000 83.0000 79.0000 80.0000 82.0000 82.0000 85.0000 80.0000 80.0000 80.0000 79.0000 80.0000 81.0000 84.0000 88.0000 86.0000 80.0000 80.0000 80.0000 80.0000 80.0000 79.0000 80.0000 80.0000 81.0000 81.0000 83.0000 86.0000 103.0000 86.0000 82.0000 81.0000 85.0000 88.0000 85.0000 83.0000 86.0000 59.0000 101.0000 91.0000 57.0000 58.0000 66.0000 61.0000 60.0000 102.0000 87.0000 80.0000 56.0000 90.0000 92.0000 85.0000 81.0000 80.0000 80.0000 81.0000 81.0000 79.0000 78.0000 79.0000 86.0000 84.0000 83.0000 83.0000 84.0000 85.0000 84.0000 78.0000 78.0000 78.0000 85.0000 84.0000 79.0000 78.0000 78.0000 79.0000 78.0000 79.0000 79.0000 78.0000 79.0000 78.0000 79.0000 79.0000 84.0000 85.0000 78.0000 84.0000 84.0000 77.0000 78.0000 78.0000 78.0000 76.0000 75.0000 74.0000 76.0000 78.0000 78.0000 77.0000 75.0000 75.0000 81.0000 78.0000 78.0000 77.0000 83.0000 82.0000 82.0000 77.0000 78.0000 77.0000 83.0000 77.0000 78.0000 84.0000 104.0000 84.0000 75.0000 76.0000 78.0000 82.0000 78.0000 79.0000 83.0000 83.0000 84.0000 78.0000 78.0000 78.0000 75.0000 76.0000 76.0000 77.0000 77.0000 77.0000 77.0000 76.0000 77.0000 78.0000 78.0000 80.0000 85.0000 83.0000 84.0000 79.0000 78.0000 78.0000 75.0000 79.0000 78.0000 78.0000 77.0000 77.0000 78.0000 78.0000 78.0000 77.0000 78.0000 77.0000 77.0000 81.0000 77.0000 75.0000 78.0000 78.0000 77.0000 80.0000 79.0000 78.0000 78.0000 77.0000 80.0000 83.0000 84.0000 84.0000 83.0000 105.0000 84.0000 89.0000 83.0000 76.0000 77.0000 77.0000 78.0000 77.0000 79.0000 79.0000 79.0000 146.0000 87.0000 89.0000 84.0000 85.0000 83.0000 83.0000 82.0000 82.0000 83.0000 81.0000 78.0000 76.0000 78.0000 78.0000 77.0000 78.0000 78.0000 78.0000 78.0000 79.0000 79.0000 81.0000 81.0000 82.0000 83.0000 85.0000 85.0000 82.0000 84.0000 84.0000 84.0000 84.0000 85.0000 83.0000 83.0000 80.0000 86.0000 90.0000 88.0000 82.0000 85.0000 85.0000 85.0000 86.0000 85.0000 84.0000 84.0000 82.0000 79.0000 78.0000 78.0000 78.0000 78.0000 78.0000 79.0000 79.0000 85.0000 81.0000 85.0000 85.0000 85.0000 85.0000 83.0000 85.0000 84.0000 83.0000 84.0000 83.0000 84.0000 83.0000 83.0000 84.0000 93.0000 87.0000 85.0000 83.0000 85.0000 83.0000 83.0000 82.0000 87.0000 85.0000 91.0000 85.0000 85.0000 77.0000 78.0000 82.0000 79.0000 81.0000 82.0000 110.0000 77.0000 76.0000 76.0000 77.0000 79.0000 78.0000 78.0000 78.0000 78.0000 78.0000 77.0000 81.0000 102.0000 78.0000 77.0000 77.0000 77.0000 78.0000 77.0000 76.0000 78.0000 77.0000 79.0000 78.0000 77.0000 78.0000 78.0000 76.0000 76.0000 78.0000 78.0000 83.0000 84.0000 79.0000 82.0000 84.0000 84.0000 83.0000 83.0000 84.0000 83.0000 84.0000 84.0000 84.0000 85.0000 83.0000 86.0000 90.0000 83.0000 83.0000 84.0000 85.0000 84.0000 84.0000 83.0000 84.0000 85.0000 85.0000 84.0000 84.0000 95.0000 102.0000 83.0000 83.0000 84.0000 84.0000 82.0000 83.0000 95.0000 83.0000 83.0000 83.0000 89.0000 83.0000 83.0000 82.0000 83.0000 83.0000 82.0000 83.0000 82.0000 85.0000 82.0000 83.0000 83.0000 78.0000 78.0000 77.0000 77.0000 78.0000 78.0000 78.0000 78.0000 78.0000 81.0000 90.0000 89.0000 83.0000 85.0000 85.0000 96.0000 116.0000 101.0000 84.0000 80.0000 90.0000 86.0000 88.0000 87.0000 87.0000 82.0000 82.0000 78.0000 77.0000 88.0000 78.0000 77.0000 76.0000 79.0000 84.0000 82.0000 83.0000 83.0000 81.0000 77.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 88.0000 76.0000 76.0000 77.0000 76.0000 77.0000 76.0000 81.0000 83.0000 85.0000 80.0000 76.0000 77.0000 76.0000 77.0000 76.0000 77.0000 77.0000 78.0000 82.0000 82.0000 83.0000 84.0000 85.0000 83.0000 82.0000 82.0000 82.0000 81.0000 84.0000 83.0000 82.0000 81.0000 82.0000 82.0000 83.0000 82.0000 81.0000 77.0000 78.0000 76.0000 76.0000 77.0000 77.0000 76.0000 77.0000 82.0000 80.0000 82.0000 83.0000 83.0000 82.0000 81.0000 80.0000 82.0000 82.0000 82.0000 81.0000 84.0000 81.0000 82.0000 83.0000 80.0000 80.0000 82.0000 76.0000 76.0000 76.0000 77.0000 77.0000 76.0000 77.0000 78.0000 76.0000 79.0000 82.0000 82.0000 82.0000 82.0000 82.0000 88.0000 85.0000 86.0000 82.0000 82.0000 107.0000 83.0000 85.0000 80.0000 78.0000 83.0000 82.0000 85.0000 81.0000 82.0000 81.0000 78.0000 76.0000 83.0000 82.0000 80.0000 80.0000 81.0000 81.0000 82.0000 81.0000 74.0000 76.0000 76.0000 77.0000 77.0000 76.0000 76.0000 78.0000 77.0000 77.0000 81.0000 82.0000 82.0000 92.0000 83.0000 82.0000 76.0000 79.0000 76.0000 77.0000 77.0000 76.0000 77.0000 81.0000 77.0000 77.0000 80.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 76.0000 77.0000 76.0000 76.0000 77.0000 77.0000FileName : Celine_091321.impDat (Current dataset : 1)
+Signal(s) Used : Ca Ka, Na Ka, K Ka, Al Ka, Mn Ka, Cr Ka, Mg Ka, Ti Ka, Si Ka, Fe Ka
+Spectrometers Conditions : Sp1 PET, Sp2 LTAP, Sp3 LPET, Sp4 TAP, Sp5 LLIF, Sp1 PET, Sp2 LTAP, Sp3 LPET, Sp4 TAP, Sp5 LLIF
+Full Spectrometers Conditions : Sp1 PET(2d= 8.75,K= 0.000144), Sp2 LTAP(2d= 25.745,K= 0.00218), Sp3 LPET(2d= 8.75,K= 0.000144), Sp4 TAP(2d= 25.745,K= 0.00218), Sp5 LLIF(2d= 4.0267,K= 5.8E-05), Sp1 PET(2d= 8.75,K= 0.000144), Sp2 LTAP(2d= 25.745,K= 0.00218), Sp3 LPET(2d= 8.75,K= 0.000144), Sp4 TAP(2d= 25.745,K= 0.00218), Sp5 LLIF(2d= 4.0267,K= 5.8E-05)
+Column Conditions : Cond 1 : 15keV 278.8476nA , Cond 2 : 15keV 280.8923nA
+ , Cond 1 : Ca Ka, Na Ka, K Ka, Al Ka, Mn Ka
+ , Cond 2 : Cr Ka, Mg Ka, Ti Ka, Si Ka, Fe Ka
+Date : Sep-15-2021
+User Name : SX-663\\SX-User
+Setup Name : C:\\SX data\\Analysis Setups\\Image&Profiles\\Celine_091321.impSet
+DataSet Comment : BOO17-5N_Map1
+Comment :
+Analysis Date : Tuesday, September 14, 2021 6:57:42 AM
+Project Name : Default Project
+Sample Name : Default Sample
+Analysis Parameters :
+Sp Elements Xtal Position Bias Gain Dtime Blin Wind Mode
+Sp1 Ca Ka PET 38388 1304 1031 3 560 Inte
+Sp2 Na Ka LTAP 46363 1311 3055 3 560 Inte
+Sp3 K Ka LPET 42765 1846 985 3 560 Inte
+Sp4 Al Ka TAP 32459 1325 3181 3 560 Inte
+Sp5 Mn Ka LLIF 52202 1820 411 3 560 Inte
+Sp1 Cr Ka PET 26172 1304 1031 3 560 Inte
+Sp2 Mg Ka LTAP 38500 1311 3055 3 560 Inte
+Sp3 Ti Ka LPET 31416 1846 985 3 560 Inte
+Sp4 Si Ka TAP 27737 1325 3181 3 560 Inte
+Sp5 Fe Ka LLIF 48083 1820 411 3 560 Inte
+Peak Position : Sp1 38388, Sp2 46363, Sp3 42765, Sp4 32459, Sp5 52202, Sp1 26172, Sp2 38500, Sp3 31416, Sp4 27737, Sp5 48083
+Start : X = -1748 Y = -29677 Z = 14
+Stop :
+Dwell Time : 0.07 Sec
+Acquisition type : Stage Grid
+Step Number : 504
+Line Number : 504
+Beam Size : N/A
+Step Size : 4.000
+
+
+
+Y Unit : cts
+
+
+ 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504
+1 1987 1954 2079 1974 2083 2103 2073 2128 2028 1850 2121 1916 1938 2013 2066 2037 2109 2075 2093 2032 2053 2087 2118 2130 2082 2047 1986 2056 1994 2006 1920 1949 1970 1981 1922 2279 1986 1984 1992 1983 1960 1958 1996 1910 1972 1939 1934 1962 1984 1948 1933 1929 1850 1921 1948 1941 1942 1978 1997 2014 1973 1939 1985 1871 1595 1935 1964 1864 1905 1894 1896 1836 1894 1908 1920 1804 1876 1871 1812 1833 1928 1830 1844 1874 1982 1853 1934 1857 1920 1837 1856 1882 1964 2920 2025 1817 1908 1967 1925 1901 1947 1895 1912 1886 1967 1898 1945 1939 1935 1975 1962 2007 1963 1930 1941 1914 1893 1994 1986 1930 1954 2039 1950 2004 2005 1976 1943 1906 2046 2070 2105 2129 2971 2420 2304 2475 2051 2036 2184 2213 2047 2049 2064 2043 2788 2104 1979 1972 2056 2022 2153 2087 2180 2124 2094 1993 1992 2097 1957 2016 2014 3758 2197 2042 2032 2024 2018 2226 2570 3155 2870 2174 2000 1983 1942 2018 1974 1944 2043 1962 2047 1954 1961 2035 2083 2085 2003 2020 2069 2153 1988 1957 2014 2087 2054 1963 2052 2043 1999 1947 2007 2109 2035 2016 2100 2000 2013 1980 2012 2051 2072 2083 1978 2015 2078 2232 3343 3864 3847 2105 3254 3823 3869 3952 4521 3751 4286 4042 3589 3693 3570 3397 3597 3553 3877 3837 3747 3997 3824 2952 2986 3676 3975 4050 4008 3699 3649 3843 3940 4099 3524 3987 4367 3461 2725 2960 2424 2145 2095 2132 1941 2070 2081 2488 2146 2270 2069 2080 2078 2062 2666 2099 2120 2152 2076 2052 2041 2033 2054 2067 2222 2012 2114 2088 2031 2142 2107 2265 2502 2852 3239 3536 3372 3914 2630 3431 3709 2810 3996 3249 3964 2999 3846 3832 3698 3362 2881 2706 2166 1969 1923 2051 2080 2138 2002 2054 2021 1992 2047 1967 2025 2007 1934 2068 1980 1990 1914 2039 1982 1995 2016 2098 2067 2048 1999 2044 3706 3266 1668 1885 1802 2099 1990 2050 1939 2059 2032 2039 2112 2146 2052 2088 2062 2041 2006 2086 2084 2090 2087 2046 1996 2078 2103 2030 2069 2018 2019 1994 1957 2042 1978 2160 2823 3063 2137 2088 2327 1857 2111 2065 2149 2063 2049 1962 1987 2056 2027 2041 1925 1945 2027 2001 1985 2006 2000 2033 1998 1951 1929 1985 1983 1939 1983 1967 1990 2063 1946 2017 1998 1990 2039 2048 2012 1996 2065 2041 2001 2053 2022 2013 1974 2074 2572 2105 2021 2040 2064 2071 2022 2020 2057 2032 2056 2095 1877 2073 1991 1990 2428 1931 2029 2031 2016 2006 2046 2024 1984 1941 2071 2061 2049 2042 1974 2010 1989 1970 2000 1984 2010 2001 1932 1921 1949 1958 2033 2998 1998 2039 2012 2023 1988 2020 2522 2108 1998 2059 2051 2008 2054 2096 2009 1914 1802 1440 1605 1981 2049 1930 2037 1900 1945 1919 1976 1969 1966 1857 1934 1934 1913 1958 1862 1945 1895 1943
+2 2332 2198 2226 2202 2107 2090 2033 2026 1757 2032 2119 1966 1903 2029 2090 2051 2144 1991 1874 2096 2019 2018 2093 2067 2154 1942 1989 2095 1990 2047 1987 2005 1986 1939 2445 2190 2034 2024 1979 1945 1995 1999 1934 2001 1962 1955 1965 1918 1902 1908 1922 1963 1921 1935 1928 1966 1953 1925 1986 1952 1993 1934 2000 1935 1301 1842 1905 1886 1910 1884 1861 1874 1887 1869 1871 1802 1881 1864 1846 1880 1891 1838 1866 1829 1805 1804 1908 1856 1972 1823 1903 1921 2755 1959 1915 1884 1873 1890 1936 1924 1959 2005 1863 1956 1926 1949 1909 2007 1982 1855 1999 1905 1855 1917 1921 1975 1917 2006 1979 1977 2020 2010 1995 1968 1944 1922 1949 1987 2004 2045 2096 2131 2339 2289 2020 2457 2116 1978 2056 2164 2286 2044 2068 2084 2587 2245 1952 1826 1988 2122 2205 1928 2096 2107 2095 2072 2077 1989 2030 1904 1963 3104 1961 2092 2102 2375 3134 3825 4043 3994 3763 3068 2383 1956 1995 2063 1986 1965 1923 2017 2094 2013 2049 2000 2006 2047 2078 2063 1979 2092 2001 1960 2014 2033 2011 2005 2002 1992 2011 1993 2067 2034 1982 2026 2088 2035 2025 2038 2055 2065 2005 2093 2043 1990 2157 2078 2801 3693 3767 3727 2846 3401 4214 4116 3366 3732 3929 4034 3755 3730 3353 3589 3412 3894 4008 4006 3844 4042 3954 3862 3095 4028 4009 3864 4144 3949 3285 3684 2972 3954 4209 3549 3351 3483 3512 2830 3129 2177 2469 2111 2167 2063 2209 2344 2205 2145 2187 2107 1978 1984 2621 2051 2018 1989 2032 2002 2061 2105 2081 2265 2051 2040 2101 2085 2115 2125 2868 3459 3643 3598 3684 3661 3244 3541 3935 3975 4090 4612 3863 3959 3461 3508 3268 3851 3696 3301 2613 2099 1899 1983 2001 2060 1986 2044 2080 1987 2019 2083 2025 1916 1930 2083 1962 2085 2002 2009 2010 2044 2010 1980 2011 2021 2049 2040 2040 2096 2394 4142 3000 1991 2024 1767 2041 2045 1954 2069 2093 1933 2081 2131 2158 2050 2027 1994 2154 2091 2202 2044 2086 2057 2024 2064 1950 2045 2007 2068 1990 1992 1937 1974 1964 2363 3343 2345 2163 2123 2050 1155 2049 2024 2059 2012 1987 2073 2126 2034 1927 2006 1949 1967 1983 1987 1990 1983 2123 1999 1909 1939 1982 1976 1991 1950 1969 2074 1954 2042 1938 2043 2038 2022 1970 2005 1995 2101 1987 2027 2063 2035 1972 1972 1955 1939 2440 2033 2080 2033 2086 2038 2053 2037 2047 2043 2110 2024 1955 2027 1997 1854 2383 2014 2019 2018 2114 2001 2047 2003 2030 2040 2086 2043 2020 1996 1996 2041 1976 1904 1922 1964 2012 1971 2012 1930 1940 2013 3141 2025 1982 2039 1976 1957 1920 2018 2133 2033 2096 1982 2105 1929 1965 2043 1920 1806 2002 1985 1959 2016 1909 1995 1971 1986 1964 1914 1917 1942 1956 1921 1917 1927 1920 2006 1977 1948 1875 1857XMapTools can import two types of spot data file that are generated by CAMECA microprobes.
DataSet/Point Na2O MgO SiO2 Al2O3 K2O FeO MnO BaO Cr2O3 Cl CaO TiO2 Total X Y Z Comment Mean Z Date
+1 / 1 . 0.000 0.047 0.000 0.000 0.012 0.049 0.102 0.000 0.000 0.019 54.334 0.000 54.562 7160 -17670 49 9.049 21.01.13 12:56
+2 / 2 . 0.043 0.020 29.964 1.474 0.019 0.716 0.118 0.435 0.014 0.013 27.921 37.349 98.086 6456 -17357 50 14.564 21.01.13 12:59In this example data from Na2O to TiO2 are imported.
Each line represents a single analysis. The first entry DataSet/Point is defined using space (noted ” ” below) whereas tabulations (“\\t”) are used to separate other entries. Only this format can be properly read by XMapTools. The correct format for each data row is:
Number” “/” “Number” “.”\\t”Number”\\t”Number”\\t”Number …Example:
DataSet/Point Na2O MgO SiO2 Al2O3 K2O FeO MnO BaO Cr2O3 Cl CaO TiO2 Total X Y Z Comment Mean Z Date
+1 / 1 . 0.000 0.047 0.000 0.000 0.012 0.049 0.102 0.000 0.000 0.019 54.334 0.000 54.562 7160 -17670 49 9.049 21.01.13 12:56
+2 / 2 . 0.043 0.020 29.964 1.474 0.019 0.716 0.118 0.435 0.014 0.013 27.921 37.349 98.086 6456 -17357 50 14.564 21.01.13 12:59Alternatively, the following input also works:
DataSet/Point Na2O MgO SiO2 Al2O3 K2O FeO MnO BaO Cr2O3 Cl CaO TiO2 Total X Y Z Comment Mean Z Date
+1 0.000 0.047 0.000 0.000 0.012 0.049 0.102 0.000 0.000 0.019 54.334 0.000 54.562 7160 -17670 49 9.049 21.01.13 12:56
+2 0.043 0.020 29.964 1.474 0.019 0.716 0.118 0.435 0.014 0.013 27.921 37.349 98.086 6456 -17357 50 14.564 21.01.13 12:59An example is provided below. In this example, data ranging from Na₂O to FeO is imported.
FileName : Celine_test_091321.qtiDat
+Signal(s) Used : Na Ka, Mg Ka, Al Ka, Si Ka, K Ka, Ca Ka, Ti Ka, Cr Ka, Mn Ka, Fe Ka
+Spectrometers Conditions : Sp2 LTAP, Sp2 LTAP, Sp4 TAP, Sp4 TAP, Sp3 LPET, Sp1 PET, Sp3 LPET, Sp1 PET, Sp5 LLIF, Sp5 LLIF
+Full Spectrometers Conditions : Sp2 LTAP(2d= 25.745,K= 0.00218), Sp2 LTAP(2d= 25.745,K= 0.00218), Sp4 TAP(2d= 25.745,K= 0.00218), Sp4 TAP(2d= 25.745,K= 0.00218), Sp3 LPET(2d= 8.75,K= 0.000144), Sp1 PET(2d= 8.75,K= 0.000144), Sp3 LPET(2d= 8.75,K= 0.000144), Sp1 PET(2d= 8.75,K= 0.000144), Sp5 LLIF(2d= 4.0267,K= 5.8E-05), Sp5 LLIF(2d= 4.0267,K= 5.8E-05)
+Column Conditions : Cond 1 : 15keV 20nA
+Date : Sep-15-2021
+User Name : SX-663\\SX-User
+Setup Name : C:\\SX data\\Analysis Setups\\Quanti\\Celine_091321.qtiSet
+DataSet Comment : BOO17-3S_map2_cpx1
+Comment :
+Analysis Date : Monday, September 13, 2021 4:45:45 PM
+Project Name : Default Project
+Sample Name : Default Sample
+Analysis Parameters :
+Sp Elements Xtal Position Bg1 Bg2 Slope Bias Gain Dtime Blin Wind Mode
+Sp2 Na Ka LTAP 46369 -700 800 1311 3117 3 560 Inte
+Sp2 Mg Ka LTAP 38513 -1150 1150 1311 3055 3 560 Inte
+Sp4 Al Ka TAP 32464 800 1.2 1324 3227 3 560 Inte
+Sp4 Si Ka TAP 27737 750 1.1 1325 3181 3 560 Inte
+Sp3 K Ka LPET 42762 -600 600 1845 981 3 560 Inte
+Sp1 Ca Ka PET 38391 700 1.1 1307 1026 3 560 Inte
+Sp3 Ti Ka LPET 31406 600 1.05 1846 985 3 560 Inte
+Sp1 Cr Ka PET 26186 -500 500 1304 1031 3 560 Inte
+Sp5 Mn Ka LLIF 52200 650 1.05 1820 412 3 560 Inte
+Sp5 Fe Ka LLIF 48084 800 1.2 1820 411 3 560 Inte
+Peak Position : Sp2 46369 (-700, 800), Sp2 38513 (-1150, 1150), Sp4 32464 (800, Slope = 1.2), Sp4 27737 (750, Slope = 1.1), Sp3 42762 (-600, 600), Sp1 38391 (700, Slope = 1.1), Sp3 31406 (600, Slope = 1.05), Sp1 26186 (-500, 500), Sp5 52200 (650, Slope = 1.05), Sp5 48084 (800, Slope = 1.2)
+Current Sample Position : X = -10413 Y = 32015 Z = 60 BeamX = 0.00 BeamX = 0.00
+Standard Name :
+Na ,Al On albite
+Mg ,Si ,Ca On Wakefield diopside
+K On kspar
+Ti On TiO2
+Cr On MgCr2O4
+Mn On Rhodon 41522 AMNH
+Fe On RKFAYb7
+Standard composition :
+albite = Na : 8.77%, Al : 10.29%, Si : 32.13%, O : 48.81%
+Wakefield diopside = Si : 25.94%, O : 44.43%, Na : 0.02%, Mg : 11.22%, Al : 0.02%, Ca : 18.53%, Ti : 0.02%, Mn : 0.02%, Fe : 0.09%
+kspar = O : 45.94%, Na : 0.85%, Al : 9.83%, Si : 30.10%, K : 12.39%, Ba : 0.70%
+TiO2 = Ti : 59.95%, O : 40.05%
+MgCr2O4 = Mg : 12.64%, Cr : 54.08%, O : 33.28%
+Rhodon 41522 AMNH = O : 37.73%, Mg : 2.36%, Si : 21.98%, Ca : 1.06%, Mn : 35.01%, Fe : 1.80%
+RKFAYb7 = Si : 13.84%, O : 31.37%, Mg : 0.06%, Al : 0.05%, Ca : 0.02%, Mn : 1.55%, Fe : 52.62%, Zn : 0.38%
+Calibration file name (Element intensity cps/nA) :
+Na ,Al : Other\\albite_15kV_NaKa-Sp2-LTAP_AlKa-Sp4-TAP_010.calDat (Na : 127.8 cps/nA, Al : 189.2 cps/nA)
+Mg ,Si ,Ca : Other\\Wakefield diopside_15kV_MgKa-Sp2-LTAP_SiKa-Sp4-TAP_CaKa-Sp1-PET_019.calDat (Mg : 315.4 cps/nA, Si : 541.5 cps/nA, Ca : 101.3 cps/nA)
+K : Other\\kspar_15kV_KKa-Sp3-LPET_017.calDat (K : 208.0 cps/nA)
+Ti : Other\\TiO2_15kV_TiKa-Sp3-LPET_029.calDat (Ti : 1589.2 cps/nA)
+Cr : Other\\MgCr2O4_15kV_CrKa-Sp1-PET_013.calDat (Cr : 247.4 cps/nA)
+Mn : Other\\Rhodon 41522 AMNH_15kV_MnKa-Sp5-LLIF_020.calDat (Mn : 192.7 cps/nA)
+Fe : Other\\RKFAYb7_15kV_FeKa-Sp5-LLIF_057.calDat (Fe : 317.7 cps/nA)
+Beam Size : N/A
+
+ Weight% Atomic% Oxide
+
+DataSet/Point Na Mg Al Si K Ca Ti Cr Mn Fe O Total Na Mg Al Si K Ca Ti Cr Mn Fe O Total Na2O MgO Al2O3 SiO2 K2O CaO TiO2 Cr2O3 MnO FeO Total X Y Z Beam X Beam Y Comment Distance (?) Mean Z Point# Date
+1 / 1 . 0.743369 7.000587 0.992360 23.675010 0.000010 15.496050 0.028793 0.050500 0.047470 7.429382 41.095420 96.558950 0.753605 6.712945 0.857190 19.646370 0.000006 9.010886 0.014010 0.022636 0.020138 3.100467 59.861750 100.000000 1.002046 11.609060 1.875058 50.649700 0.000012 21.682070 0.048028 0.073809 0.061296 9.557870 96.558950 -10413.0 32015.0 60.0 BOO17-3S_map2_cpx1 0.00 12.714160 1 Monday, September 13, 2021 4:45:45 PM
+1 / 2 . 0.683164 7.306072 1.054231 24.247130 0.005349 15.870280 0.049491 0.006556 0.033462 7.281908 42.080150 98.617790 0.676747 6.845810 0.889827 19.661410 0.003116 9.017649 0.023530 0.002872 0.013871 2.969490 59.895680 100.000000 0.920890 12.115650 1.991962 51.873660 0.006443 22.205690 0.082554 0.009582 0.043208 9.368145 98.617790 -10408.3 32015.0 60.0 BOO17-3S_map2_cpx1 4.68 12.939130 2 Monday, September 13, 2021 4:48:12 PMThe EPMA converter can be used to convert raw data for supported instruments to XMapTools format.
The following data formats are currently supported:
The converter can be accessed via Project and Imports, using the Open XMapTools' EPMA Converter button.
Main steps:
For more detailed information on the EPMA Converter, refer to the embedded documentation available within the program.
Use the Select the format dropdown menu to select data format. After selecting the data format two text boxes are displayed indicating the types of files required for the selected format. The first on the left is for the map and the second one for the spot analyses.
After selecting the data format, press the Set a destination folder button and select an empty folder to save the data in the XMapTools format. This folder must be empty and existing data will be deleted!
Note that this folder can then be transferred to a working data folder for separate storage from the raw data.
After selecting the destination folder, press the Import Maps button to select the folder containing the map files. XMapTools will convert all the maps available in this folder.
After selecting the maps, press the Validate map selection button to continue.
After selecting all files for the spot analyses, press the button generate Standards.txt. This will end the procedure and close the Converter. Note that when you return in XMapTools, the working directory has been adjusted and you can start importing your maps.
To import map data in XMapTools, select the 'Project and Import' tab and press the Import Maps ![]() button located in 'Import Maps and Images'. This will open the import module and prompt you to select files.
button located in 'Import Maps and Images'. This will open the import module and prompt you to select files.
Select the set of map files you want to import. Choose compatible files from the 'Pick Map File(s)' pop-up window. Note that multiple files can be selected at once. Any selected file that cannot be imported due to an incompatible format, for example, will be skipped during import.
Format: Map files must have the *.txt, *.asc, *.dat or *.csv extension, no header and a name compatible with the XMapTools default element names for identification. The default lists of compatible element and oxide names are given in the source of the help file below.
Selected map files are listed in the main table. More maps can be added by pressing the map selection button (see above)
File: Contains the file names
Press the Import data button to import the selected maps into XMapTools after the corrections have been applied.
For more detailed information on the EPMA Converter, refer to the embedded documentation available within the program.
This section describes how to work in XMapTools with data that have been calibrated by an other program.
The calibrated maps should be translated first into a compatible format (e.g. txt or csv file). Use the element or oxide abbreviation as file name (e.g. Si.txt, SiO2.txt, Ce.txt, etc.).
Quantitative maps can be imported in XMapTools via the Import Tool. Quantitative maps can be expressed in µg/g or wt% of elements (e.g. Si.txt, Al.txt, etc.) or oxides (e.g. SiO2.txt, Al2O3.txt, etc.). These maps are imported in the category 'Merged' data. The data format is not specified during the import.
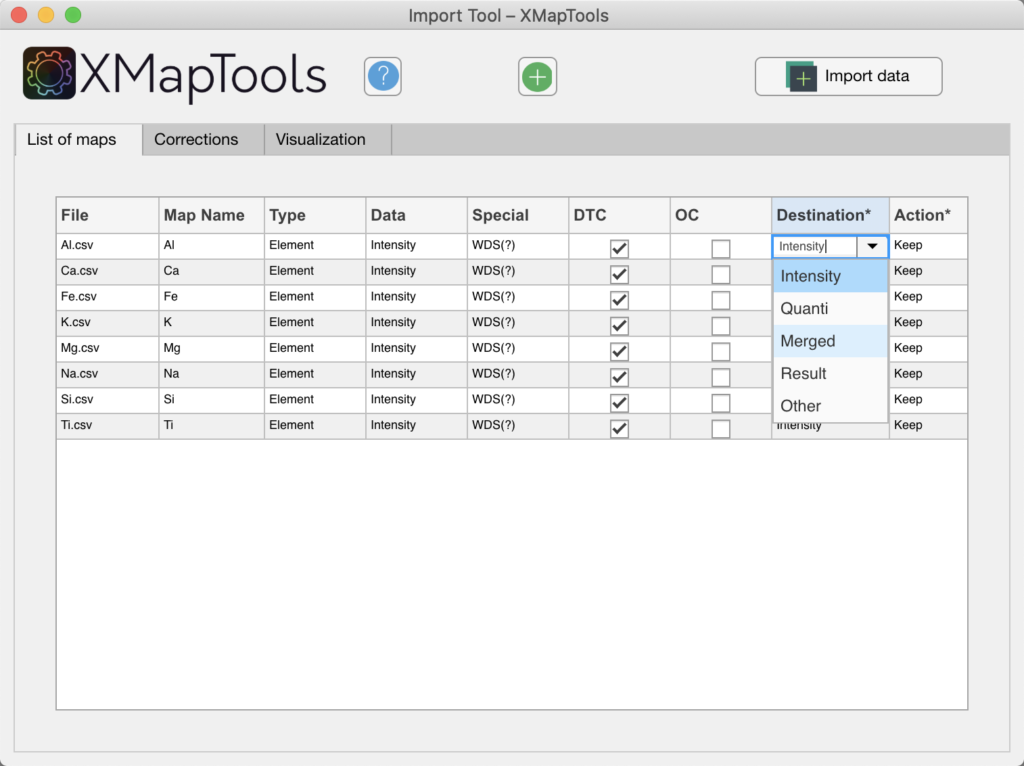
Figure: If maps are expressed in mass of elements, the destination should be changed from 'Intensity' to 'Merged'.
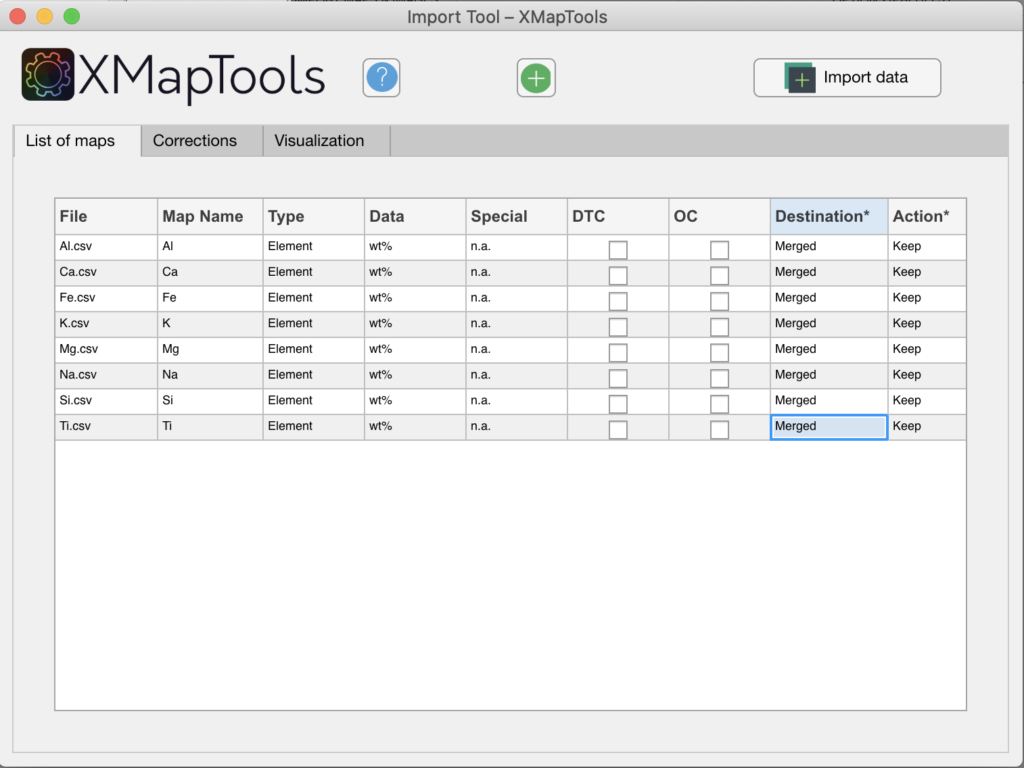
Figure: In this example the destination for each map has been set to 'Merged'.
The compositional maps should ideally be expressed in oxide wt%. if the imported maps are in µg/g or wt% of elements, a conversion step can be required to calculate structural formulas.
In the primary menu, select the dataset Imported_Maps and then right-click on its name. Select the option Convert to open the Converter module.
Select the conversion method in the Converter tool and press the Apply button.
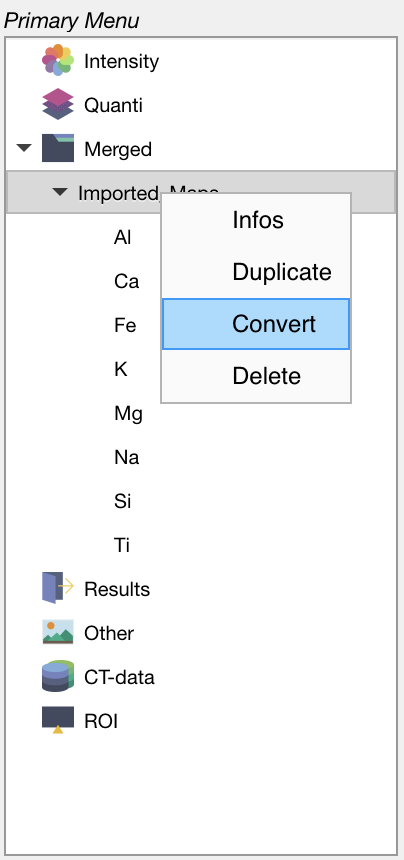
Figure: Select a dataset and then right-click to open the Converter from the primary menu.
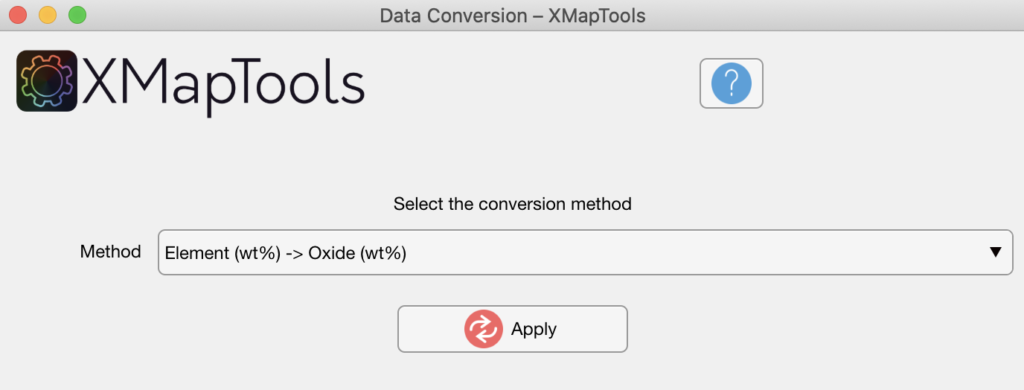
Figure: Data conversion tool. In this example data are converted from element wt% to oxide wt%.
Display a map from the imported dataset using the Primary Menu and open the Classify tab. Select Training Set (Classification) in the Secondary Menu and press the Add ![]() button in Classify and select the phases. Press again the Add
button in Classify and select the phases. Press again the Add ![]() button to create a new mask definition in the training set. This operation can be repeated until the correct number of phases is reached. Each mask definition can be deleted by right-clicking on the name and selecting Delete.
button to create a new mask definition in the training set. This operation can be repeated until the correct number of phases is reached. Each mask definition can be deleted by right-clicking on the name and selecting Delete.
You can rename each mask definition by double-clicking on his name in the Secondary Menu. Press Add ![]() in Classify when a mask definition is selected to add a region-of-interest (ROI).
in Classify when a mask definition is selected to add a region-of-interest (ROI).
Select the dataset in the category Merged of the Primary Menu and press the Add Maps for Classification ![]() button to add all the maps of the dataset in the list that will be used by the classification function.
button to add all the maps of the dataset in the list that will be used by the classification function.
Select a dataset in the Merged category of the Primary Menu and a Training Set in the Secondary Menu. Pick an algorithm in the tab Classify and press the Classify ![]() button. Note that the Classify button is only available when an appropriate dataset and training set are selected in the primary and secondary menus.
button. Note that the Classify button is only available when an appropriate dataset and training set are selected in the primary and secondary menus.
The imported dataset available in the Merged data category has to be divided into maps for each mask that will be stored under the Quanti data category.
Select a mask file in the Secondary Menu. Select the dataset in Merged and then right-click on its name. Select the Split (using maskfile) option.
The results are stored under the Quanti category as individual dataset, one for each mask. These data sets can be used to calculate maps of structural formulas or other calculations.
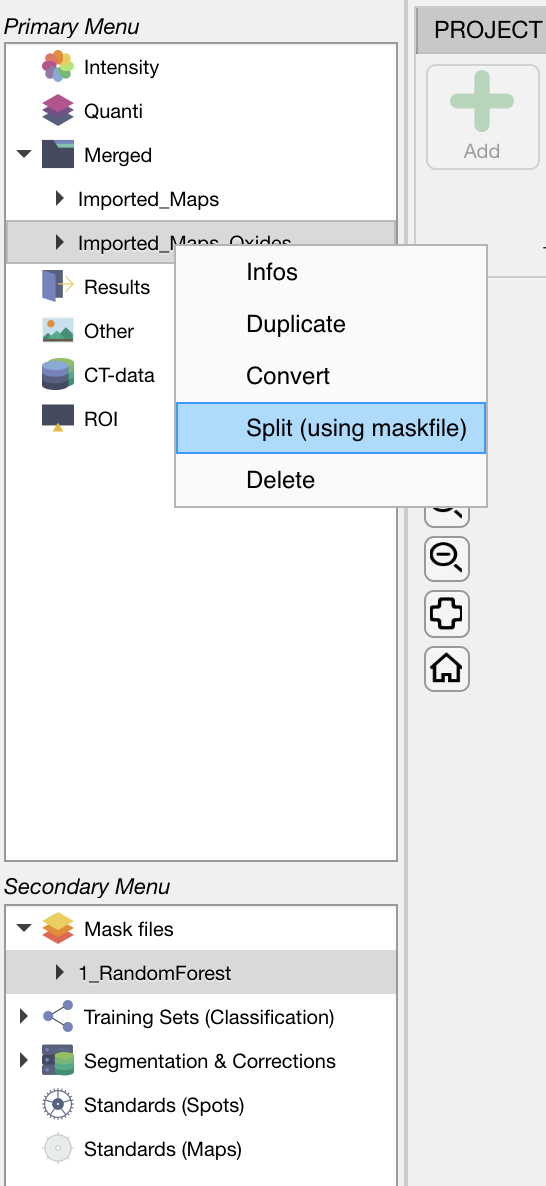
Figure: Select a mask file, then click on the dataset of interest (here Imported_Maps_Oxides) and then right-click on the name for accessing the menu.
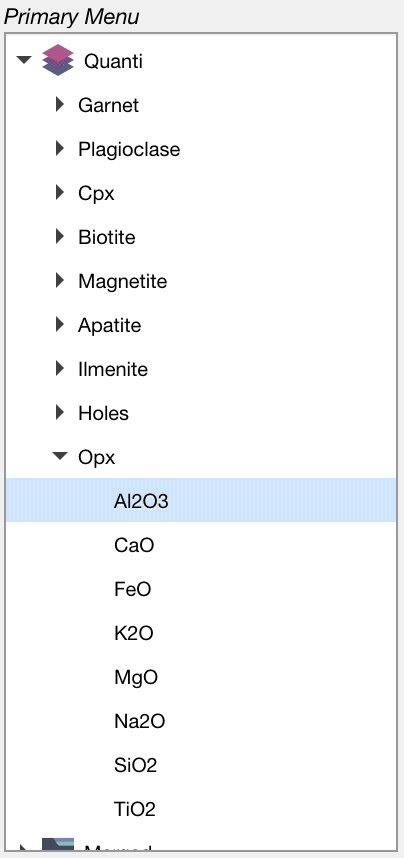
Figure: Results are stored in the category Quanti
Compositional map classification is the process of categorising and labelling groups of pixels within a dataset based on their composition. It generates a mask image showing the distribution of each mask/class (i.e. features can be mineral/epoxy/glass, etc.).
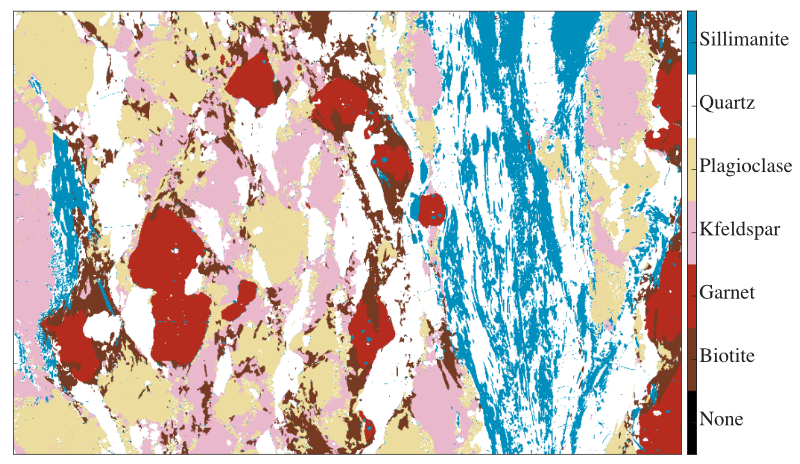

Figure: Example of mask image for a metapelite from the Himalaya published in Lanari & Duesterhoeft (2019). Each feature (mineral) is shown with a colour. Note that all the pixels of this image have been classified.
These tools are used to select an algorithm and maps to be used for classification.
The goal is to set the map input of the classification function. It is not required to use all available maps for classification; maps containing only noise are usually excluded.
The list of maps used by the classification function is displayed in the text field. The button Add Maps for Classification adds all maps available in the intensity tab of the primary menu. The button Edit Selected Map can have two modes: add (plus icon) or eliminate (minus icon) depending on whether the map selected in the primary menu is already available in the list or not. Clicking "plus" adds the selected map, whereas clicking "minus" eliminates the selected map from the list.
The machine learning algorithm used for classification can be selected via the algorithm menu available in the section Classification Parameters.
The following algorithms are available:
The principal components of a collection of points in a real coordinate space are a sequence of vectors consisting of best-fitting lines, each of them defined as one that minimizes the average squared distance from the points to the line. These directions constitute an orthonormal basis in which different individual dimensions of the data are linearly uncorrelated. The first principal component can equivalently be defined as a direction that maximizes the variance of the projected data. Principal component analysis (PCA) is the process of computing the principal components and using them to perform a change of basis on the data.
The button Generate Maps of the Principal Components (PCA) generates a map for each principal component and stores them in the section Other of the primary menu.
If the tick-box incl. PCA is selected, the maps of principal components are included as additional dimensions for the classification. Example: if 8 intensity maps are considered, a total of 14 maps of PC are added to the classification input, 7 for a normal PCA and 7 for a normalised PCA.
A training set must be selected in the secondary menu in order to activate the classification button.
The button Classify (Train a Classifier & Classify) trains a new classifier and performs the classification using the algorithm selected in the menu and the specified set of maps.
A new figure containing up to four plots will open and be continuously updated during classification. Do not close this figure until the classification is complete, otherwise the plots will not be displayed.
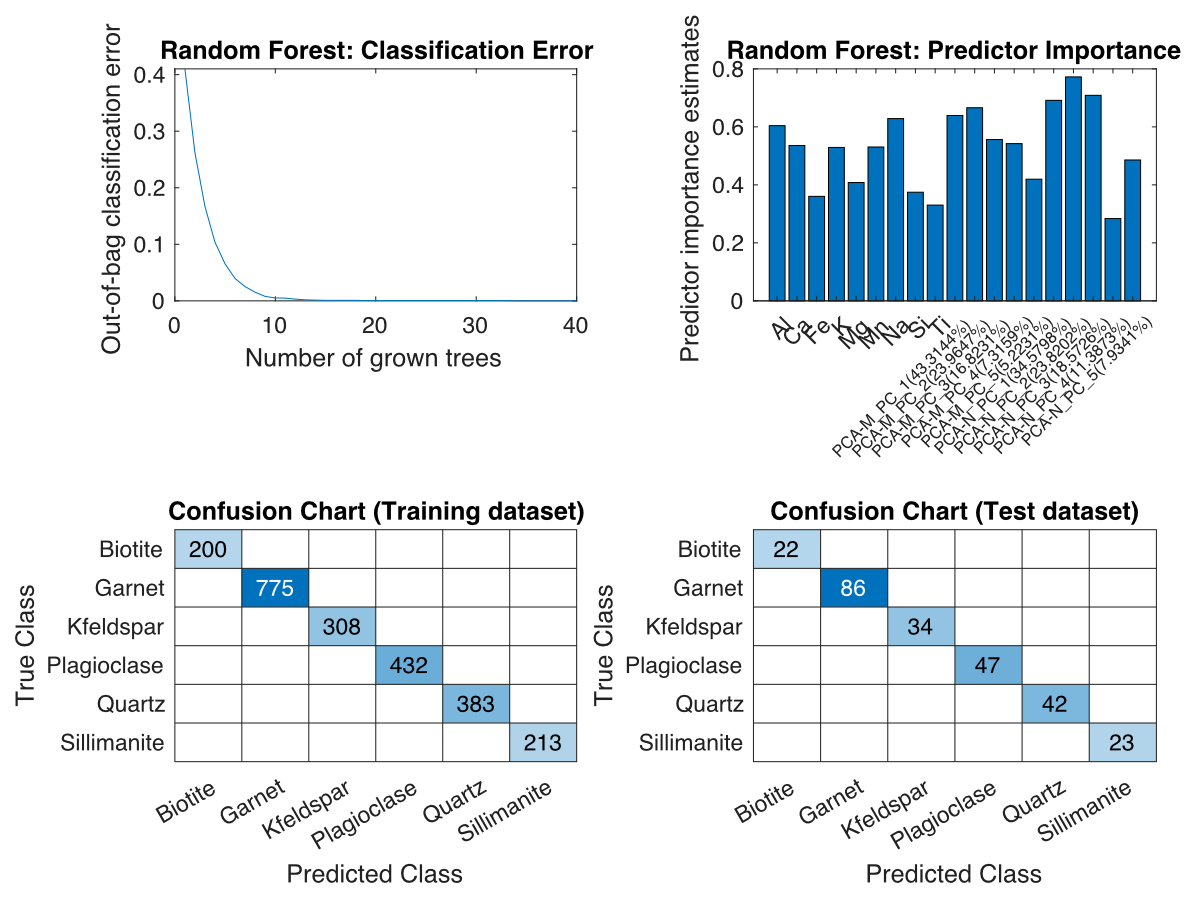
Figure: Plots for classification using the Random Forest algorithm. Top left: out-of-bag classification error vs. number of trees grown. Top right: Predictor importance. Bottom left: Confusion map of the training data set. Bottom right: Confusion plot of the test dataset.
Once the classification is achieved, a new mask file is generated and stored under Mask files in the secondary menu. The mask file is automatically selected and the mask image displayed in the main figure.
The following options apply to a given mask file. Select a mask file in the secondary menu to apply changes.
Select the option to hide pixels having a class probability lower than a given threshold for the selected mask file. Set the probability threshold used for hiding pixels. This value should range between 0 and 1.
The button Create new mask file with pixels filtered by probability generates a new mask file after filtering. This option replaces the BCR correction available in XMapTools 3 and is more adequate as only the misclassified pixels are excluded.
The modes of each class within a given region-of-interest can be exported using the tools provided in this section. Modes are given as surface percentage calculated using the number of pixels of each class. For minerals, this can be eventually extrapolated to volume fraction as discussed in Lanari & Engi (2017).
Select the ROI shape to be used to extract the local modes from the dropdown menu.
The button Add ROI allows a new region-of-interest (ROI) to be drawn on the figure. Note that a mask file must be selected before activating this mode. Results are displayed in the right window as a table containing the modes and number of pixels for each class and as a pie diagram.
The following ROI shapes are available:
The button Plot Compositions generates a plot using the data selected in the primary menu (either intensity, or a merged map) and the mask file selected in the secondary menu.
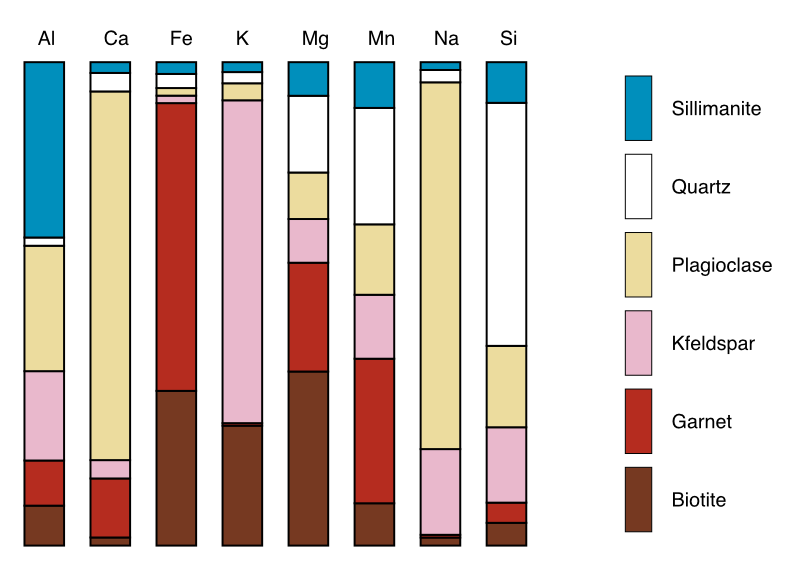
Figure: Example of compositional plot generated using a merged map (expressed in oxide wt%) and a mask file.
The following steps are required to convert raw data (e.g. X-ray maps) into maps of chemical composition:
The spot analyses used as internal standards in the calibration of EPMA data are referred to as "standards" in the following as they permit to define a calibration curve that correlates X-ray intensities to composition (e.g. expressed in oxide/element wt%).
It is necessary to import the standards from a standard file, check their positions and eventually correct, check the chemical compositions and eventually create new standards. All these steps can be achieved in XMapTools 4.
The file Standards.txt contains (i) the map coordinates and (ii) the spot analyses used for the standardisation. The map coordinates must be listed within a single row below the keyword >1. The oxide order is set below the keyword >2. X and Y must be the two last labels and must be listed in this specific order. The internal standards analyses are listed below the keyword >3 corresponding to the oxide order defined above (keyword >2).
>1 Here paste the image coordinates (Xmin Xmax Ymax Ymin)
+56.739 57.239 43.691 43.371
+
+>2 Here define the oxides order
+SiO2 MgO FeO Al2O3 X Y
+
+>3 Here paste the analyses
+25.4800 11.260 29.050 21.1400 1.4800 68.310 39.999
+52.9400 3.5300 3.0200 24.2300 0.0197 68.310 39.535
+52.5800 3.6300 2.7900 24.7200 0.0195 68.331 39.511The button Import (Import spots for Standards (from file)) is used to import standards from a file.
The box Import from Standards.txt is selected by default allowing the file 'Standards.txt' to be read automatically. If the file containing the standard data has a different name, unselect the box and it will be possible to select a file in the Pick a file pop-up window. At the moment all standards need to be stored within a single file. It is not possible to import standards from different files as existing standards will be eliminated when a new file is loaded.
Once loaded, standards are displayed on the main map with a label including the spot number and several plots are produced and shown in the category "Standards" of the live display module. You can get this global visualisation at any stage by selecting Standards (Spots) in the Secondary Menu. Three plots are produced if an element is selected in the primary menu, from top to bottom: (1) a plot showing intensity/composition versus sequence of standard to visualise if there is a good match between the standard compositions and the intensity values of the matching pixels; (2 and 3) two correlation maps, one for the selected element in the primary menu and a second one considering all elements.
To adjust the positions, display the map of a diagnostic element (Intensity) and select Standards (Spots) in the secondary menu. The two correlation maps should show a maximum value in yellow and the blue spot representing the current position should be centred on this optimum.
To adjust the position of the standards (all at once), put the mouse cursor over the blue circle showing up a transparent circle. Click on it and move the blue circle to the new position. The values in the two white fields on top will be adjusted. Then click on the button Refresh (important) to update the standard positions. The Refresh button is only available when standard positions have been changed and need to be saved.
Note: If no good correlation exists for a given element, the higher value of the second figure could not represent the optimal position.
It is possible to add new standard points directly in XMapTools. Note that these will not be saved to the file Standards.txt, and if the file is loaded again all changes will be lost.
The button Add standard point adds a new standard at selected coordinates, set by clicking on the map after pressing the button. Compositional data can be filled directly in the table, when this standard is selected.
Procedure:
A calibration step, also known as standardisation, is required to convert intensity maps into compositional maps (Lanari et al. 2019).
All minerals/objects are calibrated at the same time in XMapTools 4. Therefore it is required to select a Mask File in the Secondary Menu to activate the button Calibrate.
The approach implemented in XMapTools 4 provides a module for auto multi-phase calibration. The button Calibrate opens the Calibration Assistant for EPMA Data. This button is only available when a mask file is selected in the Secondary Menu.
For more detailed information on the Calibration Assistant, refer to the embedded documentation accessible from the assistant.
The new approach implemented in XMapTools 4 provides a module for automatic multi-phase calibration. The general procedure is described in De Andrade et al. (2006) and an advanced approach including pseudo-background correction is described in Lanari et al. (2019).
An automatic calibration is performed taking into account all spot analyses and all masks when the Calibrate button is pressed. The new algorithm first performs a general fit including all standards and then adjusts the calibration for each mineral. All calibration curves, including those for the general fit, are accessible from the tree menu.
When you press the Apply Standardisation button, all the calibrated maps for each mineral as well as a merged map are created and sent back to XMapTools.
Important
First make sure you check the quality of the calibration curves generated by the auto function!
The automatic function will work if all minerals have been measured with at least a few spot analyses and if there is at least one mineral with a composition above 1 wt% for each element. If a mineral or other feature (e.g. fracture) has no spot analyses, the program extrapolates a calibration from the general fit and thus "predicts" a composition. At this stage this composition is likely to be off because matrix effects are ignored!
INFO
The Calibration Assistant for EPMA Data opens when the Calibrate button in XMapTools is pressed.
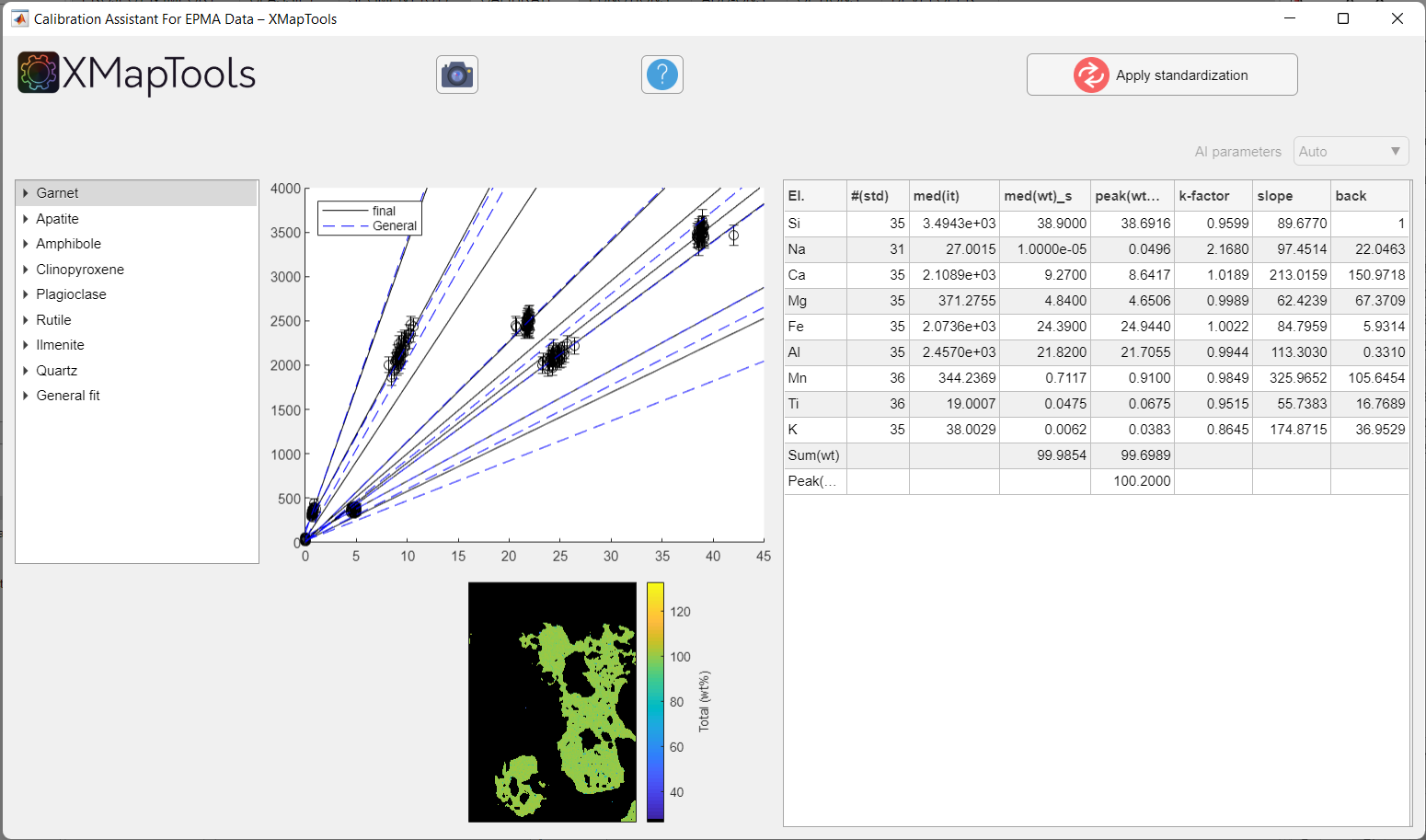
Figure: Example calibration for a clinopyroxene-garnet amphibolite metapelite of the Brasília orogen (Brazil), published in Tedeschi et al. (2017). Note that the calibration curves for all minerals are displayed when the window pops up.
Use the tree menu on the left to navigate through the list of minerals and elements and to view calibration curves.
When you select a mineral from the tree menu, a single plot showing the sum of elements/oxides (total wt%) is displayed. The plot in the middle shows all calibration curves, for all elements of the selected mineral. Some data is displayed in a table:
| Column | Description |
|---|---|
| El. | Element name (of the map); includes sum(wt%) and Peak(SumOx) labels |
| #(std) | Number of internal standards (spot analyses) used to calibrate the phase |
| med(it) | Median intensity value for all pixels of the selected mineral for each element |
| med(wt)_s | Median composition of all internal standard measurements (spot analyses) |
| mode(wt)_m | Most common composition in the calibrated pixels |
| k factor | Difference between the general fit calibration and the final mineral calibration (1 = identical) |
| Slope | Slope of the calibration curve |
| Background | Intercept of the calibration curve |
TIP
The values of mode(SumOx) and Sum(wt) may be different, in which case the median is likely to be influenced by non-Gaussian signals and may not be comparable with the median of the spot analyses. The comparison of both columns can be used to detect potential calibration problems.
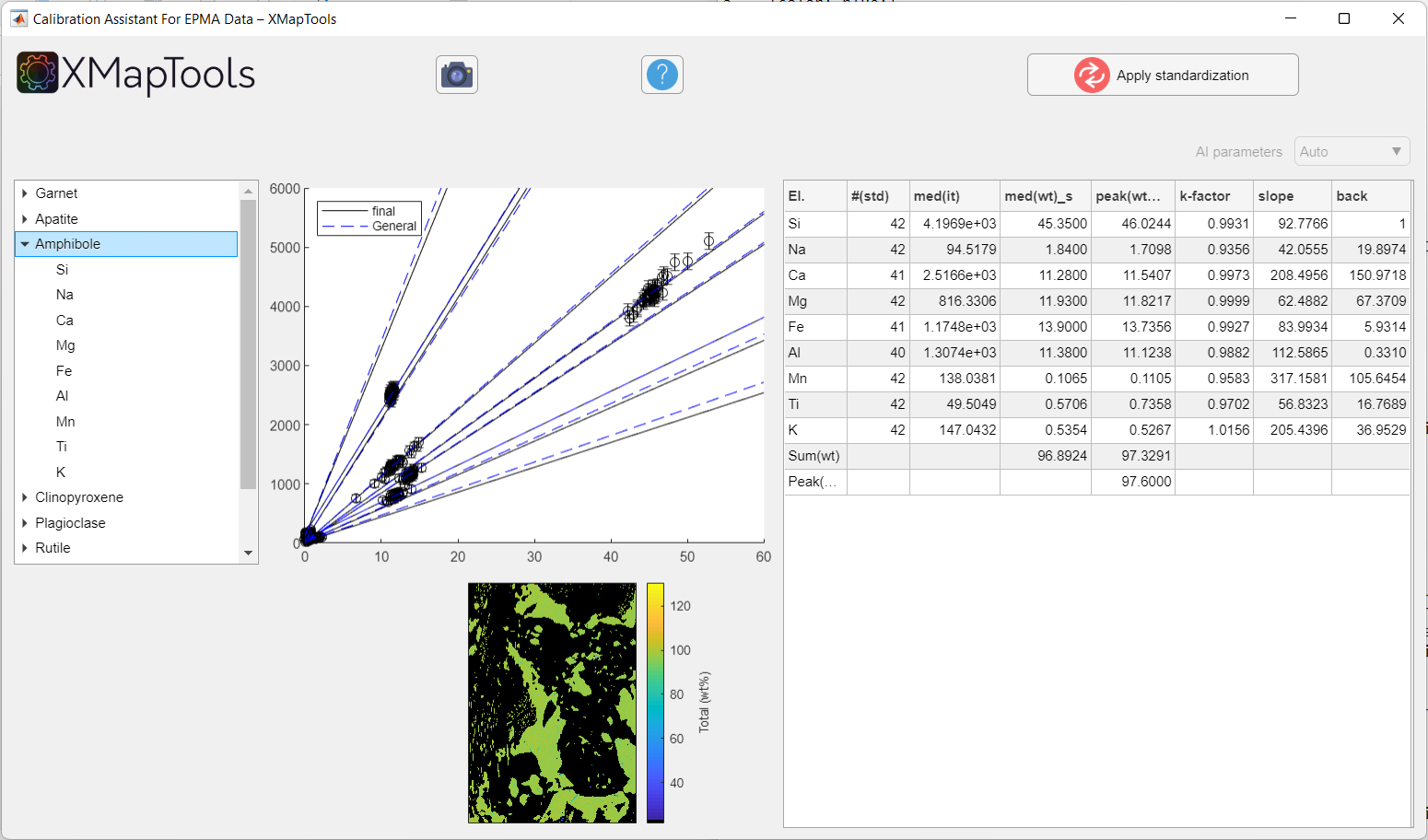
Figure: Example of calibration when a mineral is selected. All calibration curves for a given mineral are displayed.
To view a specific calibration curve (for a particular element), expand the menu by clicking on the small arrow to the left of the mineral name and select an element. The corresponding calibration curve is displayed together with the corresponding quantitative map (oxide wt-%).
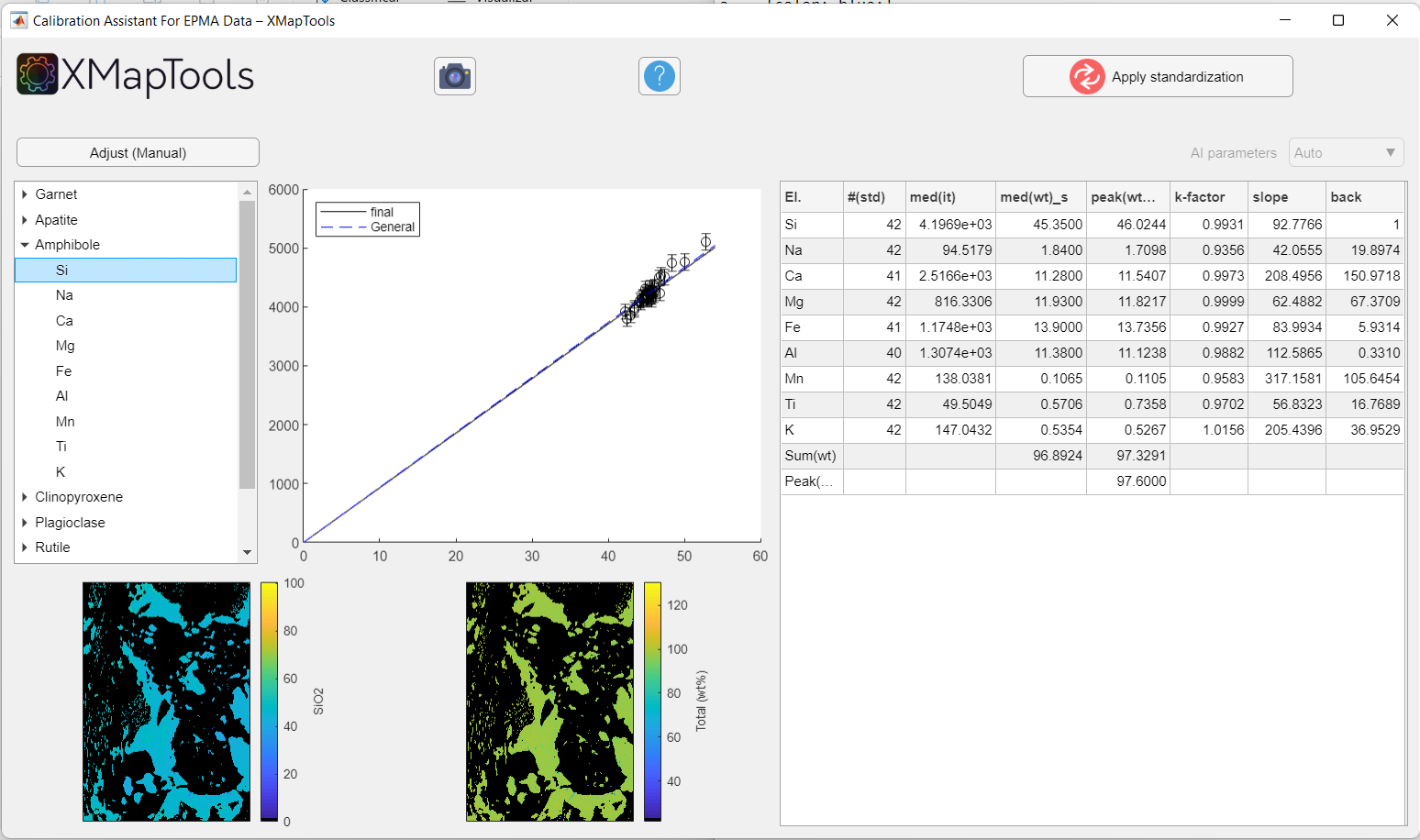
Figure: Example when an element of a given mineral phase is selected. The calibration curve and quantitative map for a given element are displayed.
When an element is selected, the Adjust ![]() button appears above the tree menu.
button appears above the tree menu.
Clicking this button displays two fields containing the values for background and slope: ![]()
Values can be changed manually by entering new values in the appropriate field. Press Enter to calculate and display the new calibration curve on the graph (this operation may take a few seconds).
Selecting General Fit (last option in the tree menu) will display a plot of the calibration curves for all elements. It is not possible to adjust the calibration curves in the general fit. This fit is automatically performed first by the program and is no longer used once the calibration of each mineral has been achieved.
After checking each calibration curve and adjusting if necessary, use the Apply Standardisation button ![]() to generate the calibrated maps.
to generate the calibrated maps.
By applying the standardisation:
A local bulk composition (abbreviation: LBC) represents the bulk composition of a spatial domain in a rock determined by integrating pixel compositions. As discussed in Lanari and Engi (2017), it is necessary to apply a density correction prior to exporting any local bulk composition.
To export local bulk compositions the following steps should be employed:
A density map is a map containing density data for each pixel of a map. It is calculated for a given mask file. Select a mask file in the secondary menu.
The button Generate Density Map (from a mask file) allows a density map to be generated from the selected mask file. A mask file should be selected to activate the button. Pressing this button opens a window with predefined average density values (provided that the mineral name was recognised and a reference value available in the internal database; when full names of minerals in English are used, the mineral should be recognised).
Note: mineral density values can be obtained from the website webmineral.com.
A density map will be created and stored under the category Other in the Primary Menu with the name 'Density [maskfile_name]'.
Merged maps are maps for which all pixels hold a chemical composition. A merged map is automatically created by the standardisation function in XMapTools 4. If you need to create a merged map manually, follow the procedure below.
In the primary menu, unfold Quanti and select a quanti map (mineral). The button Merge (Merge Quanti Data) in the section Calibrate becomes available. In the window that pops up, select the quanti maps to be merged. Select 'Ok' to generate the merged maps that will be stored in the category Merged.
A local bulk composition can be calculated from a region-of-interest (ROI). The ROI can be a rectangle or a polygon. The density correction is automatically applied.
Select a merged map in the primary menu and display an element. In the dropdown menu, select the wanted ROI shape: Rectangle ROI or Polygon ROI.
The button Add a ROI for LBC extraction allows a region-of-interest (ROI) to be drawn on the figure. If Rectangle ROI is selected, click on a corner, hold and drag to draw a rectangle. If Polygon ROI is selected, click on the image to draw the polygon and close by selecting the first point or right-clicking.
When a ROI is available, the table containing the local bulk composition appears in the live display module. The first column shows the element list, whereas compositions are listed in the column composition (unit: wt%). Below the table, a pie chart shows the repartition of the elements/oxides by weight.
The ROI can be edited and the composition values in the table and the pie chart are automatically updated.
The density map used is shown in a dropdown menu located in the live display module.
The button Copy Data to clipboard may be used to copy the LBC data from the table. The button Save may be used to save the LBC data as a .txt file.
An uncertainty approximation similar to what is described in Lanari and Engi (2017) is available.
Select a merged map and create a Rectangle ROI. Define the number of simulations Sim (default 100) and the shift Px in pixel (default 20). The shape will be randomly displaced and resized using two random variables calculated from the shift value (assuming a Gaussian distribution and the value of Px as 1 sigma expressed in number of pixels). Click on the button Calculate uncertainties using Monte-Carlo.
The areas used to approximate an uncertainty are plotted in a new figure and the result is shown in the table of the live display module. The column composition shows the original local bulk composition (selected ROI). The column 2std shows the 2 standard deviation value (note that the distributions are usually Gaussian as shown by Lanari & Engi (2017)). The last column shows the mean value of all compositions. This value should match the composition of the original ROI. If not, this means that a Gaussian distribution cannot be assumed for this element and the uncertainty is not correct.
XMapTools includes a set of external functions for calculating structural formulas, thermobarometry and other methods. Source code for all functions is available on the XMapTools GitHub repository.
The following structural formula functions are available in XMapTools. Each function calculates structural formulas and end-member proportions on a specified oxygen basis.
XMapTools External Function: structural formula of aluminosilicates. Version without loop & all Fe as Fe₂O₃. 5 Oxygen-basis.
XMapTools External Function: structural formula of Ca-amphibole. Fe₂O₃ added as possible input. Ti ordered onto M2 (Raase 1974).
| End-member | T1(4) | T2(2) | M2(2) | M13(2) | M4(2) | A(1) |
|---|---|---|---|---|---|---|
| Glaucophane (Gln) | Si₄ | Si₄ | Al,Al | Mg,Mg,Mg | Na,Na | V |
| Tremolite (Tr) | Si₄ | Si₄ | Mg,Mg | Mg,Mg,Mg | Ca,Ca | V |
| F-tremolite (Ftr) | Si₄ | Si₄ | Fe,Fe | Fe,Fe,Fe | Ca,Ca | V |
| Tschermakite (Ts) | Si₄ | Si₂,Al₂ | Al,Al | Mg,Mg,Mg | Ca,Ca | V |
| Pargasite (Prg) | Si₄ | Si₂,Al₂ | Mg,Al | Mg,Mg,Mg | Ca,Ca | Na |
Other: Ti, Mn. 23 Oxygen-basis.
XMapTools External Function: structural formula of Ca-amphibole with Fe³⁺ estimation. 23 Oxygen-basis. Not tested.
XMapTools External Function: structural formula of biotite. Fe₂O₃ added as possible input. Ti assumed to be ordered onto M2, Al onto M1, Mn added (ordered onto M12).
| End-member | T1(2) | T2(2) | M1(1) | M2(2) | A(1) |
|---|---|---|---|---|---|
| Phlogopite | Si,Si | Si,Al | Mg | Mg,Mg | K |
| Annite | Si,Si | Si,Al | Fe | Fe,Fe | K |
| Eastonite | Si,Si | Al,Al | Al | Mg,Mg | K |
| Siderophyllite | Si,Si | Al,Al | Al | Fe,Fe | K |
| Ti-biotite | Si,Si | Si,Al | Mg | Ti,Mg | K |
| Mn-biotite | Si,Si | Si,Al | Mn | Mn,Mn | K |
11 Oxygen-basis.
XMapTools External Function: structural formula of brucite. Version without loop & Fe₂O₃ as possible input. As₂O₃ is used instead of As₂O₅ in XMapTools 3.4.2. 1 Oxygen-basis.
XMapTools External Function: structural formula of chlorite. Ferri-sudoite added; Trincal & Lanari (2016) Clay Min. 51, 675–689. Fe₂O₃ added as possible input. 1 Input: XFe3 = Fe3/(Fe2+Fe3).
| End-member | T1(2) | T2(2) | M1(1) | M2-M3(4) | M4(1) |
|---|---|---|---|---|---|
| Amesite (Ame) | Si,Si | Al,Al | Al | Mg₄ | Al |
| Fe-Amesite (fAme) | Si,Si | Al,Al | Al | Fe₄ | Al |
| Clinochlore (Clc) | Si,Si | Si,Al | Mg | Mg₄ | Al |
| Daphnite (Dph) | Si,Si | Si,Al | Fe | Fe₄ | Al |
| Sudoite (Sud) | Si,Si | Si,Al | V | Mg₂,Al₂ | Al |
| FerriSudoite (fSud) | Si,Si | Si,Al | V | Mg₂,Fe₂ | Al |
| AlfreeChl (afchl) | Si,Si | Si,Si | Mg | Mg₄ | Mg |
14 Oxygen-basis.
XMapTools External Function: structural formula of chloritoid. Version without loop & Fe₂O₃ as possible input.
| End-member | T1(2) | T2(3) | M1(1) | M2(2) |
|---|---|---|---|---|
| Mg-chloritoid (Ctd) | Si,Si | Al₃ | Al | Mg,Mg |
| Fe-chloritoid (fCtd) | Si,Si | Al₃ | Al | Fe,Fe |
| Mn-chloritoid (mCtd) | Si,Si | Al₃ | Al | Mn,Mn |
12 Oxygen-basis.
XMapTools External Function: structural formula of chromite. Version without loop & Fe₂O₃ as possible input. Fe³⁺ estimated using Droop (1987). 32 Oxygen-basis. Not tested in this version.
XMapTools External Function: structural formula of clinopyroxene. Version without loop & Fe₂O₃ as possible input.
| End-member | T1(2) | M1(1) | M2(1) |
|---|---|---|---|
| Jadeite (Jd) | Si,Si | Al | Na |
| Diopside (Di) | Si,Si | Mg | Ca |
| Hedenbergite (Hd) | Si,Si | Fe | Ca |
| Ca-Tschermak (Cats) | Si,Al | Al | Ca |
6 Oxygen-basis.
XMapTools External Function: structural formula of clinopyroxene with Fe³⁺ estimation.
| End-member | T1(2) | M1(1) | M2(1) |
|---|---|---|---|
| Jadeite (Jd) | Si,Si | Al | Na |
| Diopside (Di) | Si,Si | Mg | Ca |
| Hedenbergite (Hd) | Si,Si | Fe² | Ca |
| Ca-Tschermak (Cats) | Si,Al | Al | Ca |
| Acmite (Acm) | Si,Si | Fe³ | Na |
6 Oxygen-basis.
XMapTools External Function: structural formula of cordierite. Version without loop & Fe₂O₃ as possible input.
| End-member | T1(2) | T2(3) | M1(2) |
|---|---|---|---|
| Mg-Cordierite (Crd) | Si₅ | Al₄ | Mg,Mg |
| Fe-Cordierite (fCrd) | Si₅ | Al₄ | Fe,Fe |
| Mn-Cordierite (mCrd) | Si₅ | Al₄ | Mn,Mn |
18 Oxygen-basis.
XMapTools External Function: structural formula of epidote. Version without loop & all iron as Fe³⁺.
| End-member | T1(3) | M1(1) | M2(1) | M3(1) | A1(2) |
|---|---|---|---|---|---|
| Epidote (Ep) | Si₃ | Al | Al | Fe | Ca,Ca |
| Fe-epidote (fEp) | Si₃ | Fe | Al | Fe | Ca,Ca |
| Zoisite (Zo) | Si₃ | Al | Al | Al | Ca,Ca |
| Mn-epidote (mEp) | Si₃ | Al | Al | Mn | Ca,Ca |
12.5 Oxygen-basis.
XMapTools External Function: structural formula of feldspar. Version without loop & Fe₂O₃ as possible input.
| End-member | T1(2) | T2(2) | M1(1) |
|---|---|---|---|
| Albite (Ab) | Si,Si | Si,Al | Na |
| Anorthite (An) | Si,Si | Al,Al | Ca |
| Sanidine (Sn) | Si,Si | Si,Al | K |
8 Oxygen-basis.
XMapTools External Function: structural formula of garnet. Version without loop & Fe₂O₃ as possible input.
| End-member | X(3) | Y(2) | O/T(3) |
|---|---|---|---|
| Almandine (Alm) | Fe₃ | Al₂ | (SiO₄)₃ |
| Pyrope (Prp) | Mg₃ | Al₂ | (SiO₄)₃ |
| Grossular (Grs) | Ca₃ | Al₂ | (SiO₄)₃ |
| Spessartine (Sps) | Mn₃ | Al₂ | (SiO₄)₃ |
12 Oxygen-basis.
XMapTools External Function: structural formula of garnet with Fe³⁺ estimation. Method of Droop (1987) implemented.
| End-member | X(3) | Y(2) | O/T(3) |
|---|---|---|---|
| Almandine (Alm) | Fe₃ | Al₂ | (SiO₄)₃ |
| Pyrope (Prp) | Mg₃ | Al₂ | (SiO₄)₃ |
| Grossular (Grs) | Ca₃ | Al₂ | (SiO₄)₃ |
| Spessartine (Sps) | Mn₃ | Al₂ | (SiO₄)₃ |
| Andradite (And) | Ca₃ | Fe₂ | (SiO₄)₃ |
12 Oxygen-basis.
XMapTools External Function: structural formula of ilmenite. Approximation of Fe³⁺ and Fe²⁺; works fine with Ca-Na-K ignored.
| End-member | Formula |
|---|---|
| Ilmenite (Ilm) | Fe(1)Ti(1)O(3) |
| Mn-Ilmenite (mIlm) | Mn(1)Ti(1)O(3) |
| Hematite (Hem) | Fe(2)O(3) |
3 Oxygen-basis.
XMapTools External Function: structural formula of magnetite. Version without loop & Fe₂O₃ as possible input. Normalized to 3 cations (assuming 4 oxygen). 4 Oxygen-basis.
XMapTools External Function: structural formula of olivine. Version without loop & Fe₂O₃ as possible input.
| End-member | Formula |
|---|---|
| Forsterite (Fo) | Mg(2)Si(1)O(4) |
| Fayalite (Fa) | Fe(2)Si(1)O(4) |
| Tephroite (Tep) | Mn(2)Si(1)O(4) |
4 Oxygen-basis.
XMapTools External Function: structural formula of orthopyroxene. Version without loop & Fe₂O₃ as possible input.
| End-member | T1(1) | M1(1) |
|---|---|---|
| Enstatite (En) | Si₂ | Mg₂ |
| Ferrosilite (Fs) | Si₂ | Fe₂ |
| Tschermak-px (Mgts) | Si,Al | Al,Mg |
3 Oxygen-basis.
XMapTools External Function: structural formula of rutile. Version without loop & Fe₂O₃ as possible input. 2 Oxygen-basis.
XMapTools External Function: structural formula of serpentine. Number of oxygen changed from 14.5 to 14. Ratio Si/(Si+Fe+Mg) added. Version without loop & Fe₂O₃ as possible input. 14 Oxygen-basis.
XMapTools External Function: structural formula of spinel. Version without loop & Fe₂O₃ as possible input. Fe³⁺ estimated using Droop (1987).
Al spinels (AlSpinel):
| End-member | X(1) | Y(2) |
|---|---|---|
| Hercynite (Hc) | Fe | Al₂ |
| Spinel (Spl) | Mg | Al₂ |
| Galaxite (Glx) | Mn | Al₂ |
| Gahnite (Ghn) | Zn | Al₂ |
Cr spinels (CrSpinel):
| End-member | X(1) | Y(2) |
|---|---|---|
| Chromite (Chr) | Fe | Cr₂ |
| Magnesiochromite (mChr) | Mg | Cr₂ |
| Nichromite (Nic) | Ni | Cr₂ |
Fe spinels (FeSpinel):
| End-member | X(1) | Y(2) |
|---|---|---|
| Magnesioferrite (Mfr) | Mg | Fe₂ |
| Magnetite (Mag) | Fe | Fe₂ |
| Ulvospinel (tSpl) | Ti | Fe₂ |
| Jacobsite (Jac) | Mn | Fe₂ |
| Franklinite (Frk) | Zn | Fe₂ |
| Trevorite (Tre) | Ni | Fe₂ |
Olivine spinels (OlSpinel):
| End-member | X(1) | Y(2) |
|---|---|---|
| Ringwoodite (Rwd) | Si | Mg₂ |
4 Oxygen-basis. Calculated assuming equipartition of Fe²⁺/Mg²⁺/Mn²⁺/Zn²⁺/Ni. Not tested.
XMapTools External Function: structural formula of staurolite. Version without loop & Fe₂O₃ as possible input.
| End-member | Z(11) | Y(2) | X(4) |
|---|---|---|---|
| Staurolite (St) | Si(4)Al(7) | Al,Al | Fe₄ |
| Mg-Staurolite (Mst) | Si(4)Al(7) | Al,Al | Mg₄ |
| Mn-Staurolite (MnSt) | Si(4)Al(7) | Al,Al | Mn₄ |
24 Oxygen-basis.
XMapTools External Function: structural formula of white mica (generic). Version without loop & Fe₂O₃ as possible input.
| End-member | T1(2) | T2(2) | M1(1) | M2(2) | A(1) |
|---|---|---|---|---|---|
| Celadonite-Mg (Cel) | Si,Si | Si,Si | V | Al,Mg | K |
| Celadonite-Fe (fCel) | Si,Si | Si,Si | V | Al,Fe | K |
| Muscovite (Ms) | Si,Si | Si,Al | V | Al,Al | K |
| Paragonite (Pg) | Si,Si | Si,Al | V | Al,Al | Na |
| Pyrophyllite (Prl) | Si,Si | Si,Si | V | Al,Al | V |
Not considered: Biotite (Si,Si | Si,Al | MgFe | MgFeTi | K). 11 Oxygen-basis.
Function headers for thermobarometry and other methods. Source code is available on the XMapTools GitHub repository. Several functions are taken from the PTtoolbox package of Joshua Laughton.
XMapTools External Function: amphibole thermobarometry. 23 Oxygen-basis.
Output variables:
XMapTools External Function: biotite thermobarometry. 22 Oxygen-basis.
Output variables:
XMapTools External Function: chlorite thermometry (all calibrations). Fe₂O₃ added as possible input. 14 Oxygen-basis.
Output variables:
XMapTools External Function: monazite geochronology.
Output variables:
Input variables: ThO₂, UO₂ and PbO in wt% (use the converter if your data were calibrated in a different format).
XMapTools External Function: quartz thermobarometry (all calibrations).
Output variables:
Input variables: TiO₂ (wt%).
XMapTools External Function: rutile thermobarometry (all calibrations).
Output variables:
Input variables: ZrO₂ (wt%).
XMapTools External Function: titanite thermometry (all calibrations).
Output variables:
XMapTools External Function: phengite barometry. 11 Oxygen-basis.
Output variables:
XMapTools External Function: zircon thermobarometry (all calibrations).
Output variables:
XMapTools External Function: Garnet-Hornblende thermometry. 12 Oxygen-basis for Garnet, 23 Oxygen-basis for Hornblende.
Output variables:
XMapTools External Function: Garnet-Biotite thermometry. 12 Oxygen-basis for Garnet, 11 Oxygen-basis for Biotite.
Output variables:
XMapTools External Function: Garnet-Chlorite thermometry. 12 Oxygen-basis for Garnet, 14 Oxygen-basis for Chlorite.
Output variables:
XMapTools External Function: Garnet-Clinopyroxene thermometry. 12 Oxygen-basis for Garnet, 6 Oxygen-basis for Clinopyroxene.
Output variables:
XMapTools External Function: Garnet-Cordierite thermometry. 12 Oxygen-basis for Garnet, 18 Oxygen-basis for Cordierite.
Output variables:
XMapTools External Function: Garnet-Ilmenite thermometry. 12 Oxygen-basis for Garnet, 3 Oxygen-basis for Ilmenite. Not tested.
Output variables:
XMapTools External Function: Garnet-Orthopyroxene thermometry. 12 Oxygen-basis for Garnet, 6 Oxygen-basis for Orthopyroxene.
Output variables:
XMapTools External Function: Garnet-Phengite thermometry. 12 Oxygen-basis for Garnet, 11 Oxygen-basis for Phengite.
Output variables:
Based on XMapTools 4.5 embedded documentation – Help file version 14.01.2024
This page describes the Images module, which provides tools to create and manage publication-ready images from XMapTools data.
Images are plots stored in the project file and can be accessed from the primary menu under the Image category. These plots can be created using the tools available in the Image menu. They can be edited at any time and changes are saved.
Four types of images can be created:
| Type | Description |
|---|---|
| Single plot image | A single map (e.g. Ca_apfu in garnet) |
| Multi-plot image | A mosaic of maps (e.g. SiO2, Al2O3, FeO, Na2O); can also be combined into a GIF |
| Multi-layer image (shared scale) | Multiple layers plotted with the same colour bar (e.g. XMg in biotite and garnet) |
| Multi-layer image (multi-scale) | Multiple layers, each plotted with its own colour scale and palette |
Use the main menu to view a map and, if necessary, adjust the colour contrast using the live histogram. Select Image >> Add Current Image from the main menu. A new image object will be created and placed in the Image category at the bottom of the primary menu.
You can view this image at any time by selecting Img (single-layer) #X from the primary menu (where X is the number of the image).
To edit the image, expand the image object and select the map. You can adjust the colour settings (borders, colour palette, etc.); changes are saved within the image object.
To export the image, select the image item in the primary menu and choose File > Save Image or Edit > Copy Image from the menu.
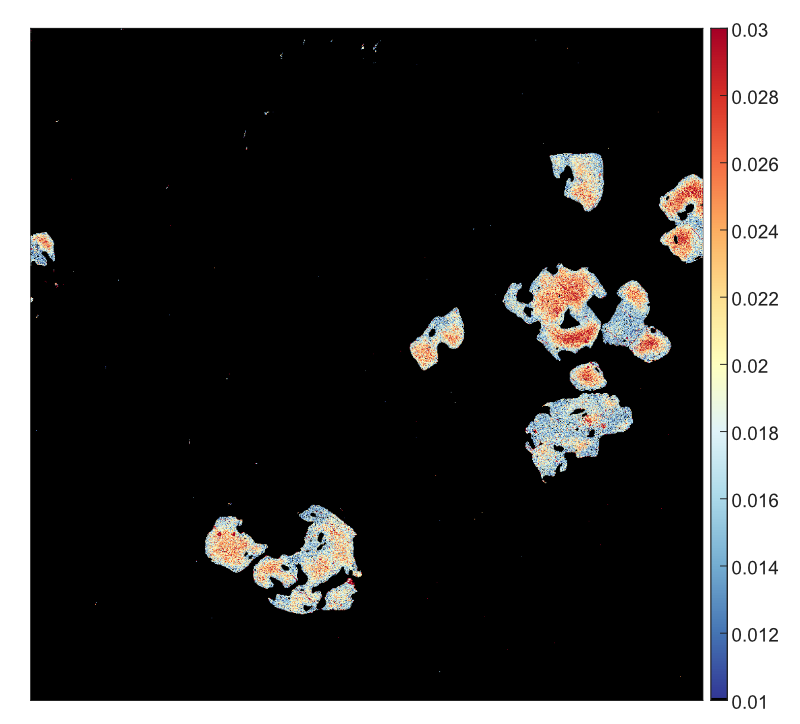
Figure 1: Example of an image generated with XMapTools for Ca (apfu) in garnet.
You can expand the Img (multi-plot) #X item in the primary menu and select the maps one by one to adjust the colour contrast. You can also change the colour palette in the Options panel.
WARNING
It is not recommended to use XMapTools with multi-selection mode enabled for normal operation — it should only be used to generate images.
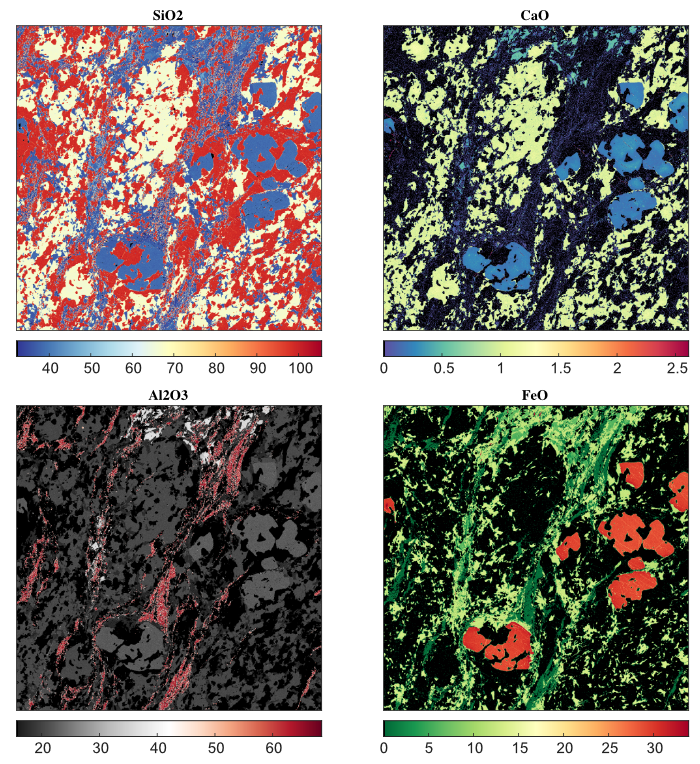
Figure 2: Example of a multi-plot image generated by XMapTools. Maps are expressed in oxide wt%. Note that each map is plotted using a different colour palette.
A GIF can be created from a multi-plot image. Select a multi-plot object, right-click on the name and select Save as GIF.
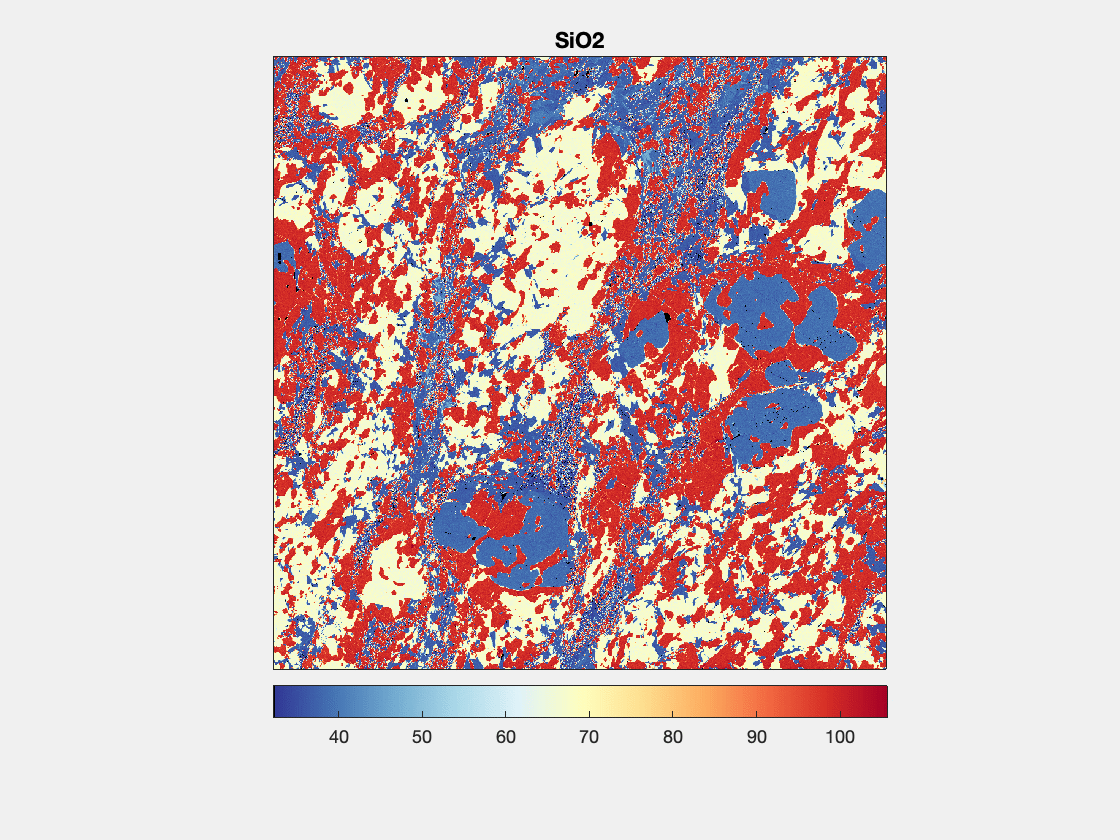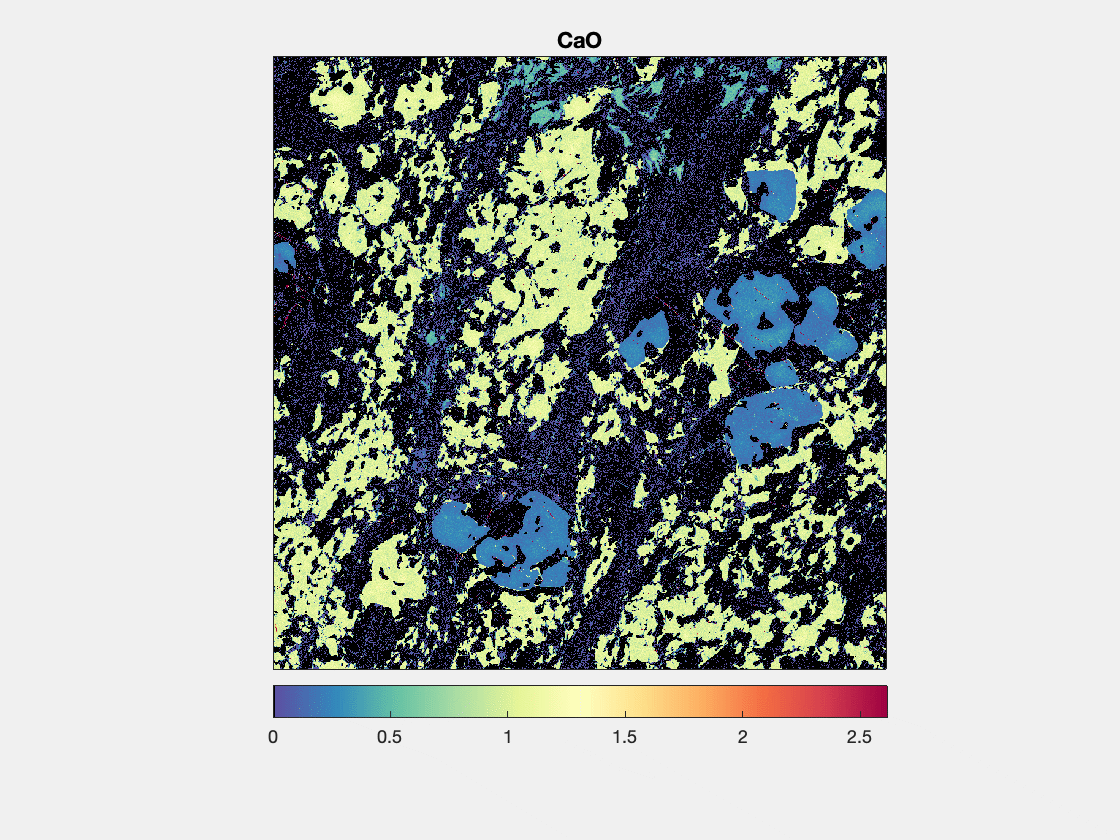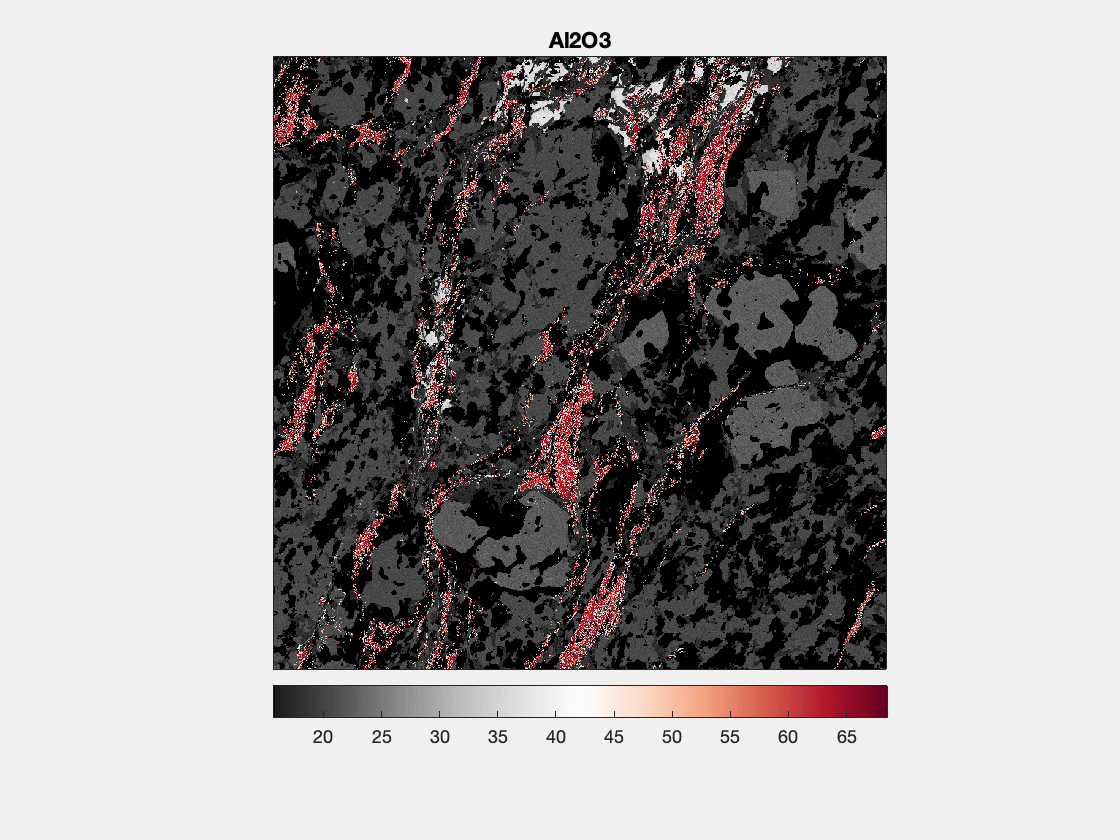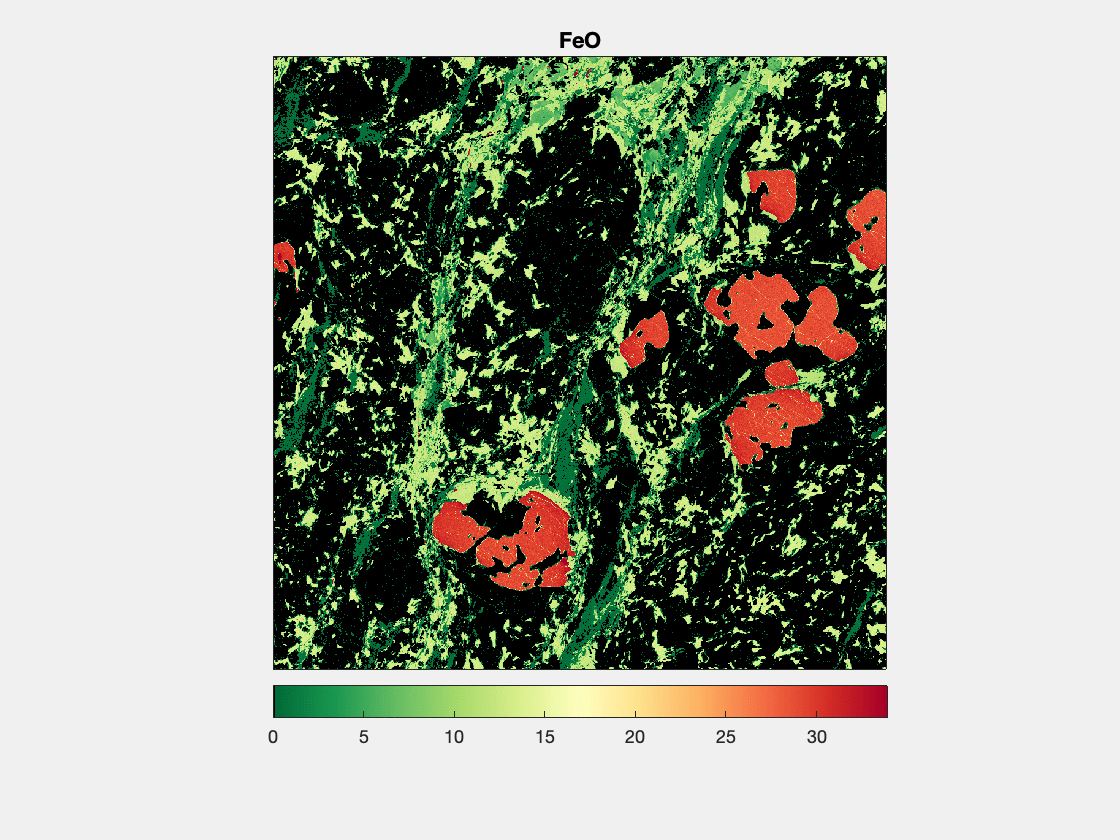
Figure 3: Example of GIF generated with the images shown in Figure 2.
You can expand the Img (multi-layer ShS) #X item in the primary menu and select the maps one by one to adjust the colour contrast. The same minimum and maximum values are applied to both images.
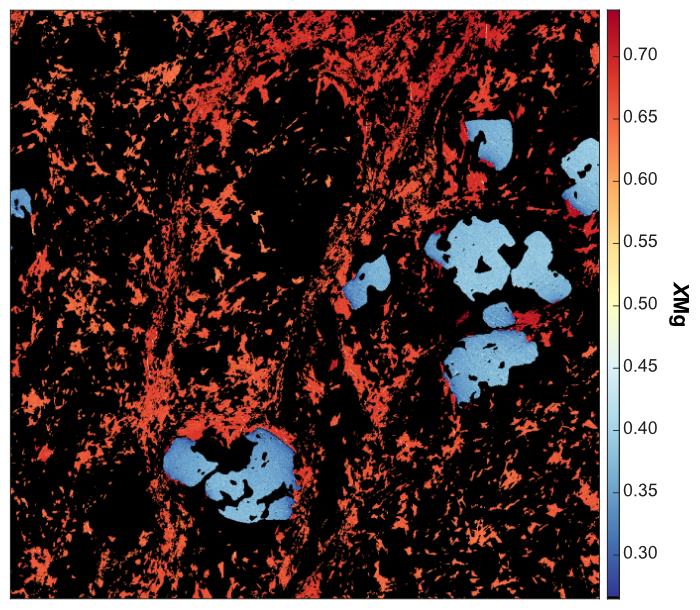
Figure 4: Example of multi-layer image with common colour scale for XMg in garnet and biotite.
It is strongly recommended to use a different colour map for each map, as multiple colour bars will be created.
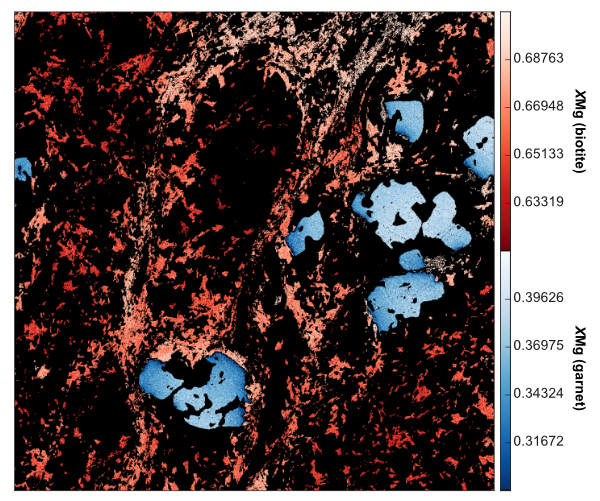
Figure 5: Example of multi-layer image with multi-scale for XMg in garnet and biotite. Note that the contrast is much better than on Figure 4 because two colour bars are used.
',41)])])}const u=t(l,[["render",m]]);export{p as __pageData,u as default}; diff --git a/.vitepress/dist/assets/doc-images.md.CLTo80uE.lean.js b/.vitepress/dist/assets/doc-images.md.CLTo80uE.lean.js new file mode 100644 index 0000000..d525bf1 --- /dev/null +++ b/.vitepress/dist/assets/doc-images.md.CLTo80uE.lean.js @@ -0,0 +1 @@ +import{_ as t,o as a,c as i,ae as o}from"./chunks/framework.DIXg7aqb.js";const p=JSON.parse('{"title":"Images","description":"","frontmatter":{},"headers":[],"relativePath":"doc-images.md","filePath":"doc-images.md"}'),l={name:"doc-images.md"};function m(r,e,n,s,g,d){return a(),i("div",null,[...e[0]||(e[0]=[o("",41)])])}const u=t(l,[["render",m]]);export{p as __pageData,u as default}; diff --git a/.vitepress/dist/assets/doc-import-maps.md.B-cz6IVV.js b/.vitepress/dist/assets/doc-import-maps.md.B-cz6IVV.js new file mode 100644 index 0000000..f5be129 --- /dev/null +++ b/.vitepress/dist/assets/doc-import-maps.md.B-cz6IVV.js @@ -0,0 +1 @@ +import{_ as e,o as d,c as a,ae as o}from"./chunks/framework.DIXg7aqb.js";const h=JSON.parse('{"title":"Import Maps Tool","description":"","frontmatter":{},"headers":[],"relativePath":"doc-import-maps.md","filePath":"doc-import-maps.md"}'),r={name:"doc-import-maps.md"};function i(n,t,l,s,m,p){return d(),a("div",null,[...t[0]||(t[0]=[o('Based on XMapTools 4.5 embedded documentation – Help file version 14.01.2024 by M. Tedeschi & P. Lanari
This page describes the Import Tool, which can be used to import maps, set the destination, and make basic corrections. Closing the Import Tool will stop the import process (no data will be imported).
The Select ![]() button allows a set of map files to be imported. Select compatible files from the pick map file(s) pop-up window. Multiple files can be selected together. If a selected file cannot be imported (e.g. due to an incompatible format), it will be skipped during the import.
button allows a set of map files to be imported. Select compatible files from the pick map file(s) pop-up window. Multiple files can be selected together. If a selected file cannot be imported (e.g. due to an incompatible format), it will be skipped during the import.
Map files must have the *.txt, *.asc, *.dat or *.csv extension, no header, and a name compatible with the XMapTools default element names for identification. The default lists of compatible element and oxide names are listed below.
Selected map files are listed in the main table. Additional maps can be added by pressing the Add ![]() button.
button.
| Column | Description |
|---|---|
| File | Contains the file names |
| Map Name | Contains the name of the corresponding element in the database |
| Type | Element or oxide |
| Data | Intensity or wt% |
| Special | EDS or WDS(?); in the case of WDS(?) a DTC is automatically selected |
| DTC | Dead time correction |
| OC | Orientation correction (do not use for EPMA and LA-ICPMS using the approach of Markmann et al. 2024) |
| Destination | In XMapTools, can be: Intensity, Quanti, Merged, Other; drop-down menu, can be edited |
| Action | Keep or eliminate; drop-down menu, can be edited |
Settings such as the dwell time can be changed in the Corrections section.
Press Import data ![]() to import the selected maps into XMapTools after the corrections have been applied.
to import the selected maps into XMapTools after the corrections have been applied.
XMapTools supports all standard element symbols from H (1.00797) to Lr (260), including all lanthanides and actinides. Map files should be named with the corresponding element symbol for automatic identification.
The following oxides are supported (non-exhaustive list of common oxides):
| Oxide | Element | Conversion Factor |
|---|---|---|
| SiO2 | Si | 0.467 |
| TiO2 | Ti | 0.600 |
| Al2O3 | Al | 0.529 |
| FeO | Fe | 0.777 |
| Fe2O3 | Fe | 0.699 |
| MnO | Mn | 0.774 |
| MgO | Mg | 0.603 |
| CaO | Ca | 0.715 |
| Na2O | Na | 0.742 |
| K2O | K | 0.830 |
| P2O5 | P | 0.436 |
| Cr2O3 | Cr | 0.684 |
| NiO | Ni | 0.786 |
| ZnO | Zn | 0.803 |
| ZrO2 | Zr | 0.740 |
| BaO | Ba | 0.896 |
| SrO | Sr | 0.846 |
| H2O | H | 0.112 |
| CO2 | C | 0.273 |
| SO3 | S | 0.400 |
Additional oxides are supported including rare earth element oxides (La2O3, Ce2O3, CeO2, Nd2O3, Sm2O3, Eu2O3, Gd2O3, Dy2O3, Er2O3, Yb2O3, Lu2O3, etc.), high field strength element oxides (Nb2O5, Ta2O5, HfO2, ThO2, UO2, U3O8, WO3, etc.) and many more.
',21)])])}const b=e(r,[["render",i]]);export{h as __pageData,b as default}; diff --git a/.vitepress/dist/assets/doc-import-maps.md.B-cz6IVV.lean.js b/.vitepress/dist/assets/doc-import-maps.md.B-cz6IVV.lean.js new file mode 100644 index 0000000..8570c1d --- /dev/null +++ b/.vitepress/dist/assets/doc-import-maps.md.B-cz6IVV.lean.js @@ -0,0 +1 @@ +import{_ as e,o as d,c as a,ae as o}from"./chunks/framework.DIXg7aqb.js";const h=JSON.parse('{"title":"Import Maps Tool","description":"","frontmatter":{},"headers":[],"relativePath":"doc-import-maps.md","filePath":"doc-import-maps.md"}'),r={name:"doc-import-maps.md"};function i(n,t,l,s,m,p){return d(),a("div",null,[...t[0]||(t[0]=[o("",21)])])}const b=e(r,[["render",i]]);export{h as __pageData,b as default}; diff --git a/.vitepress/dist/assets/doc-laicpms.md.DD1IZLH8.js b/.vitepress/dist/assets/doc-laicpms.md.DD1IZLH8.js new file mode 100644 index 0000000..e64d71b --- /dev/null +++ b/.vitepress/dist/assets/doc-laicpms.md.DD1IZLH8.js @@ -0,0 +1,40 @@ +import{_ as a,o as t,c as n,ae as i}from"./chunks/framework.DIXg7aqb.js";const m=JSON.parse('{"title":"XMapTools documentation for LA-ICP-MS","description":"","frontmatter":{},"headers":[],"relativePath":"doc-laicpms.md","filePath":"doc-laicpms.md"}'),o={name:"doc-laicpms.md"};function s(r,e,l,p,d,c){return t(),n("div",null,[...e[0]||(e[0]=[i(`The importer module should be able to read files in the following format. Note that some format require a conversion before to be imported in XMapTools.
The date and time are read from the third row.
D:\\Agilent\\ICPMH\\1\\DATA\\Name\\map.b\\map.d
+Intensity Vs Time,CPS
+Acquired : 2022-01-24 11:11:45 AM using Batch map.b
+Time [Sec],Li7,Na23,Mg25
+0.6196,50.00,82227.81,0.00
+1.1516,50.00,79613.57,0.00
+[...]The Date and time are read from the first row.
Standards run :02-27-2024 11:16:40 AM;
+Software: [...]
+Configuration: [...]
+S-SQ-N%2FA: [...]
+RF Generator: [...]
+Ion Optics: [...]
+Vacuum: [...]
+Detector: [...]
+Cooling System: [...]
+Power Supply: [...]
+Gas Supply: [...]
+Pulse Counting: [...]
+
+Time,7Li,23Na,24Mg,25Mg,
+,dwell time=0.01;xcal factor=59347.84333,dwell time=0.001;xcal factor=72480.60121,dwell time=0.01;xcal factor=73115.42471
+0.01326,400.006400102402,89317.9719802497,0
+0.41544,0,67180.0425139374,0
+0.81759,0,76231.74450329,0
+[...]Warning: This file must be converted to CSV format using the 'Tools > Convert FIN2 to CSV' option in the converter.
The Date and time are read from the first row.
Finnigan MAT ELEMENT Raw Data
+Friday, July 04, 2025 11:31:12
+2025_07_04_Name1.FIN
+231
+0
+16,16,16
+CPS
+Time,Li7,B10,B11
+3.236000,340192.875000,585.555542,887.700012
+4.044000,342415.125000,459.555542,934.299988
+[...]The date and time are read from the file timestamp.
Intensity Vs Time, Counts per Second
+Time in Seconds ,Mg/25,Al/27,P/31,Ca/42
+
+0.,200.0016000128,1000.0400016001,3067.0428905946,31639.99295109
+0.28,200.0016000128,1600.102406554,3267.0935668927,30837.992406645
+[...]The Converter for LA-ICP-MS data can read LA-ICP-MS mapping data natively and generate intensity and standard maps for XMapTools (Markmann et al. 2024).
Main steps to import LA-ICP-MS data:
The import tools are located at the top of the module. The data format can be selected from the first dropdown menu and the file format (single or multiple) from the second dropdown menu. The data from a measurement session can be contained in a single file or multiple files. If multiple files are selected, it is recommended that you select the last file first and then hold SHIFT while selecting the first file, not vice versa. For the laser, deselect the log file option if no compatible log file is available. In this case, the Log Generator module will appear when the Load Datafiles button is pressed.
Once appropriate options have been selected, press the Load Datafiles button to select: (1) a single data file from the mass spectrometer containing timestamps and intensity data (e.g. Data.csv) or multiple files (see warning above), and (2) if the Log File option is enabled, immediately after a corresponding log file (e.g. Log.csv) from the laser system (tested with RESONETICS only).
The value of the time shift can be adjusted to synchronise the data file with the log file. In the main figure, the total signal is plotted together with the laser on/off signals given in the log file (vertical red lines). The shift value is automatically adjusted by XMapTools when the data is imported. If necessary, adjust the value until both signals are synchronised.
Once the value of the time shift is optimised, press the Extract Integrations button to automatically extract all the analyses listed in the log file. They should appear in the Integration tree menu on the left of the window after extraction.
The integrations for fitting a background correction are automatically selected for each measurement. They are listed in a tree menu (integration menu) located on the left and can be plotted by selecting an item under the category Background.
To view and edit integrations, select the first measurement in the tree menu. The display is automatically adjusted to zoom in on the selected integration. If you select a different measurement in the tree menu, the integration will be displayed in the centre of the plot. The program automatically excludes a fraction of sweeps at the beginning and at the end of each background measurement (default value is 10 %). This value can be changed manually if required. It is also possible to manually edit the duration of an integration by changing the limit values available as Sweep (min and max).
Select a method for fitting the background in the BACKGROUND section below the plot. It is recommended to display the signal Raw_Sum and to adjust the display to show the entire background signal. The following functions are available: (1) linear, (2) polynomial, (3) step function, (4) spline. Step function is recommended for background correction. Press the button Apply to apply the background correction. You cannot change the background correction once it has been applied.
The background-corrected (BackCorr) signal is displayed when a background correction has been applied. Corrected data are available from the Plot menu. Select the measurement to be used as the primary standard from the PRIMARY STANDARD dropdown menu. The integration name should match the name of a standard file containing the composition of this standard (e.g. NIST612 or GSD-1g). It is also possible to add custom files using the options available in the lower left panel, but this process must be repeated each time the converter is used. Integrations can be edited using the same strategy as for background integration (see above).
Then select an interpolation method. A spline function is usually the best method to approximate instrument drift during the measurement, especially when reconstructing maps (Markmann et al. 2024). A step function can also be used in some cases, but this can result in sharp transitions between lines in the final map. Press the Apply button to validate and interpolate the primary standard signal. If the standard is not automatically recognised, a window will open with a list of all available standards. It is possible to define additional interpolations to be used as primary standards. After pressing the Apply button it is possible to define a new interpolation for the same material or for a different material by selecting the Yes option. If you do not wish to define any further interpolations to be used as primary standards, select No (Continue).
Select the measurements to be used as the secondary standard using the dropdown menu under SECONDARY STANDARD. Integrations can be edited using the same strategy as for background integration and primary standard (see above). Use the Int. Std dropdown menu to select an element to be used as a reference for calculating the composition of the secondary standard. This choice has no effect at this stage on the map calibration performed in XMapTools (another element can be used). But here it is possible to quickly check the calibration using the secondary standard for several elements used as internal standard.
Maps can be generated from raster measurements after background correction, adjustment of the primary reference material(s) and checking of the secondary reference material(s). Use the dropdown menu in MAPS (SCANS) to select the raster measurements to be used to construct the map. Integrations cannot be edited. Press the Apply button to validate and generate the maps. Three buttons become available.
Markmann, T.A., Lanari, P., Piccoli, F., Pettke, T., Tamblyn, R., Tedeschi, M., Lueder, M., Kunz, B., Riel, N., and Laughton, J. (2024). Multi-phase quantitative compositional mapping by LA-ICP-MS: analytical approach and data reduction protocol implemented in XMapTools. Chemical Geology, 646, 121895.
For more detailed information on the LA-ICPMS Converter, refer to the embedded documentation available within the program.
When the log generator module is used the time shift should be set to zero.
Intensity maps obtained by LA-ICPMS can be calibrated to element/oxide compositions. The following steps can be followed:
Only standard maps generated by the Converter for LA-ICPMS data can be imported and used to calibrate LA-ICPMS maps in XMapTools 4.
The button Import (Import Maps for Standards (from file)) is used to open a window in which the file 'MapStandards_Import.mat' must be selected.
Once standard data are imported they become available under the category Standards (Maps) in the Secondary Menu. Select an element to plot a map corresponding to the intensity of the primary standard material.
As for EPMA, the calibration tool for LA-ICPMS data requires a mask file, so the Calibrate button is only available when a mask file is selected in the Secondary Menu.
The button Calibrate (Open LA-ICP-MS Calibration Tools) opens the calibration assistant.
For more detailed information on the Calibration Assistant, refer to the embedded documentation accessible from the assistant.
The approach implemented in XMapTools 4, described in Markmann et al. (2024), provides a mineral calibration module based on the reference composition of an element (internal standard). In contrast to other programs, the composition of the reference element in the mineral can be variable.
Markmann, T.A., Lanari, P., Piccoli, F., Pettke, T., Tamblyn, R., Tedeschi, M., Lueder, M., Kunz, B., Riel, N., and Laughton, J. (2024). Multi-phase quantitative compositional mapping by LA-ICP-MS: analytical approach and data reduction protocol implemented in XMapTools. Chemical Geology, 646, 121895.
This mode is useful when an element is chemically unzoned in the selected mineral and you want to use it as an internal standard (e.g. SiO₂ in garnet).
You can use the Internal Composition Converter tool to convert the composition of the internal standard into µg/g of element. Enter a value in oxide wt% and the converter will automatically convert this value to µg/g. Note that the value in the Fixed Composition field changes when a new conversion is performed; there is no need to copy the value from one field to the other.
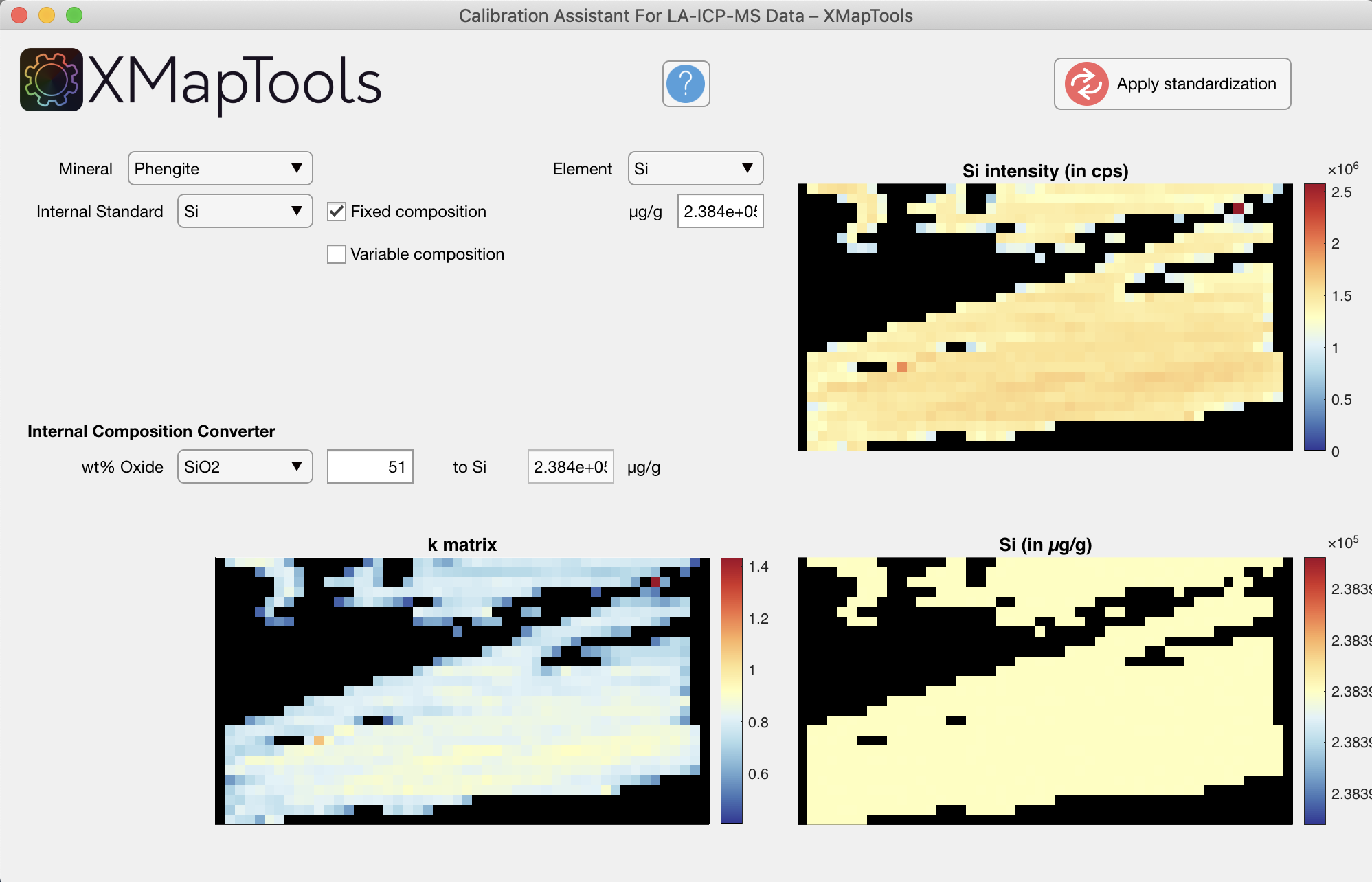
Figure 1: Calibration Assistant for LA-ICPMS data. Example of internal calibration of phengite using Si as internal standard and a reference composition of SiO2 = 51 wt%. The value of Si = 2.384e+05 µg/g has been automatically set in the "Fixed Composition" field. Two maps are displayed at the bottom: the k-matrix calculated using the internal standard and the calibrated map for Si expressed in µg/g.
Press the Apply Standardisation button to generate the compositional maps available in the Quantity category of the Primary Menu. The maps are expressed in µg/g of elements. You can convert them to wt% of oxides using the internal converter: right-click on a Quanti file and select Convert.
TIP
After converting the maps, the name of the maps can be changed (this is not done by the converter).
This mode is optimal for minerals that are zoned in all major elements (e.g. phengite). Average intensities of several ROIs can be extracted from the map and assigned to different compositions.
The Pick a ROI (circle) ![]() button activates the mode to draw a ROI with a circle shape. Draw a circle on the map in a region of constant intensity.
button activates the mode to draw a ROI with a circle shape. Draw a circle on the map in a region of constant intensity.
Set the average composition in wt% oxide in the Internal Composition Converter tool. Then click on the first and second row of Pt-1 in the table; values are automatically added.
Move the ROI to another area (e.g. with lower intensity/composition values), adjust the value for wt% oxide, and click in both rows of the table to create Pt-2.
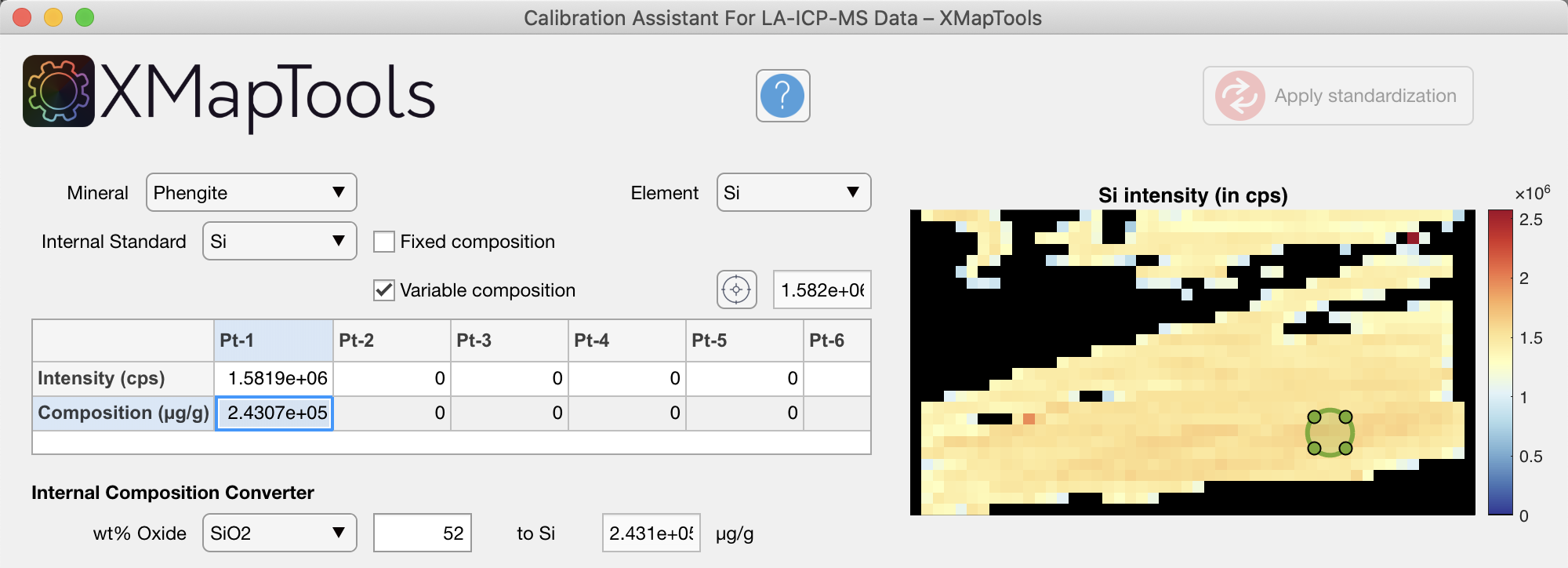
Figure 2: Example of internal calibration of phengite using Si as internal standard and a variable reference composition. A value of SiO2 = 52 wt% is assigned to the phengite core (Pt-1).
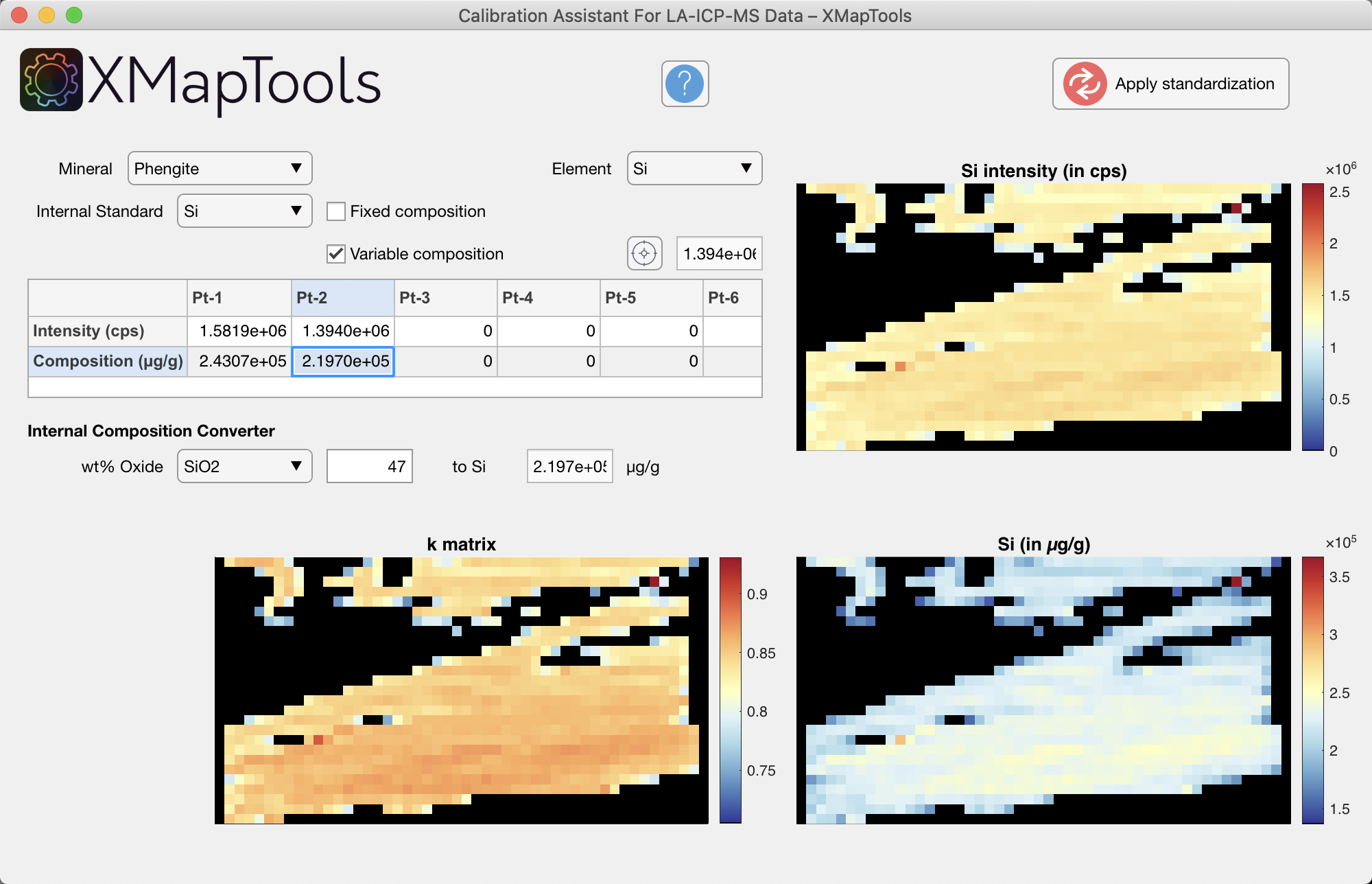
Figure 3: Example with a value of SiO2 = 47 wt% assigned to the muscovite rim (Pt-2). Two maps are shown below: the k-matrix calculated using the internal standard with variable composition and the calibrated map for Si expressed in µg/g. Note that compared to Figure 1, Si is not homogeneous throughout the mica.
Press the Apply Standardisation button to generate the compositional maps available in the Quantity category of the Primary Menu. The maps are expressed in µg/g of elements. You can convert them to wt% of oxides using the internal converter: right-click on a Quanti file and select Convert.
TIP
After converting the maps, the name of the maps can be changed (this is not done by the converter).
The button Spider (Open Spider Module) opens the module Spider Plot. This button is only available when a quanti file or any map within a quanti file is selected in the primary menu.
For more detailed information on the Spider Module, see the dedicated Spider Module page.
`,82)])])}const u=a(o,[["render",s]]);export{m as __pageData,u as default}; diff --git a/.vitepress/dist/assets/doc-laicpms.md.DD1IZLH8.lean.js b/.vitepress/dist/assets/doc-laicpms.md.DD1IZLH8.lean.js new file mode 100644 index 0000000..048949b --- /dev/null +++ b/.vitepress/dist/assets/doc-laicpms.md.DD1IZLH8.lean.js @@ -0,0 +1 @@ +import{_ as a,o as t,c as n,ae as i}from"./chunks/framework.DIXg7aqb.js";const m=JSON.parse('{"title":"XMapTools documentation for LA-ICP-MS","description":"","frontmatter":{},"headers":[],"relativePath":"doc-laicpms.md","filePath":"doc-laicpms.md"}'),o={name:"doc-laicpms.md"};function s(r,e,l,p,d,c){return t(),n("div",null,[...e[0]||(e[0]=[i("",82)])])}const u=a(o,[["render",s]]);export{m as __pageData,u as default}; diff --git a/.vitepress/dist/assets/doc-options.md.BUc_pidR.js b/.vitepress/dist/assets/doc-options.md.BUc_pidR.js new file mode 100644 index 0000000..f6e092e --- /dev/null +++ b/.vitepress/dist/assets/doc-options.md.BUc_pidR.js @@ -0,0 +1 @@ +import{_ as o,o as a,c as t,ae as r}from"./chunks/framework.DIXg7aqb.js";const u=JSON.parse('{"title":"Options","description":"","frontmatter":{},"headers":[],"relativePath":"doc-options.md","filePath":"doc-options.md"}'),i={name:"doc-options.md"};function l(n,e,s,p,h,c){return a(),t("div",null,[...e[0]||(e[0]=[r('Based on XMapTools 4.5 embedded documentation – Help file version 11.01.2024
This page describes the tools available in the Options section.

Figure 1: Option tools in XMapTools.
Use these options to change the colour palette, change the resolution of the colour map (number of colours), change the scale of the colour bar to logarithmic and add a black or white layer at the bottom and/or top of the colour palette.
![]() Set the colour palette to be used for the colour bar throughout XMapTools.
Set the colour palette to be used for the colour bar throughout XMapTools.
The recommended colour palettes are largely inspired by www.ColorBrewer.org by Cynthia A. Brewer, Geography, Pennsylvania State University. The following abbreviations are used in the drop-down menu: Colourblind friendly (C), Linear (L), Divergent (D) and Print friendly (P).
Default palettes:
Scientific colour maps by Simon Garnier (viridisLite):
Scientific colour maps by Fabio Crameri:
![]() Check the Log (Use a log colormap) box to set a logarithmic axis for the colour bar. If this option is not selected, a linear scale is used instead.
Check the Log (Use a log colormap) box to set a logarithmic axis for the colour bar. If this option is not selected, a linear scale is used instead.
![]() Sets the number of colours used in the colormap (i.e. the resolution of the colour bar). Colour palettes defined with more colours are resampled. The default value is 256.
Sets the number of colours used in the colormap (i.e. the resolution of the colour bar). Colour palettes defined with more colours are resampled. The default value is 256.
![]() Adds a black or white layer at the bottom (lower) and/or top (upper) of the colour palette. When selected, the layer can be set to either black or white in the corresponding drop down menu.
Adds a black or white layer at the bottom (lower) and/or top (upper) of the colour palette. When selected, the layer can be set to either black or white in the corresponding drop down menu.
![]() If selected, negative values are displayed in the plotted map. By default, XMapTools ignores negative values in plots.
If selected, negative values are displayed in the plotted map. By default, XMapTools ignores negative values in plots.
The Apply Auto Contrast option activates automatic contrast enhancement each time a new map is displayed.
![]() Defines the value of the median filter to be applied when plotting a new 3D surface (see main menu Edit > Plot 3D surface). Changing this option will not update plots already generated.
Defines the value of the median filter to be applied when plotting a new 3D surface (see main menu Edit > Plot 3D surface). Changing this option will not update plots already generated.
This is the current resolution of the programme. Please submit this value if you are experiencing display problems with XMapTools.
',33)])])}const d=o(i,[["render",l]]);export{u as __pageData,d as default}; diff --git a/.vitepress/dist/assets/doc-options.md.BUc_pidR.lean.js b/.vitepress/dist/assets/doc-options.md.BUc_pidR.lean.js new file mode 100644 index 0000000..1f3b734 --- /dev/null +++ b/.vitepress/dist/assets/doc-options.md.BUc_pidR.lean.js @@ -0,0 +1 @@ +import{_ as o,o as a,c as t,ae as r}from"./chunks/framework.DIXg7aqb.js";const u=JSON.parse('{"title":"Options","description":"","frontmatter":{},"headers":[],"relativePath":"doc-options.md","filePath":"doc-options.md"}'),i={name:"doc-options.md"};function l(n,e,s,p,h,c){return a(),t("div",null,[...e[0]||(e[0]=[r("",33)])])}const d=o(i,[["render",l]]);export{u as __pageData,d as default}; diff --git a/.vitepress/dist/assets/doc-project-import.md.Dwd43zVv.js b/.vitepress/dist/assets/doc-project-import.md.Dwd43zVv.js new file mode 100644 index 0000000..9fd7915 --- /dev/null +++ b/.vitepress/dist/assets/doc-project-import.md.Dwd43zVv.js @@ -0,0 +1 @@ +import{_ as t,o as a,c as o,ae as i}from"./chunks/framework.DIXg7aqb.js";const h=JSON.parse('{"title":"Project & Import","description":"","frontmatter":{},"headers":[],"relativePath":"doc-project-import.md","filePath":"doc-project-import.md"}'),s={name:"doc-project-import.md"};function r(n,e,l,c,m,p){return a(),o("div",null,[...e[0]||(e[0]=[i('Based on XMapTools 4.5 embedded documentation
This page describes the tools available in the Project & Import workspace. Each section is described separately below.
Figure 1: Project & Import tools in XMapTools.
The progress of a session can be saved to a project file (*.mat) and restored later. All current data and settings will be saved. Project files are generally compatible with all new versions of XMapTools. Project management tools described in the following are available in the Project & Import workspace or from the File menu.
INFO
XMapTools 4 uses a different strategy for internal data organisation and storage. Projects created with previous versions of XMapTools are not compatible with the new format and a conversion step is required. When a project file created with XMapTools 3 is opened in XMapTools 4, an automatic conversion to the new format is performed. Further explanation can be found here.
The Save (Save Project) ![]() button saves the active project as a MATLAB M-file containing all variables used by XMapTools. If no project has been saved, a new project will be saved.
button saves the active project as a MATLAB M-file containing all variables used by XMapTools. If no project has been saved, a new project will be saved.
The Save Project As... ![]() button saves the active project as a new MATLAB M-file.
button saves the active project as a new MATLAB M-file.
![]() Display the name or the full path of the active project (if saved) depending on the selected option in the dropdown menu (File or Path).
Display the name or the full path of the active project (if saved) depending on the selected option in the dropdown menu (File or Path).
TIP
Remember that the current working directory changes automatically when a new project file is opened. It is also possible to change the working directory using the main menu: XMapTools > Set Working Directory.
Several tools are available to convert raw data into an XMapTools-friendly format and generate mosaics.
Data formats currently supported:
EPMA - JEOL (SUN) – JEOL microprobes running on SUN-OS
EPMA - JEOL (WIN) – JEOL microprobes running on WINDOWS
EPMA - CAMECA – for recent CAMECA microprobes
The Open XMapTools' LA-ICP-MS converter ![]() button opens the converter for LA-ICPMS data. A detailed description of how to convert LA-ICP-MS data can be found in the LA-ICP-MS documentation.
button opens the converter for LA-ICPMS data. A detailed description of how to convert LA-ICP-MS data can be found in the LA-ICP-MS documentation.
XMapTools can combine maps to create mosaics. There are two types of mosaic: grid mosaic, where the maps are stitched together in a grid, and real mosaic, where the maps are merged in the same coordinate system.
How to create a mosaic using a predefined grid:
01_Map34, 02_Map4, 03_Map1 to define the order before importing-), the folder will be skipped and not imported when the mosaic is created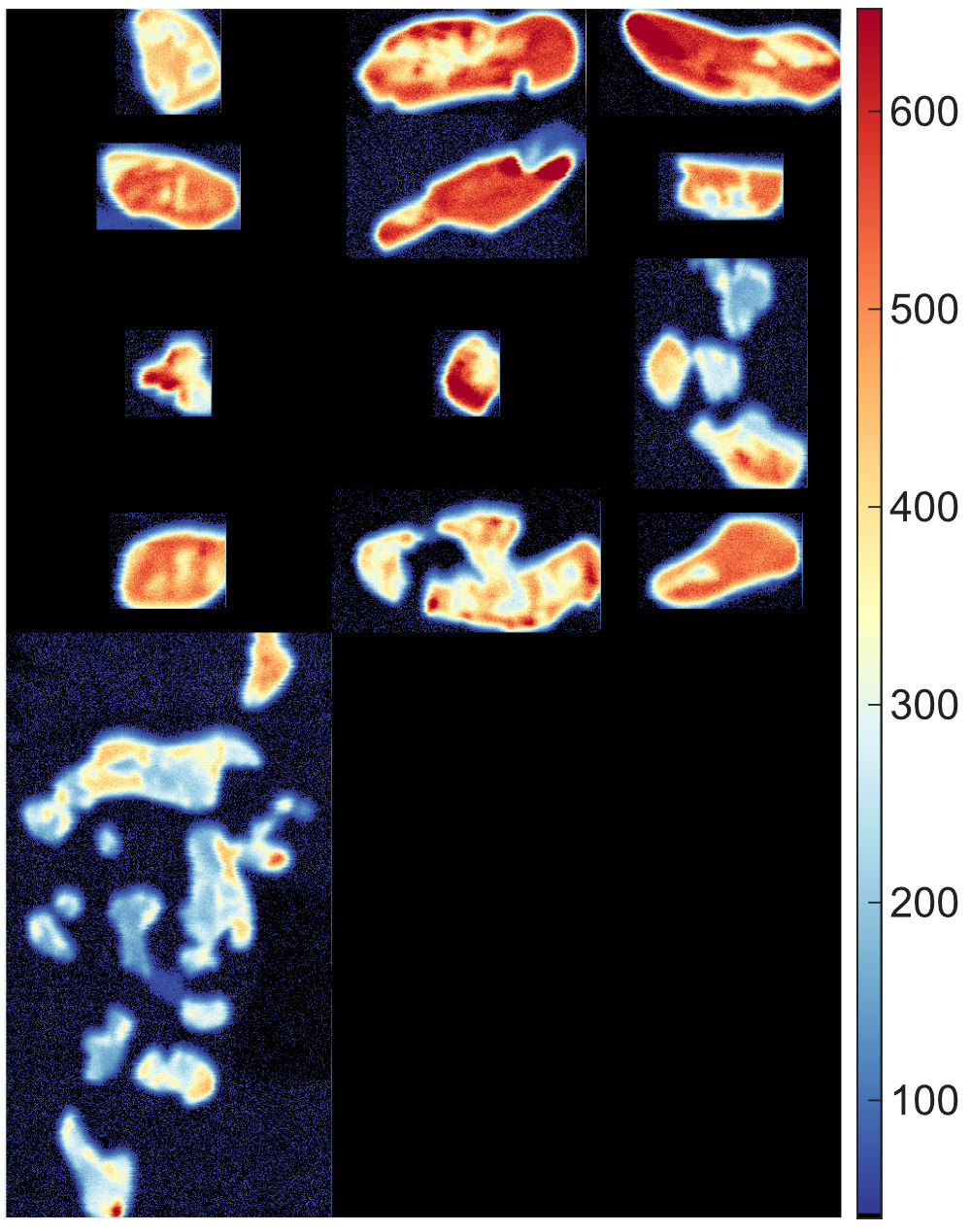
Figure: This is an example of a mosaic grid for 13 maps, arranged in three columns (map: Pb in counts). This allows the maps to be merged and displayed with the same colour scale.
How to create a referenced mosaic:
01_Map34, 02_Map4, 03_Map1 to define the order before importing. Each folder must contain a Standards.txt file containing the map coordinates. If a Classification.txt file exists, it will be skipped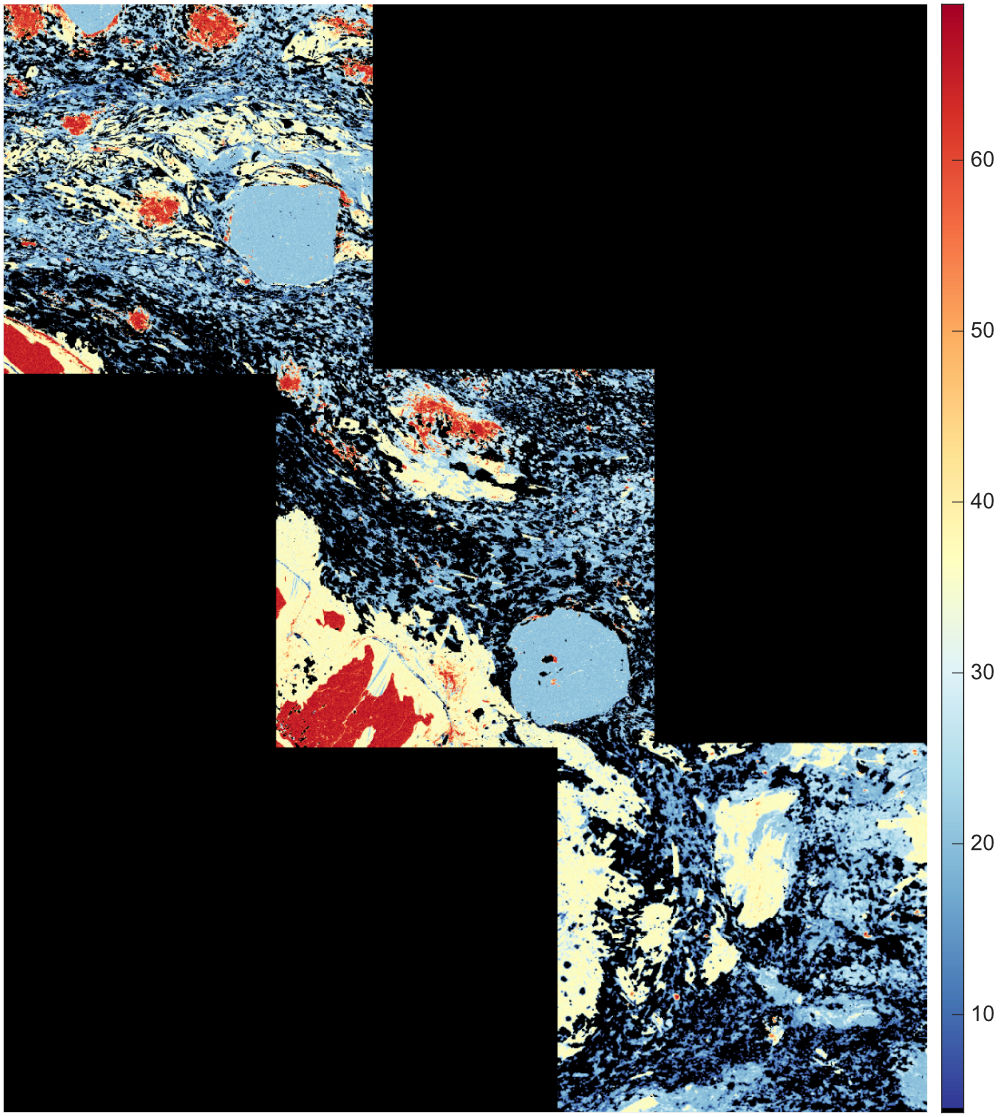
Figure: This is an example of a real mosaic for 3 maps, arranged using the map coordinates (map: Al2O3 in wt%).
This section describes the tools available for importing maps and other images into XMapTools. See also the dedicated Import Maps Tool documentation.
Format: Map files must have the extension *.txt, *.asc, *.dat or *.csv, no header and a name compatible with the default element names of XMapTools for identification.
*.jpg and *.png. It is recommended to select the files from the last one to the first one while pressing MAJ. Importing large datasets can take several minutes.This section describes the display options for the map being viewed. These options are saved for each project.
![]() Set the map resolution corresponding to the pixel size, also known as the step size on EPMA instruments. The default value is 20µm (arbitrary). The resolution value is automatically calculated from the map coordinates defined in Standards.txt when internal standards (spot analyses) are imported.
Set the map resolution corresponding to the pixel size, also known as the step size on EPMA instruments. The default value is 20µm (arbitrary). The resolution value is automatically calculated from the map coordinates defined in Standards.txt when internal standards (spot analyses) are imported.
![]() Activate and display the scale bar using the pixel size defined in Resolution. Two options are available from the drop down menu: white (default) or black.
Activate and display the scale bar using the pixel size defined in Resolution. Two options are available from the drop down menu: white (default) or black.
The Rotate view by 90° ![]() button can be used to adjust the orientation of the map. The rotation angle is displayed next to the button with possible values: 0 (default orientation), 90, 180 or 270. Note that some of the display features, such as drag-for-zoom, are not available when a rotation is applied.
button can be used to adjust the orientation of the map. The rotation angle is displayed next to the button with possible values: 0 (default orientation), 90, 180 or 270. Note that some of the display features, such as drag-for-zoom, are not available when a rotation is applied.
Based on XMapTools 4.5 embedded documentation – Help file version 14.01.2024
This page describes the Sampling Tools. These tools are available at any stage of the data procedure and apply to the map shown in the main figure.
The following sampling functions are available from the Sampling menu or from the Sampling section of the live output display, located below the live histogram:
| Tool | Icon | Description |
|---|---|---|
| Circle | Average value and standard deviation of selected pixels | |
| Area (polygon) | Average value and standard deviation of selected pixels | |
| Transect | Composition/intensity changes along a path | |
| Strip | Averaged compositional/intensity changes along a strip |
Once a ROI has been selected, the results are displayed in the Sampling Viewer to the right of the main window. Data can be saved using the tools available in the Sampling menu under 'Save Results'. The 'Single Map' option saves data for the map selected from the primary menu, while the 'Multiple Map' option saves all element maps available in the selected data category. Data is automatically saved to an 'Exported-Sampling' folder in the working directory.
Select Sampling > Circle from the main menu and draw a circle on the map by clicking on a location, holding the mouse down and dragging over an area. A crosshair cursor becomes available when the mouse pointer is over the map and the Sampling option is enabled.
Once you have drawn the circle, click and hold one of the four filled dots inside the circle to change its size. Inside the circle a cross cursor can be used to move the position of the circle.
TIP
If it is not possible to edit or move the ROI, select XMapTools, press Ctrl and try editing the ROI again.
The result of the data extraction is shown by the green bar in the graph at the top right of the main window and the corresponding numbers immediately below. A histogram is also plotted using all the pixel data contained within the ROI.
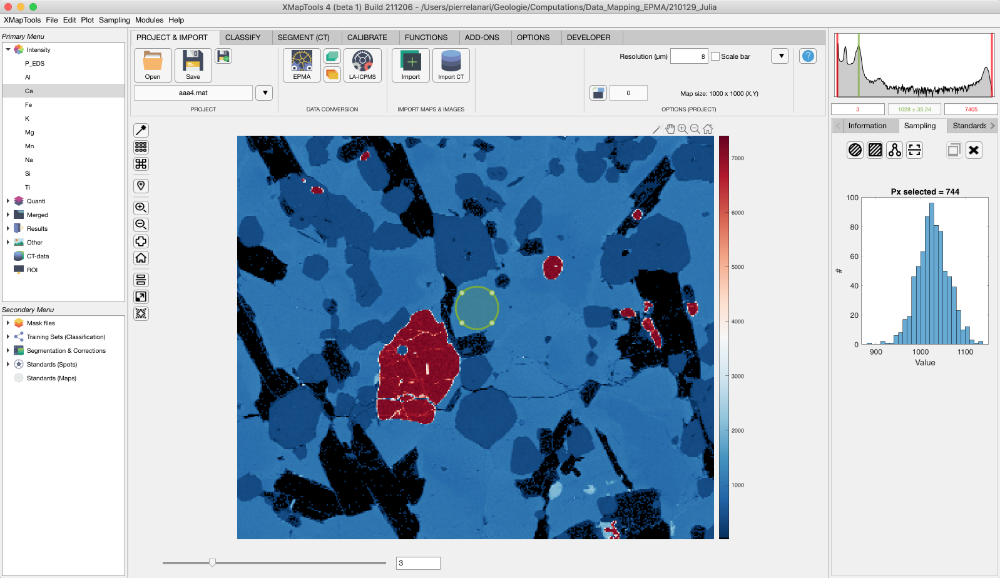
Figure 1: Example sampling of an intensity map (Ca) using the circle shape. Note that in this example, the average value of the selected pixels (Nb = 774) is 1028 ± 35.24 (stdev).
Select Sampling > Area (Polygon) from the main menu and draw a polygon on the map by clicking on several points. Click on the first point to close the polygon (a circle cursor appears).
The polygon shape can be edited.
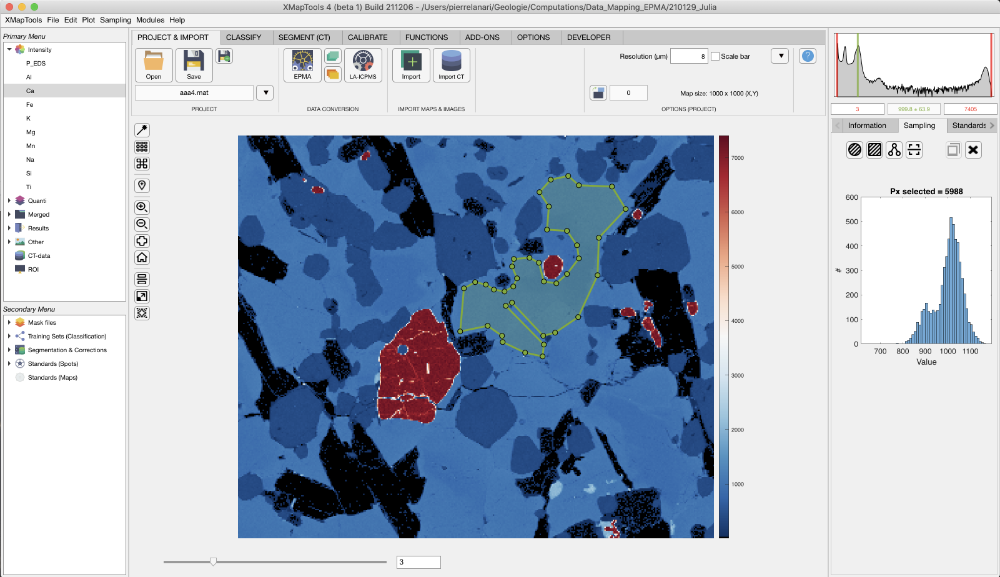
Figure 2: Example sampling of an intensity map (Ca) using the polygon shape. Note that in this example, the average value of the selected pixels (Nb = 774) is 999.8 ± 63.9 (stdev), but two plagioclase compositions are visible on the histogram (at 900 and 1030 counts).
Select Sampling > Transect from the main menu and draw a polygon on the map by clicking on several positions. Right-click to complete the selection. You must select at least two positions to obtain a valid transect.
The resulting profile is shown in the live display in the category sampling (Fig. 3).
The transect position can be edited and the plot will be updated automatically. Note that changing the colour bar boundaries will change the profile as the colours from the colour bar are used to plot the points.
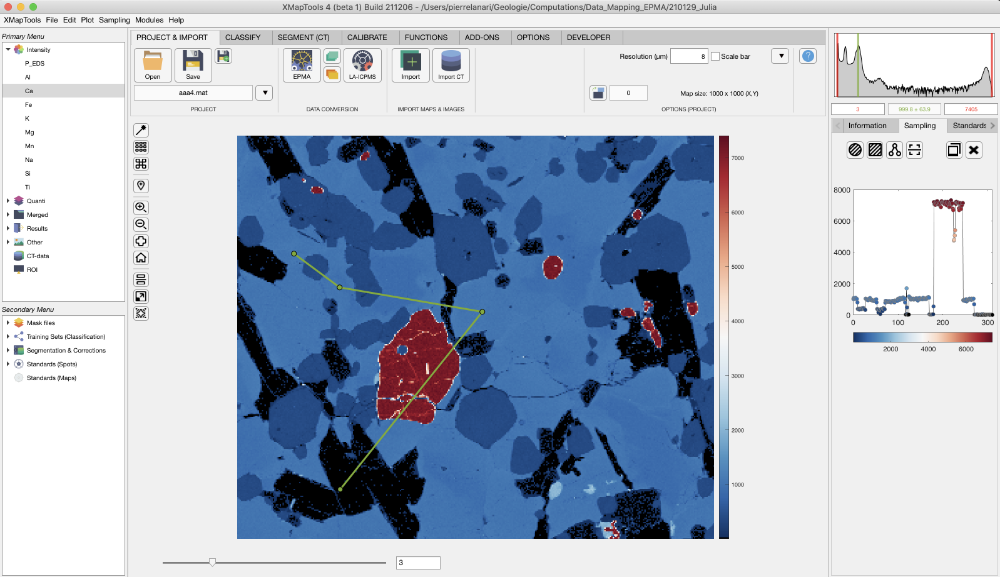
Figure 3: Example of sampling an intensity map (Ca) using a transect starting at the point located to the left.
This tool is used to extract the mean and median profile from a rectangular strip, using the composition of each transect. For a given pixel on the central transect, the value is calculated as the mean and median of the pixel compositions perpendicular to the reference transect.
Select Sampling > Strip from the main menu and draw a horizontal rectangle on the map by clicking on the corner position (top left), holding the mouse down and dragging over an area.
To move the rectangle, position the cursor in the centre of the rectangle, left-click, hold and drag, and drop it into the new position.
INFO
The rectangle must be drawn with its extension horizontal. However, it can be rotated later by positioning the cursor on the perimeter of the rectangle, near one of its edges, and then activating the rotation cursor.
Once the rectangle has been created (or edited), the plots in the Live Display Sampling category are updated. Two plots are available:
Figure 4: Example sampling of an intensity map (Ca) using a strip from bottom left to top right.
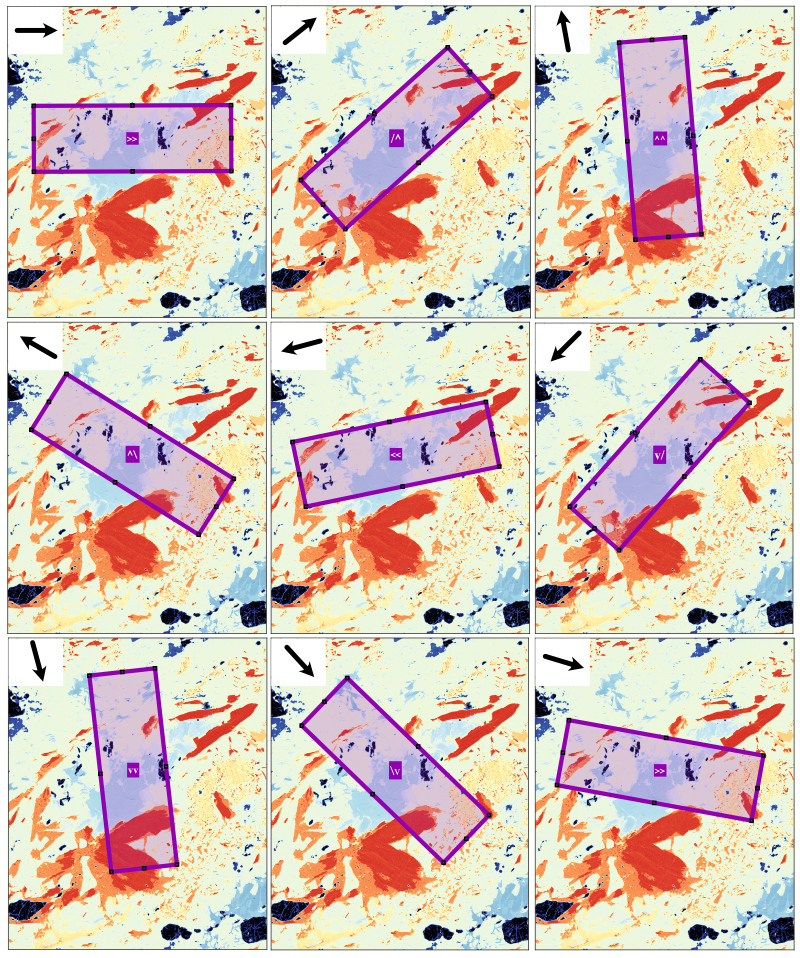

Figure 5: Orientation of the sampling strip.
',41)])])}const g=t(n,[["render",l]]);export{d as __pageData,g as default}; diff --git a/.vitepress/dist/assets/doc-sampling.md.DpW5G8we.lean.js b/.vitepress/dist/assets/doc-sampling.md.DpW5G8we.lean.js new file mode 100644 index 0000000..4bb9d83 --- /dev/null +++ b/.vitepress/dist/assets/doc-sampling.md.DpW5G8we.lean.js @@ -0,0 +1 @@ +import{_ as t,o as a,c as i,ae as o}from"./chunks/framework.DIXg7aqb.js";const d=JSON.parse('{"title":"Sampling Tools","description":"","frontmatter":{},"headers":[],"relativePath":"doc-sampling.md","filePath":"doc-sampling.md"}'),n={name:"doc-sampling.md"};function l(r,e,s,p,c,h){return a(),i("div",null,[...e[0]||(e[0]=[o("",41)])])}const g=t(n,[["render",l]]);export{d as __pageData,g as default}; diff --git a/.vitepress/dist/assets/doc-segment.md.xn7xnz3f.js b/.vitepress/dist/assets/doc-segment.md.xn7xnz3f.js new file mode 100644 index 0000000..a4064b6 --- /dev/null +++ b/.vitepress/dist/assets/doc-segment.md.xn7xnz3f.js @@ -0,0 +1 @@ +import{_ as t,o as a,c as n,ae as i}from"./chunks/framework.DIXg7aqb.js";const d=JSON.parse('{"title":"CT Segmentation","description":"","frontmatter":{},"headers":[],"relativePath":"doc-segment.md","filePath":"doc-segment.md"}'),o={name:"doc-segment.md"};function s(r,e,l,m,g,h){return a(),n("div",null,[...e[0]||(e[0]=[i('Based on XMapTools 4.5 embedded documentation – Help file version 11.01.2024
This page describes the tools available in the Segment section. These tools are optimised for CT data and should not be used with other data types.
The processing of CT data for segmentation is divided into five steps:
A segmentation scheme is a set of recipes containing constraints that are applied to perform data segmentation. It is similar to the training sets for classification, although it does not (yet) contain training data.
Segmentation schemes are available in the secondary menu under Segmentation & Corrections in Schemes (Segmentation).
Select a CT image under the CT Data node in the primary menu and select the Schemes (Segmentation) item in the secondary menu.
The Add (Scheme / Group / Range) ![]() button has various functions depending on the item selected in the secondary menu:
button has various functions depending on the item selected in the secondary menu:
INFO
Selected regions may overlap, but the algorithm will always consider the group in a sequential order. Pixels may be ignored by the segmentation algorithm if they are out of range.
Figure 1: Example of selecting a range of values (here from 128 to 212.6) for garnet. Note that this range has been stored in the segmentation scheme shown in the secondary menu.
These parameters can be set in the Segment (CT) and Segmentation Parameters & Test sections. There are two main algorithms implemented: Filter GB to remove grain boundaries and Interp GB to remove growth phases and re-interpolate filtered pixels.
Select a CT image from the CT Data node in the primary menu and a segmentation scheme from the secondary menu.
Four algorithms are available and can be selected from a drop-down menu:
You can set the threshold (range between 0 and 1) above which a grain boundary is detected and the order (for the local range).
The Calculate GB map ![]() button opens a new figure containing two plots (e.g. Fig. 2): the result of the algorithm on the left and the same map with the filtered grain boundaries shown in red on the right. The axes of the two plots are aligned, and zooming in on one plot automatically adjusts the view of the second plot.
button opens a new figure containing two plots (e.g. Fig. 2): the result of the algorithm on the left and the same map with the filtered grain boundaries shown in red on the right. The axes of the two plots are aligned, and zooming in on one plot automatically adjusts the view of the second plot.
Figure 2: Example of a grain boundary filter using the Local Gradient algorithm and a filter threshold of 0.15. Zoom has been used to show a small part of the image.
An algorithm based on a median filter is used to re-interpolate the grain boundary (an equivalent was available in XMapTools 3). The order of interpolation is adjustable.
You can test the segmentation on the selected image using the Segment Selected Image ![]() (small) button available in Segmentation Parameters & Test.
(small) button available in Segmentation Parameters & Test.
The resulting ROI is available in the ROI category of the primary menu.
The Segment All Images ![]() button applies the segmentation to all images available in CT-data. The resulting ROI is available in the ROI category of the primary menu.
button applies the segmentation to all images available in CT-data. The resulting ROI is available in the ROI category of the primary menu.
Select a multi-slice ROI (3D ROI) from the primary menu.
The Plot Phase Proportions ![]() button generates several plots to examine the spatial distribution of the segmented groups:
button generates several plots to examine the spatial distribution of the segmented groups:
Based on XMapTools 4.5 embedded documentation – Help file version 03.12.2021
This page describes the Spider Module which can be used to generate spider plots for LA-ICP-MS data.
Step 1: Select the normalisation data from the Normalisation drop-down menu ![]()
Step 2 (optional): Use the Auto Contrast ![]() button to automatically adjust the colour range.
button to automatically adjust the colour range.
Step 3: Select pixels to plot using one of the ROI types:
Step 4 (optional): Activate the Hold on ![]() mode to select multiple regions of interest. Simply repeat Step 3 to add more selections (circle or ROI). It is possible to edit and move any selection and the plot will be updated.
mode to select multiple regions of interest. Simply repeat Step 3 to add more selections (circle or ROI). It is possible to edit and move any selection and the plot will be updated.
TIP
Uncertainties are plotted on the graph. It is possible to choose to display '1se' (1sd/√N) or '1sd' ![]() .
.
Based on XMapTools 4.5 embedded documentation – Help file version 09.02.2026
This page describes the tools available in the Spot Data workspace. This feature was added in XMapTools 4.5.
In XMapTools, spot data is stored in datasets. A dataset is a collection of spot positions, and each position can be linked to external data.
New datasets can be created by selecting the "Spot Dataset" item in the secondary menu and by clicking on the Add ![]() button. New datasets are added to the list of datasets in the secondary menu on the left. The name of the dataset is automatically generated using the name provided in the text field (Fig. 1). A dataset can be renamed by double-clicking on its name in the list. To delete a dataset, first select it in the list, then right-click on its name and select Delete.
button. New datasets are added to the list of datasets in the secondary menu on the left. The name of the dataset is automatically generated using the name provided in the text field (Fig. 1). A dataset can be renamed by double-clicking on its name in the list. To delete a dataset, first select it in the list, then right-click on its name and select Delete.
Spots can be automatically added while creating a dataset if the Randomly populate option is selected. In this case the number of spots and their names is defined in the Nb and Spot Name fields. Spots are automatically distributed using the display image and only the pixels with a value between Cmin and Cmax. This allows users to select one mask/submask or a specific composition of the map.

Figure 1: Dataset tools for spot data in XMapTools 4.5.
Each spot corresponds to a set of X,Y coordinates, but the compositional data can be extracted from taking the median and MAD values of the surrounding pixels. If the Integrate Pixels option is activated, the compositional data is extracted by integrating the values of the surrounding pixels. The number of pixels to be integrated is defined in the Nb x Nb fields corresponding to the number of pixels along X and Y directions. These two numbers must be odd to ensure that the central pixel is included in the integration.

Figure 2: Spot tools for spot data in XMapTools 4.5.
Additional spots can be added by pressing the Add Spot ![]() button and by immediately clicking on the map to define the spot position. The spots can be deleted by selecting them in the secondary menu and then by right-clicking on their name and selecting Delete. The position of the spots can be edited by selecting their name in the list and dragging them on the map. The coordinates of the spots are automatically updated.
button and by immediately clicking on the map to define the spot position. The spots can be deleted by selecting them in the secondary menu and then by right-clicking on their name and selecting Delete. The position of the spots can be edited by selecting their name in the list and dragging them on the map. The coordinates of the spots are automatically updated.
In the example reported in Figure 3 below, 10 spots were automatically added to the newly created dataset 'Garnet'. The default names were used for the spots (Spot_#). The compositional data of the spots are extracted by integrating the values of the surrounding pixels in a 5x5 grid (Nb = 25). Note that the spot coordinates are available in a table on the right of the interface when the dataset garnet is selected (Fig. 3).
Figure 3: Spot data interface in XMapTools 4.5.
If a spot is selected in the secondary menu, the names, coordinates and value of each pixel are listed in the table on the right of the interface. The compositional data of the spot is extracted by taking the median and MAD values of the surrounding pixels. You can delete pixels by selecting them in the table and by pressing the Delete button. You can reset to the original n×n grid of pixels by pressing the Reset button.
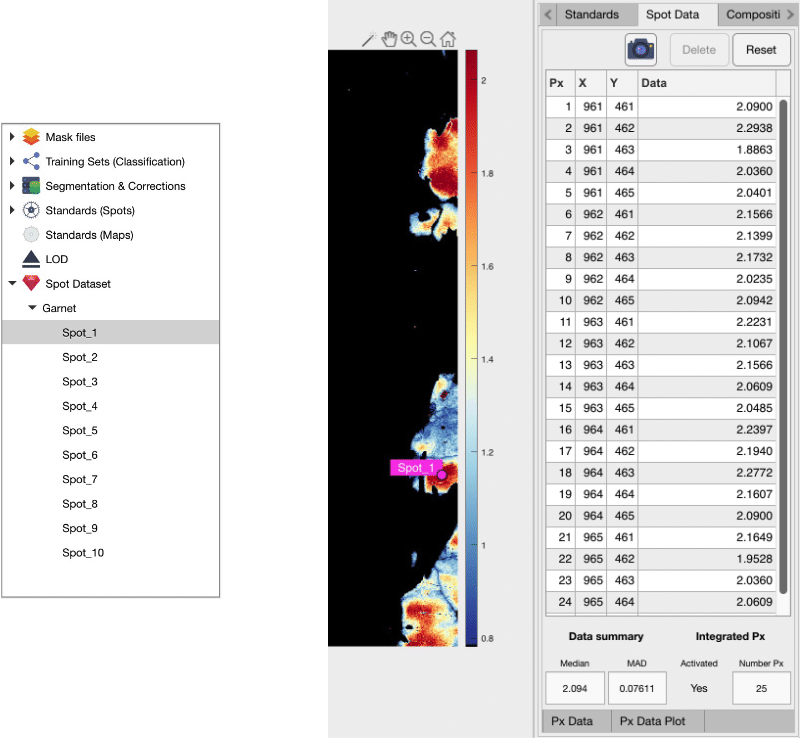
Figure 4: Spot data pixel details in XMapTools 4.5.
External data for the spots can be imported and visualised in XMapTools. The following procedure is recommended:
Import the spot external data by using the Import ![]() button available in the Spot External Data section (Fig. 5). The number of columns in the CSV file (except the name column) corresponds to the number of variables that will be imported and visualised in XMapTools (e.g. one variable in Fig. 5). You can use the display button to open a table containing the spot names and external values. The external data can be visualised on the map by selecting the variable to be displayed in the Add to plot dropdown menu in the Plot external data section.
button available in the Spot External Data section (Fig. 5). The number of columns in the CSV file (except the name column) corresponds to the number of variables that will be imported and visualised in XMapTools (e.g. one variable in Fig. 5). You can use the display button to open a table containing the spot names and external values. The external data can be visualised on the map by selecting the variable to be displayed in the Add to plot dropdown menu in the Plot external data section.

Figure 5: Spot External Data tools in XMapTools 4.5.
Two options Apply a colour gradient and Apply spot size gradient can be selected in the Plot external data section (e.g. colour gradient for Fig. 6).
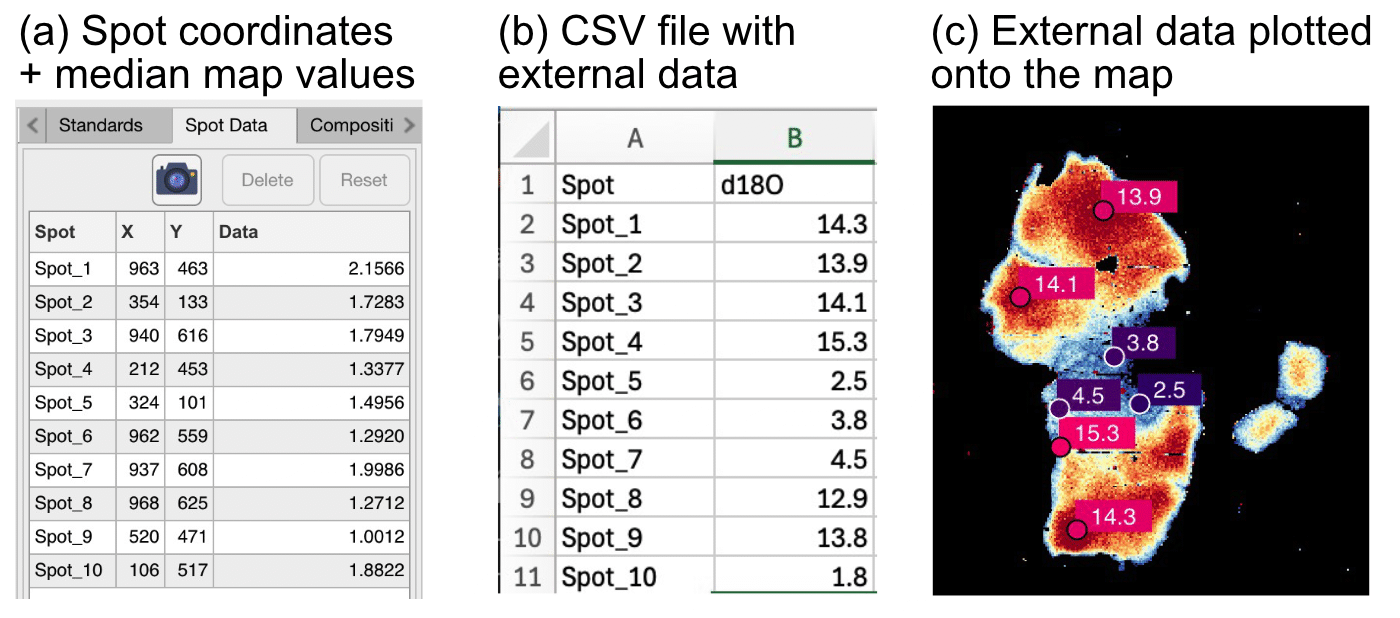
Figure 6: Example of (a) spot data in XMapTools, (b) external data for δ18O saved in a CSV file, (c) external spot data (δ18O) plotted onto the map.
Spot data can be exported using the Export module available in the Add-on section. Open the module and select the Export Spot Data mode. Select your data source (e.g. Quanti in Fig. 7) and press the Generate & Save button.

Figure 7: Export module for Spot Data in XMapTools 4.5.
',42)])])}const g=t(n,[["render",s]]);export{c as __pageData,g as default}; diff --git a/.vitepress/dist/assets/doc-spot-data.md.BaQTjTtE.lean.js b/.vitepress/dist/assets/doc-spot-data.md.BaQTjTtE.lean.js new file mode 100644 index 0000000..825e7c3 --- /dev/null +++ b/.vitepress/dist/assets/doc-spot-data.md.BaQTjTtE.lean.js @@ -0,0 +1 @@ +import{_ as t,o as a,c as o,ae as i}from"./chunks/framework.DIXg7aqb.js";const c=JSON.parse('{"title":"Spot Data","description":"","frontmatter":{},"headers":[],"relativePath":"doc-spot-data.md","filePath":"doc-spot-data.md"}'),n={name:"doc-spot-data.md"};function s(l,e,p,r,d,m){return a(),o("div",null,[...e[0]||(e[0]=[i("",42)])])}const g=t(n,[["render",s]]);export{c as __pageData,g as default}; diff --git a/.vitepress/dist/assets/doc-visualization.md.CwED1vHD.js b/.vitepress/dist/assets/doc-visualization.md.CwED1vHD.js new file mode 100644 index 0000000..22320cb --- /dev/null +++ b/.vitepress/dist/assets/doc-visualization.md.CwED1vHD.js @@ -0,0 +1 @@ +import{_ as a,o as i,c as t,ae as o}from"./chunks/framework.DIXg7aqb.js";const h=JSON.parse('{"title":"Data Visualization Tools","description":"","frontmatter":{},"headers":[],"relativePath":"doc-visualization.md","filePath":"doc-visualization.md"}'),l={name:"doc-visualization.md"};function n(s,e,r,p,d,m){return i(),t("div",null,[...e[0]||(e[0]=[o('Based on XMapTools 4.5 embedded documentation – Help file version 09.05.2025
This page describes the Data Visualization module, which can be used to display map data in histograms, binary plots, ternary plots and as an RGB composite image.
The Data Visualization module has access to the data selected in the primary menu of XMapTools when it is opened. To change the type of data, close the module and re-open it after selecting different data.
The type of plot to be generated can be selected in the Plot section. The following modes are available:
By default, XMapTools generates a binary diagram when the module is opened if at least two variables (maps) are available.
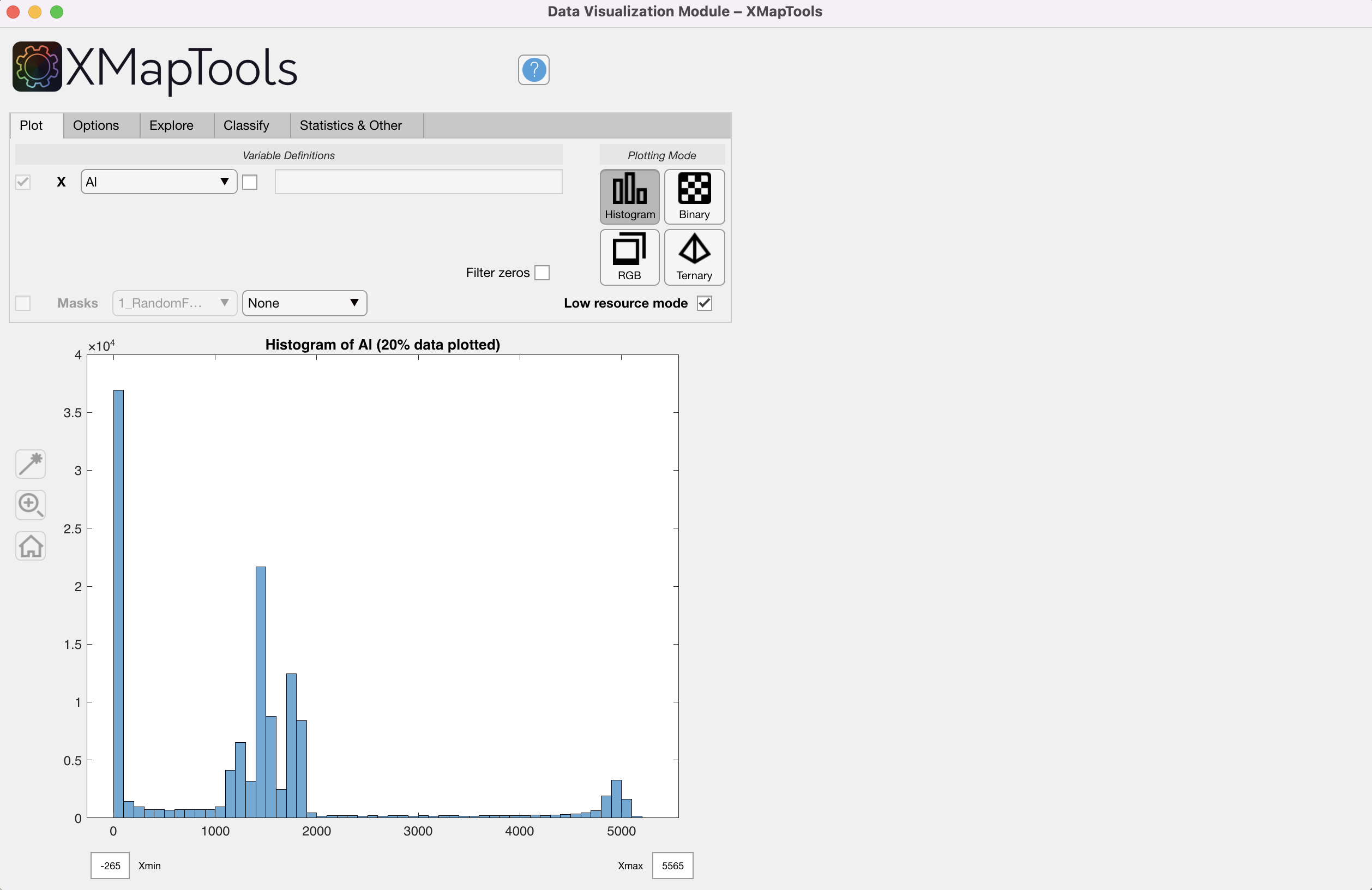
Figure 1: Data Visualization Module, histogram mode.
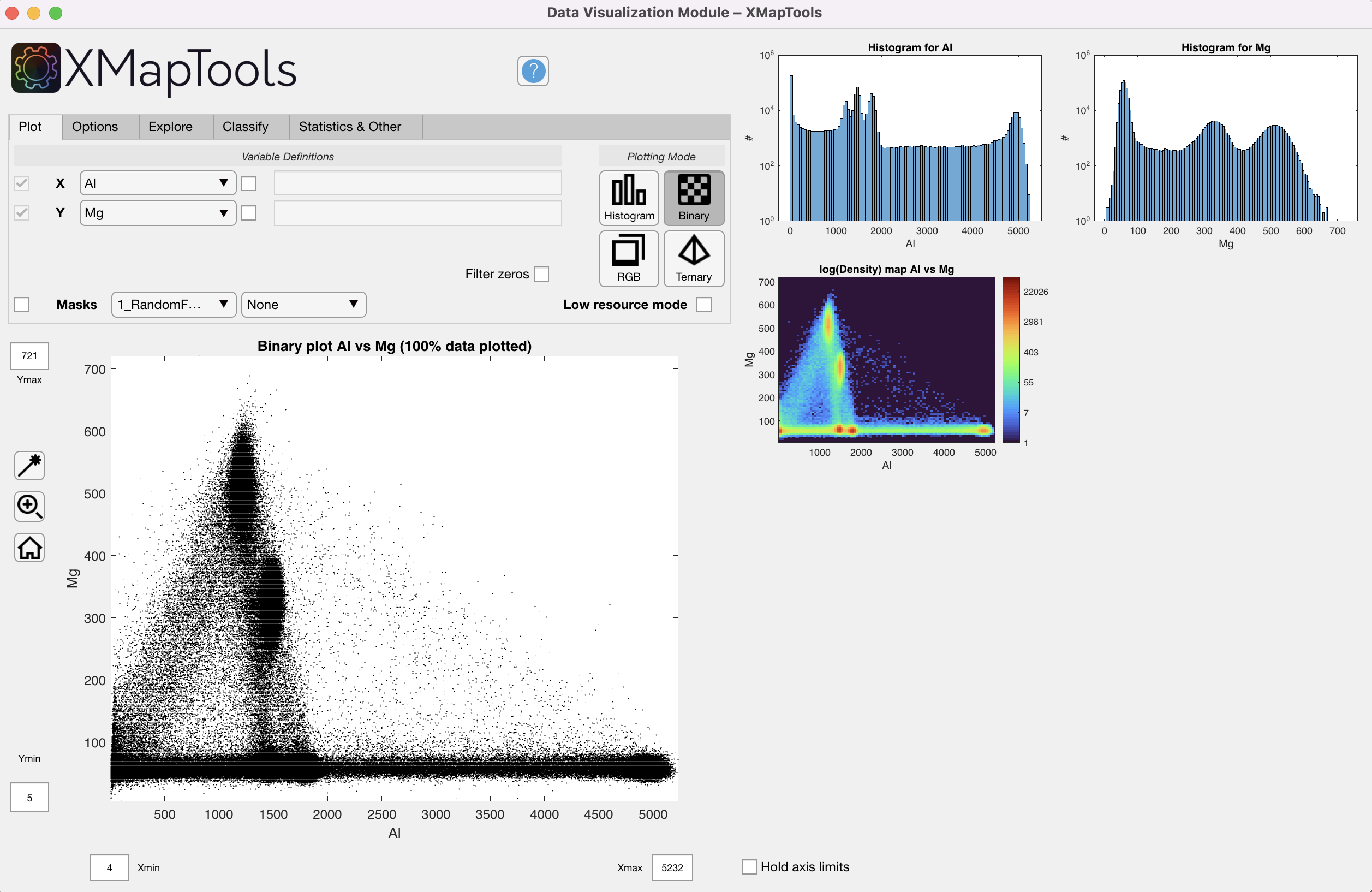
Figure 2: Data Visualization Module, binary mode.
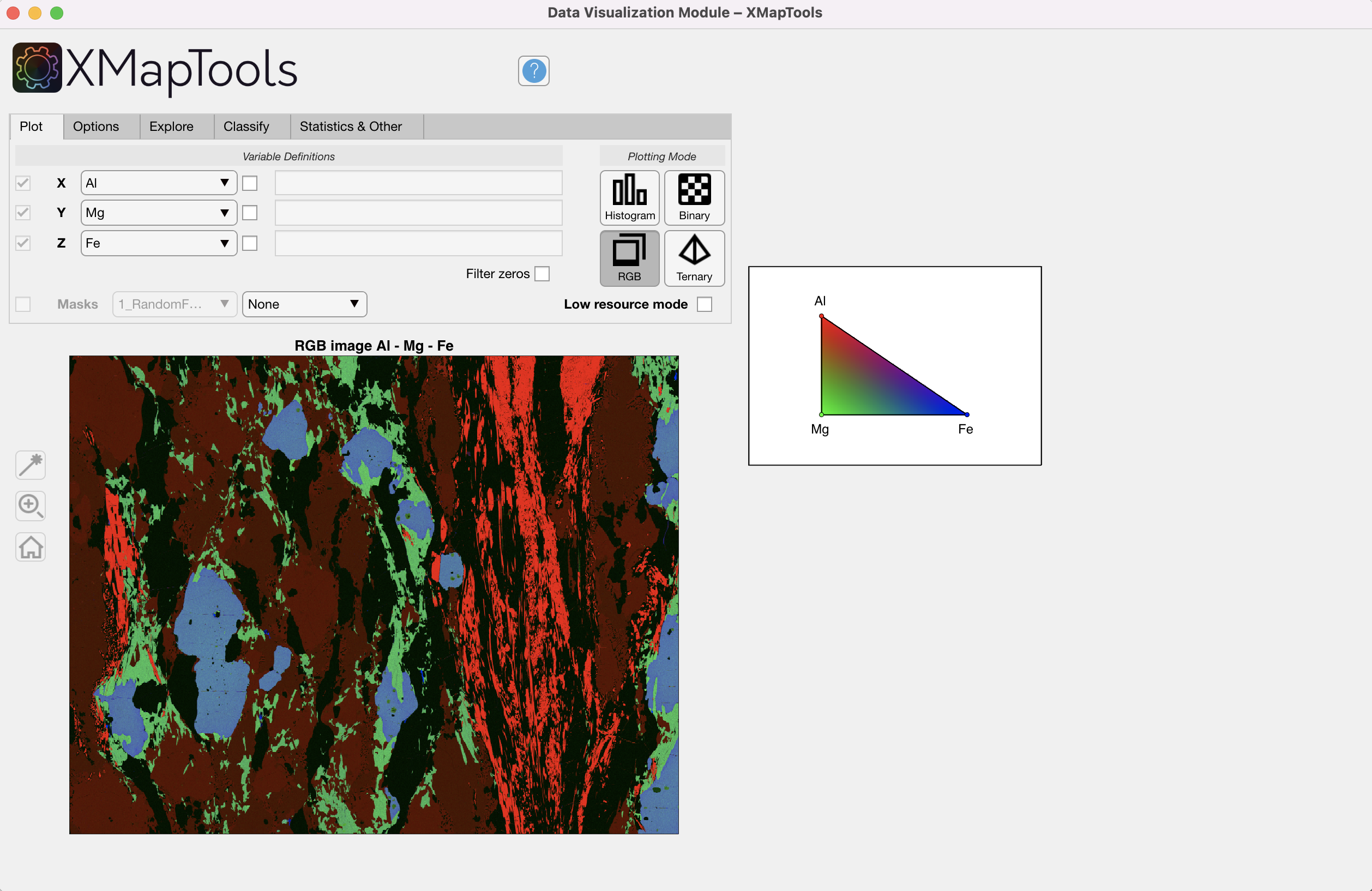
Figure 3: Data Visualization Module, RGB mode.
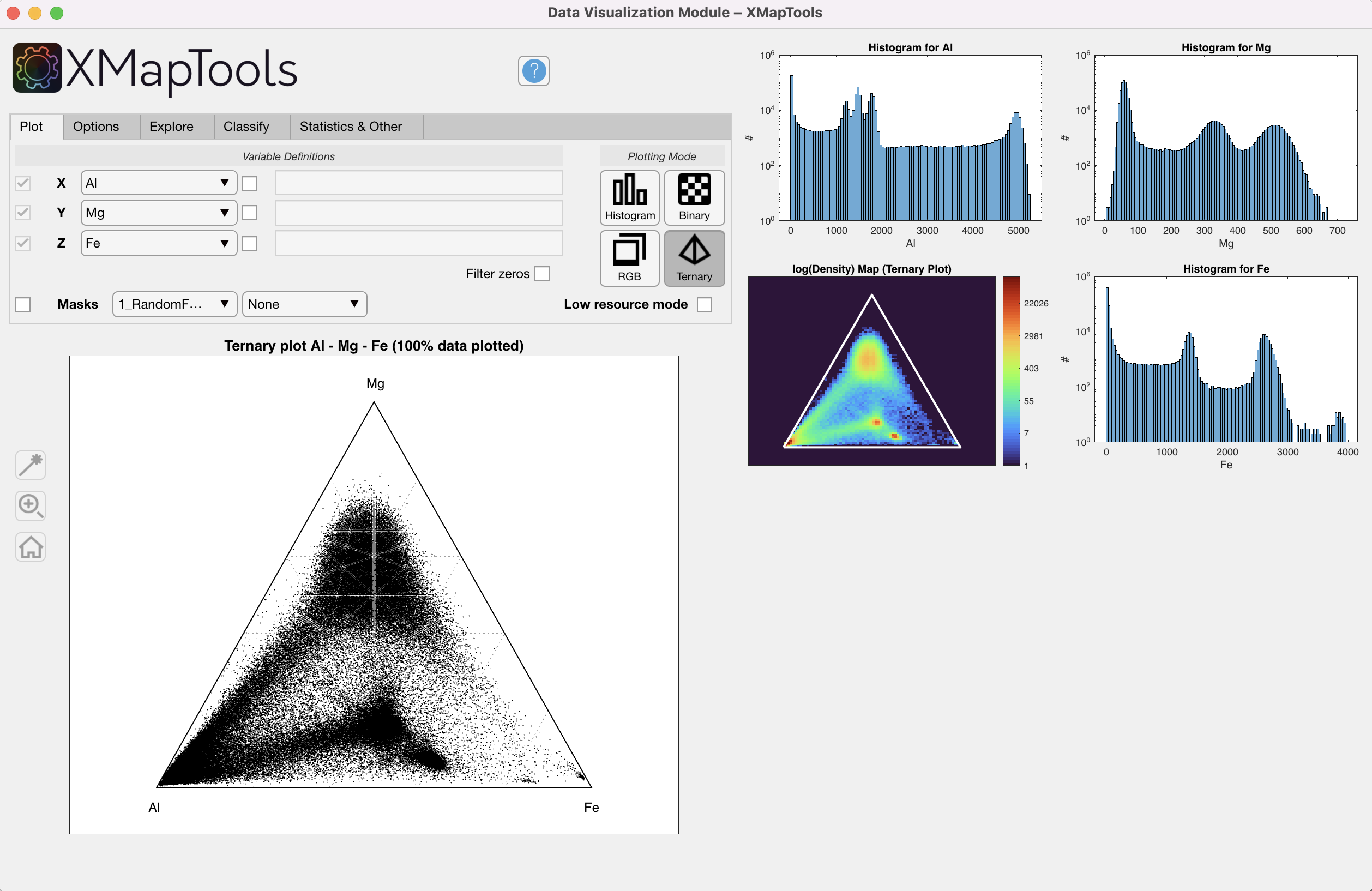
Figure 4: Data Visualization Module, ternary mode.
On the left hand side of the Plot section you can select the maps to be displayed. These can be elements, groups of elements or any other variable (e.g. structural formula).

Figure 5: Variable definition.
Whatever the plot type, variables are selected in Variable Definitions, either using the drop-down menus listing all the available maps, or using any combination of them.
Standard MATLAB code format is used to combine existing maps and create new variables to plot. Use an arithmetic operation defined by the rules of linear algebra or array arithmetic. The operation can include spaces between variables, digits, and operators. Array operators (with the dot character ".") must be used when maps are divided or multiplied by each other.
TIP
Element-wise multiplication or division should use .* or ./ to multiply or divide the two arrays element by element.
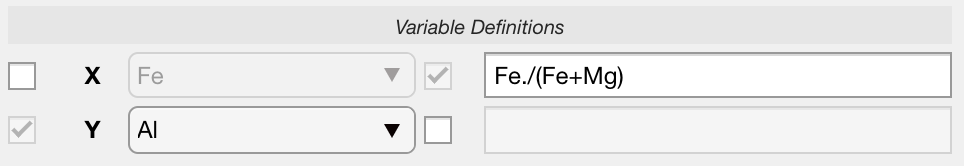
Figure 6: Variable definition using code.
A basic pixel identification tool is only available for binary and ternary maps. Use it to create polygons that identify the location of points on the map.
When a polygon is drawn, the pixel locations of all points within the region of interest are displayed on the map at the bottom right. The shape can be edited and the plot updated accordingly. An identical polygon is plotted on the density map when low resource mode is off.
The Polygon ![]() button activates a cross cursor and you can select vertices of a polygon around any region of interest in the main plot. Right-click to close the polygon.
button activates a cross cursor and you can select vertices of a polygon around any region of interest in the main plot. Right-click to close the polygon.
The selection can be saved as a mask file.
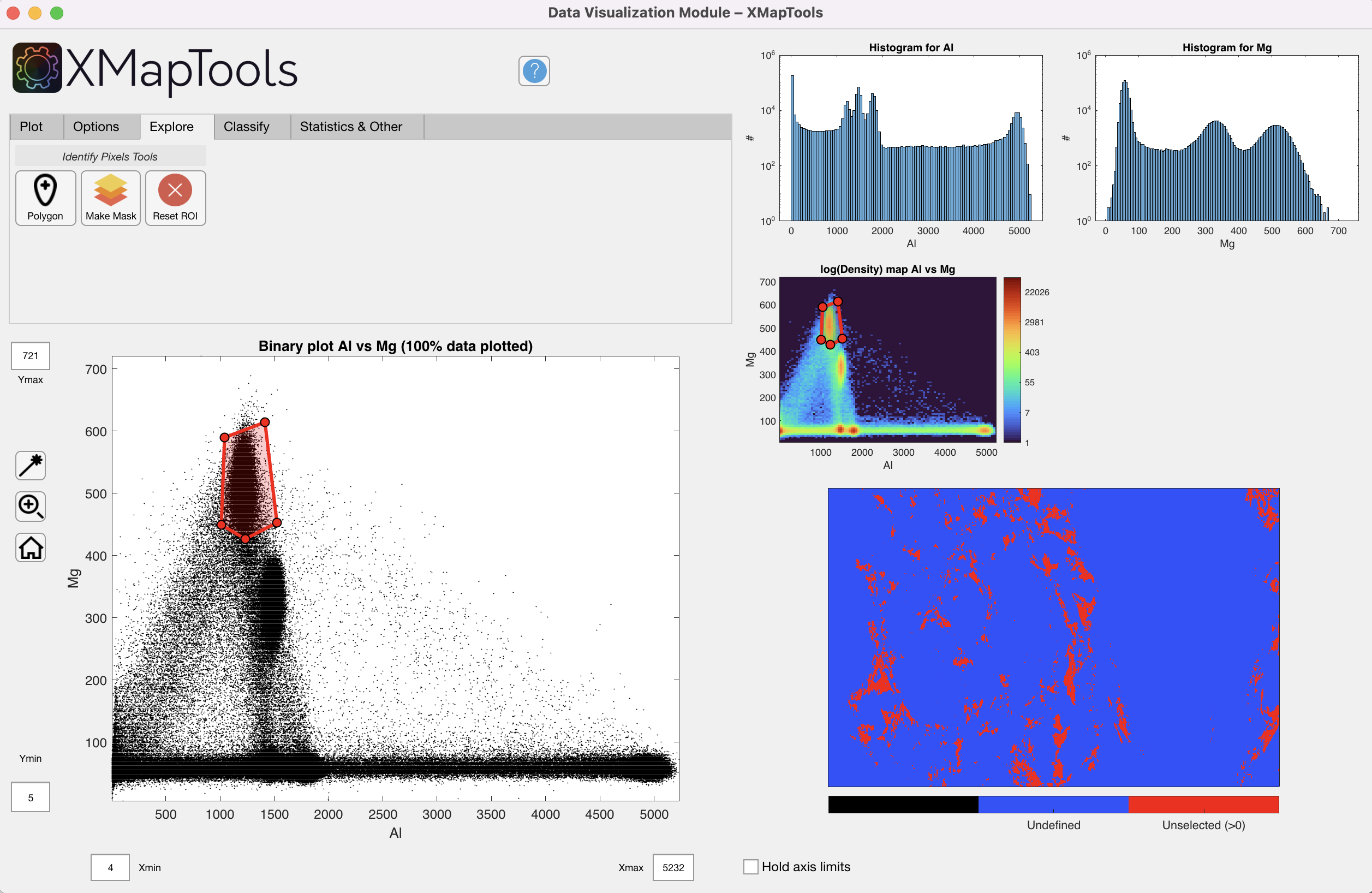
Figure 7: Data Visualization Module — Identify pixels using a polygonal selection in a binary plot.
![]() Selecting the Masks check box will colour the data displayed in the binary or ternary plots according to the mask file displayed in the drop down menu.
Selecting the Masks check box will colour the data displayed in the binary or ternary plots according to the mask file displayed in the drop down menu.
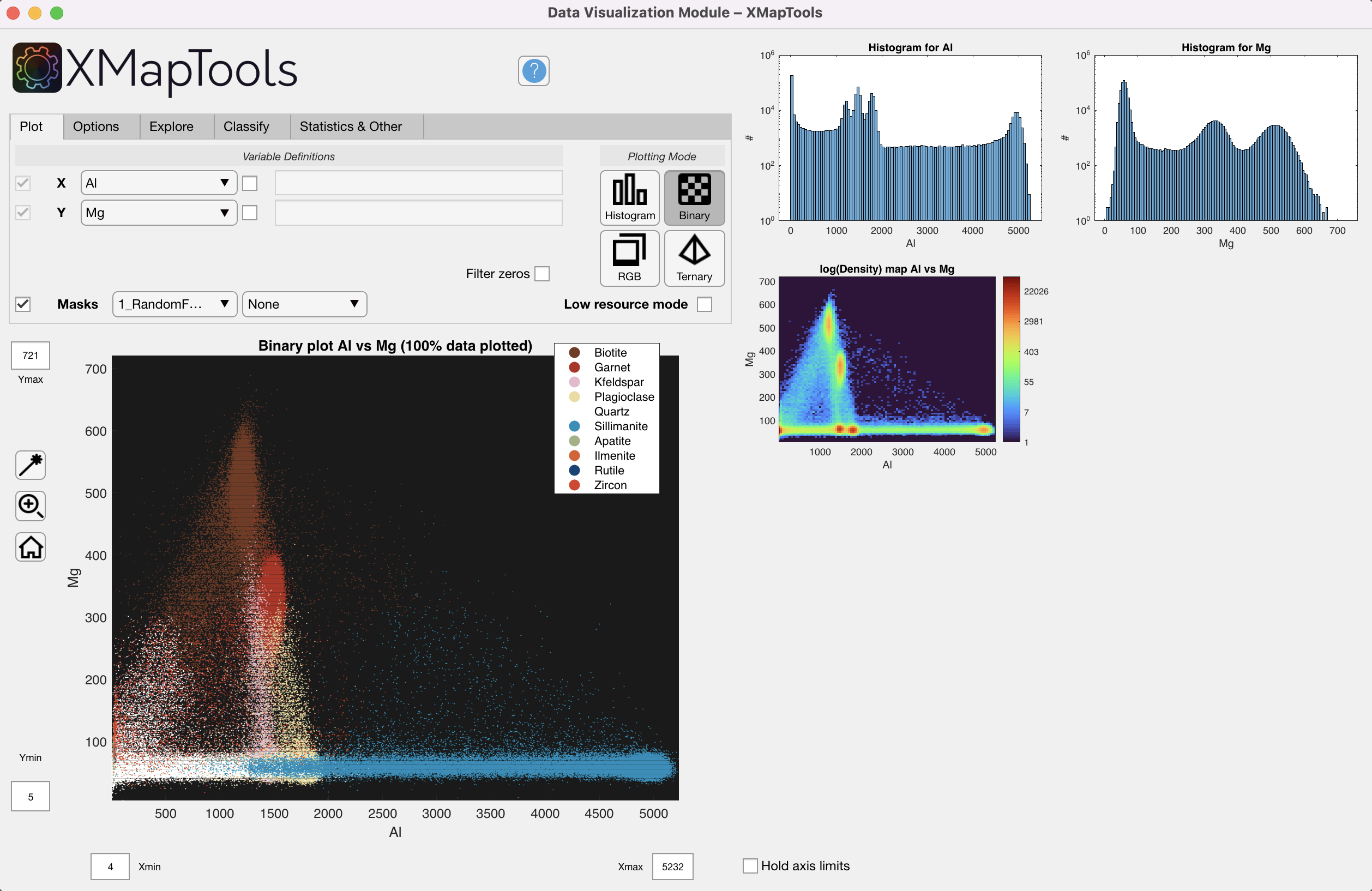
Figure 8: Intensity data coloured according to minerals retrieved from the selected mask file.
This mode is optimal for computers with limited memory and is enabled by default when the Data Visualization module is opened. When this mode is activated, only 40% of the data is plotted (random selection).
The "Bulk Value" (Average) is calculated from the data plotted in the figure. There is no density correction applied, so deviations from the true bulk composition are expected if phases have different densities.
Statistics are reported in the Statistics section and are updated each time the plot is changed:
The plotting report section can contain the following information:
Open the XMapTools app. Note: it may take up to a minute to start the first time while MATLAB libraries are loaded. XMapTools will start slightly faster on subsequent launches (until you restart your computer).
A dialogue box opens — select your working directory and press Open. A working directory contains the data for a specific project. XMapTools automatically generates files during use that are stored there. Although you can change the working directory later, you must select one at startup. A log file is also automatically created in the selected working directory.
The current working directory is displayed in the top bar of the main window.
You are now ready to go! You can start converting raw data or importing maps.
XMapTools creates many files during data processing and it is better to store these files together with the data they belong to.
It is strongly recommended to have a folder for each project that contains only the map and standard files needed by XMapTools. Raw data should be stored in a separate folder.
All the MATLAB libraries need to be loaded, which takes some time. The second time you run XMapTools it will be faster if you haven't restarted your computer. Use this time to think about your project and perhaps grab a cup of tea or coffee.
You can find articles on this website and help files in the program. We also have tutorial videos. Do not hesitate to click on the help button available in each section to access a description of the main steps. If you feel that something is missing or unclear, or if you find an error, do not hesitate to contact us on Discord or via the developer repository on GitHub.
',13)])])}const u=t(i,[["render",s]]);export{m as __pageData,u as default}; diff --git a/.vitepress/dist/assets/getting-started-2.md.CEzqsAHA.lean.js b/.vitepress/dist/assets/getting-started-2.md.CEzqsAHA.lean.js new file mode 100644 index 0000000..7fe8d09 --- /dev/null +++ b/.vitepress/dist/assets/getting-started-2.md.CEzqsAHA.lean.js @@ -0,0 +1 @@ +import{_ as t,o,c as a,ae as r}from"./chunks/framework.DIXg7aqb.js";const m=JSON.parse('{"title":"Opening XMapTools for the First Time","description":"","frontmatter":{},"headers":[],"relativePath":"getting-started-2.md","filePath":"getting-started-2.md"}'),i={name:"getting-started-2.md"};function s(n,e,l,h,d,p){return o(),a("div",null,[...e[0]||(e[0]=[r("",13)])])}const u=t(i,[["render",s]]);export{m as __pageData,u as default}; diff --git a/.vitepress/dist/assets/getting-started.md.BLdsliy4.js b/.vitepress/dist/assets/getting-started.md.BLdsliy4.js new file mode 100644 index 0000000..9cc77b4 --- /dev/null +++ b/.vitepress/dist/assets/getting-started.md.BLdsliy4.js @@ -0,0 +1 @@ +import{_ as e,o as a,c as o,ae as n}from"./chunks/framework.DIXg7aqb.js";const u=JSON.parse('{"title":"Getting Started","description":"","frontmatter":{},"headers":[],"relativePath":"getting-started.md","filePath":"getting-started.md"}'),r={name:"getting-started.md"};function i(s,t,d,l,h,p){return a(),o("div",null,[...t[0]||(t[0]=[n('Welcome to the XMapTools documentation! Whether you are a new or returning user, this guide will help you get XMapTools 4.5 up and running.
The installation and update procedures have changed in XMapTools 4.5. Most binaries now require MATLAB Runtime 25.1, which is installed automatically by the XMapTools installer. Older Runtime versions (e.g. v99 or v912) are no longer needed and can be safely deleted after upgrading.
XMapTools is distributed via shell scripts for macOS and Windows that automate the installation, update, download, extraction, and configuration of the program.
Before installing, make sure your system meets the requirements below:
| Version | Processor | Operating System | Runtime |
|---|---|---|---|
| macOS | Apple Silicon (M1, M2, M3, M4, …) | macOS Sequoia (15), Sonoma (14), or Ventura (13) | R2025a (25.1) |
| macOS Intel | Intel | macOS Sequoia (15), Sonoma (14), or Ventura (13) | R2020b (v99) |
| Windows | x86-64 | Windows 10 or Windows 11 | R2025a (25.1) |
Note: The macOS version is not compatible with Intel processors, and the macOS Intel version is not stable with Apple Silicon processors. The installer auto-detects your architecture.
Ready to install? Head over to the Installation & Update Guide for step-by-step instructions.
Is XMapTools already installed? If so, go straight here.
',11)])])}const g=e(r,[["render",i]]);export{u as __pageData,g as default}; diff --git a/.vitepress/dist/assets/getting-started.md.BLdsliy4.lean.js b/.vitepress/dist/assets/getting-started.md.BLdsliy4.lean.js new file mode 100644 index 0000000..e0753d0 --- /dev/null +++ b/.vitepress/dist/assets/getting-started.md.BLdsliy4.lean.js @@ -0,0 +1 @@ +import{_ as e,o as a,c as o,ae as n}from"./chunks/framework.DIXg7aqb.js";const u=JSON.parse('{"title":"Getting Started","description":"","frontmatter":{},"headers":[],"relativePath":"getting-started.md","filePath":"getting-started.md"}'),r={name:"getting-started.md"};function i(s,t,d,l,h,p){return a(),o("div",null,[...t[0]||(t[0]=[n("",11)])])}const g=e(r,[["render",i]]);export{u as __pageData,g as default}; diff --git a/.vitepress/dist/assets/index.md.Doinpoc4.js b/.vitepress/dist/assets/index.md.Doinpoc4.js new file mode 100644 index 0000000..59aefaf --- /dev/null +++ b/.vitepress/dist/assets/index.md.Doinpoc4.js @@ -0,0 +1 @@ +import{_ as o,o as i,c as e,j as a,a as n}from"./chunks/framework.DIXg7aqb.js";const g=JSON.parse('{"title":"","description":"","frontmatter":{"layout":"home","hero":{"name":"XMapTools 4.5","text":"Compositional mapping","tagline":"Free and versatile software solution for quantitative chemical maps analysis.","actions":[{"theme":"brand","text":"Installation","link":"/installation_guide"},{"theme":"alt","text":"Official website","link":"https://xmaptools.ch"},{"theme":"alt","text":"Developer repository","link":"https://github.com/xmaptools/XMapTools_Developers"}],"image":{"src":"/assets/Logo_2025_square_512.png","alt":"XMapTools"}},"features":[{"icon":"📚","title":"Documentation","details":"Powerful tools for quantitative compositional mapping and analysis","link":"/getting-started"},{"icon":"💻","title":"Tutorials","details":"Video tutorials and demonstrations of XMapTools functionality","link":"/tutorials"},{"icon":"🚀","title":"Bingo-Antidote","details":"Step-by-step tutorial for using Bingo-Antidote in XMapTools","link":"/bingo-antidote"}]},"headers":[],"relativePath":"index.md","filePath":"index.md"}'),s={name:"index.md"};function l(r,t,p,d,m,u){return i(),e("div",null,[...t[0]||(t[0]=[a("h2",{id:"what-is-xmaptools",tabindex:"-1"},[n("What is XMapTools? "),a("a",{class:"header-anchor",href:"#what-is-xmaptools","aria-label":'Permalink to "What is XMapTools?"'},"")],-1),a("p",null,"XMapTools is an advanced analysis software for quantitative chemical analysis of solids in 1D, 2D and 3D.",-1),a("p",null,"It provides numerical tools and packages implemented in a guided and versatile environment that allows you to explore and visualise data in your own way.",-1),a("p",null,"For example, XMapTools includes a wide range of data processing options including routines for classification, segmentation, calibration and visualisation via single and multi-channel maps or via binary, ternary and spider diagrams.",-1),a("br",null,null,-1),a("img",{src:"https://raw.githubusercontent.com/xmaptools/Documentation_Images/main/Logo/XMapTools_Transition.jpg",alt:"XMapTools Logo",style:{"max-width":"60%",height:"auto",display:"block",margin:"0 auto"}},null,-1),a("br",null,null,-1),a("img",{src:"https://raw.githubusercontent.com/xmaptools/Documentation_Images/main/Logo/Logo_2025_square_512.png",alt:"XMapTools Logo",style:{"max-width":"10%",height:"auto",display:"block",margin:"0 auto"}},null,-1)])])}const h=o(s,[["render",l]]);export{g as __pageData,h as default}; diff --git a/.vitepress/dist/assets/index.md.Doinpoc4.lean.js b/.vitepress/dist/assets/index.md.Doinpoc4.lean.js new file mode 100644 index 0000000..59aefaf --- /dev/null +++ b/.vitepress/dist/assets/index.md.Doinpoc4.lean.js @@ -0,0 +1 @@ +import{_ as o,o as i,c as e,j as a,a as n}from"./chunks/framework.DIXg7aqb.js";const g=JSON.parse('{"title":"","description":"","frontmatter":{"layout":"home","hero":{"name":"XMapTools 4.5","text":"Compositional mapping","tagline":"Free and versatile software solution for quantitative chemical maps analysis.","actions":[{"theme":"brand","text":"Installation","link":"/installation_guide"},{"theme":"alt","text":"Official website","link":"https://xmaptools.ch"},{"theme":"alt","text":"Developer repository","link":"https://github.com/xmaptools/XMapTools_Developers"}],"image":{"src":"/assets/Logo_2025_square_512.png","alt":"XMapTools"}},"features":[{"icon":"📚","title":"Documentation","details":"Powerful tools for quantitative compositional mapping and analysis","link":"/getting-started"},{"icon":"💻","title":"Tutorials","details":"Video tutorials and demonstrations of XMapTools functionality","link":"/tutorials"},{"icon":"🚀","title":"Bingo-Antidote","details":"Step-by-step tutorial for using Bingo-Antidote in XMapTools","link":"/bingo-antidote"}]},"headers":[],"relativePath":"index.md","filePath":"index.md"}'),s={name:"index.md"};function l(r,t,p,d,m,u){return i(),e("div",null,[...t[0]||(t[0]=[a("h2",{id:"what-is-xmaptools",tabindex:"-1"},[n("What is XMapTools? "),a("a",{class:"header-anchor",href:"#what-is-xmaptools","aria-label":'Permalink to "What is XMapTools?"'},"")],-1),a("p",null,"XMapTools is an advanced analysis software for quantitative chemical analysis of solids in 1D, 2D and 3D.",-1),a("p",null,"It provides numerical tools and packages implemented in a guided and versatile environment that allows you to explore and visualise data in your own way.",-1),a("p",null,"For example, XMapTools includes a wide range of data processing options including routines for classification, segmentation, calibration and visualisation via single and multi-channel maps or via binary, ternary and spider diagrams.",-1),a("br",null,null,-1),a("img",{src:"https://raw.githubusercontent.com/xmaptools/Documentation_Images/main/Logo/XMapTools_Transition.jpg",alt:"XMapTools Logo",style:{"max-width":"60%",height:"auto",display:"block",margin:"0 auto"}},null,-1),a("br",null,null,-1),a("img",{src:"https://raw.githubusercontent.com/xmaptools/Documentation_Images/main/Logo/Logo_2025_square_512.png",alt:"XMapTools Logo",style:{"max-width":"10%",height:"auto",display:"block",margin:"0 auto"}},null,-1)])])}const h=o(s,[["render",l]]);export{g as __pageData,h as default}; diff --git a/.vitepress/dist/assets/installation_guide.md.DyFiVWdL.js b/.vitepress/dist/assets/installation_guide.md.DyFiVWdL.js new file mode 100644 index 0000000..4b4961e --- /dev/null +++ b/.vitepress/dist/assets/installation_guide.md.DyFiVWdL.js @@ -0,0 +1,10 @@ +import{_ as a,o as i,c as t,ae as e}from"./chunks/framework.DIXg7aqb.js";const u=JSON.parse('{"title":"XMapTools Installation & Update Guide","description":"","frontmatter":{},"headers":[],"relativePath":"installation_guide.md","filePath":"installation_guide.md"}'),n={name:"installation_guide.md"};function l(o,s,r,h,p,d){return i(),t("div",null,[...s[0]||(s[0]=[e(`This guide covers how to install and update XMapTools 4.5 and future releases on macOS and Windows using the official bootstrap scripts. These scripts automate downloading, extracting, and configuring XMapTools and the required MATLAB Runtime.
| Requirement | macOS | Windows |
|---|---|---|
| OS | macOS (Intel or Apple Silicon) | Windows 10/11 |
| Admin rights | Yes (script prompts for sudo password) | Yes (run PowerShell as Administrator) |
| Internet connection | Required | Required |
| Shell | Terminal (zsh or bash) | PowerShell |
Note: The MATLAB Runtime is installed automatically during a full installation. You do not need a MATLAB licence.
Compatibility: XMapTools 4.5 requires MATLAB Runtime 25.1. Older versions of the Runtime (e.g. v99, or v912) can be deleted after updating to XMapTools 4.5.
Security: The code executed by the install command is available at https://xmaptools.ch/install.sh (macOS) and https://xmaptools.ch/install.ps1 (Windows). Do not use any other domain name. The
xmaptools.chdomain is official and the only domain used to distribute XMapTools. If you are behind a firewall that blocks access, ask your system administrator to allow HTTPS traffic to this domain.
All macOS commands are run in the Terminal app. The script is fetched directly from the XMapTools server and executed locally.
A full installation downloads XMapTools and the MATLAB Runtime. Use this when installing XMapTools for the first time or when you need to reinstall the runtime.
Auto-detect architecture (recommended):
curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --installThe script automatically detects whether your Mac uses Apple Silicon or Intel and downloads the correct version.
Specify architecture manually:
# Apple Silicon (M1, M2, M3, M4, …)
+curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --install AppleSilicon
+
+# Intel
+curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --install IntelWhat happens during installation:
/Applications/XMapTools is removed (you may be prompted for your administrator password).xmaptools is created so you can launch the app from any terminal window.Graphical installer steps (macOS):
/Applications/XMapTools to allow automatic updates in the future and press Next./Applications/MATLAB/MATLAB_Runtime and press Next.XMapTools.app.An update replaces the XMapTools application bundle without reinstalling the MATLAB Runtime. This is faster than a full installation and preserves your runtime setup.
Prerequisite: XMapTools must already be installed in
/Applications/XMapTools.
Auto-detect architecture (recommended):
curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --updateSpecify architecture manually:
# Apple Silicon
+curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --update AppleSilicon
+
+# Intel
+curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --update IntelWhat happens during an update:
XMapTools.app) is replaced with the new version.To display the available versions, remote file timestamps, detected MATLAB Runtime installations, and usage instructions:
curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --info| Option | When to use |
|---|---|
AppleSilicon | Macs with M1, M2, M3, M4, or later Apple chips (native performance) |
Intel | Macs with Intel processors |
| (omitted) | The script auto-detects the architecture — Apple Silicon is selected on arm64 hardware |
Tip: If you are unsure which architecture your Mac uses, click > About This Mac. If the Chip field says "Apple M…", use
AppleSilicon. If it says "Intel", useIntel.
After installation or update, the script creates a command-line launcher at /usr/local/bin/xmaptools. You can start XMapTools from any terminal by typing:
xmaptoolsThe script also updates your shell configuration file (.zshrc, .bash_profile, or .profile) to ensure /usr/local/bin is on your PATH.
All Windows commands are run in PowerShell. You should run PowerShell as Administrator (right-click → Run as Administrator).
A full installation downloads XMapTools and the MATLAB Runtime.
iex "& { $(irm https://xmaptools.ch/install.ps1) } --install"What happens during installation:
C:\\Program Files\\XMapTools is removed.XMapToolsInstaller_Windows.exe) is launched — follow the on-screen instructions to complete the setup.An update replaces the XMapTools executable and splash screen without reinstalling the MATLAB Runtime.
Prerequisite: XMapTools must already be installed in
C:\\Program Files\\XMapTools.
iex "& { $(irm https://xmaptools.ch/install.ps1) } --update"What happens during an update:
XMapTools.exe (and splash.png if present) are copied into the existing installation directory.To display the available version, remote file timestamps, detected MATLAB Runtime installations, and usage instructions:
iex "& { $(irm https://xmaptools.ch/install.ps1) } --info"This error occurs when running --update without a prior installation. Run a full installation first:
curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --installiex "& { $(irm https://xmaptools.ch/install.ps1) } --install"If XMapTools reports an invalid MATLAB Runtime version after an update, the runtime may have changed between versions. Perform a full reinstallation using --install to get the matching runtime.
The scripts check for the MATLAB Runtime in:
/Applications/MATLAB/MATLAB_RuntimeC:\\Program Files\\MATLAB\\MATLAB RuntimeIf the runtime is missing, a full installation (--install) is required.
| Runtime version | Platform |
|---|---|
v99 (R2020b) | macOS Intel |
R2025a | macOS Apple Silicon (native), Windows |
The installation script automatically clears Gatekeeper quarantine flags. If you still see a warning, run manually:
sudo xattr -cr /Applications/XMapTools/application/XMapTools.appIf PowerShell refuses to run the script, you may need to adjust the execution policy for the current session:
Set-ExecutionPolicy -Scope Process -ExecutionPolicy BypassThen re-run the installation or update command.
Make sure you are running PowerShell as Administrator. Right-click the PowerShell icon and select Run as Administrator.
If XMapTools was working previously and suddenly stops opening, this can be caused by a corrupted MATLAB Runtime installation or a Windows update that modified system paths. Try the following:
--install flagIf XMapTools becomes unresponsive during an operation, a dialog box may appear with OK and Cancel buttons. Click Cancel to cancel the current operation and unfreeze XMapTools. Clicking OK may restart the operation and keep XMapTools frozen.
To fully uninstall XMapTools on Windows:
| Action | macOS | Windows |
|---|---|---|
| Install | curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --install | iex "& { $(irm https://xmaptools.ch/install.ps1) } --install" |
| Update | curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --update | iex "& { $(irm https://xmaptools.ch/install.ps1) } --update" |
| Info | curl -fsSL https://xmaptools.ch/install.sh | bash -s -- --info | iex "& { $(irm https://xmaptools.ch/install.ps1) } --info" |
| Install dir | /Applications/XMapTools | C:\\Program Files\\XMapTools |
| Runtime dir | /Applications/MATLAB/MATLAB_Runtime | C:\\Program Files\\MATLAB\\MATLAB Runtime |
After upgrading to XMapTools 4.5, older MATLAB Runtime versions (v99, v912) are no longer needed and can be safely removed to free disk space.
macOS:
sudo rm -rf /Applications/MATLAB/MATLAB_Runtime/v99
+sudo rm -rf /Applications/MATLAB/MATLAB_Runtime/v912Windows:
Use Settings → Apps & Features (or Add or Remove Programs) to uninstall the older MATLAB Runtime versions (R2020b / v912).
`,99)])])}const k=a(n,[["render",l]]);export{u as __pageData,k as default}; diff --git a/.vitepress/dist/assets/installation_guide.md.DyFiVWdL.lean.js b/.vitepress/dist/assets/installation_guide.md.DyFiVWdL.lean.js new file mode 100644 index 0000000..bbb8de8 --- /dev/null +++ b/.vitepress/dist/assets/installation_guide.md.DyFiVWdL.lean.js @@ -0,0 +1 @@ +import{_ as a,o as i,c as t,ae as e}from"./chunks/framework.DIXg7aqb.js";const u=JSON.parse('{"title":"XMapTools Installation & Update Guide","description":"","frontmatter":{},"headers":[],"relativePath":"installation_guide.md","filePath":"installation_guide.md"}'),n={name:"installation_guide.md"};function l(o,s,r,h,p,d){return i(),t("div",null,[...s[0]||(s[0]=[e("",99)])])}const k=a(n,[["render",l]]);export{u as __pageData,k as default}; diff --git a/.vitepress/dist/assets/inter-italic-cyrillic-ext.r48I6akx.woff2 b/.vitepress/dist/assets/inter-italic-cyrillic-ext.r48I6akx.woff2 new file mode 100644 index 0000000..b6b603d Binary files /dev/null and b/.vitepress/dist/assets/inter-italic-cyrillic-ext.r48I6akx.woff2 differ diff --git a/.vitepress/dist/assets/inter-italic-cyrillic.By2_1cv3.woff2 b/.vitepress/dist/assets/inter-italic-cyrillic.By2_1cv3.woff2 new file mode 100644 index 0000000..def40a4 Binary files /dev/null and b/.vitepress/dist/assets/inter-italic-cyrillic.By2_1cv3.woff2 differ diff --git a/.vitepress/dist/assets/inter-italic-greek-ext.1u6EdAuj.woff2 b/.vitepress/dist/assets/inter-italic-greek-ext.1u6EdAuj.woff2 new file mode 100644 index 0000000..e070c3d Binary files /dev/null and b/.vitepress/dist/assets/inter-italic-greek-ext.1u6EdAuj.woff2 differ diff --git a/.vitepress/dist/assets/inter-italic-greek.DJ8dCoTZ.woff2 b/.vitepress/dist/assets/inter-italic-greek.DJ8dCoTZ.woff2 new file mode 100644 index 0000000..a3c16ca Binary files /dev/null and b/.vitepress/dist/assets/inter-italic-greek.DJ8dCoTZ.woff2 differ diff --git a/.vitepress/dist/assets/inter-italic-latin-ext.CN1xVJS-.woff2 b/.vitepress/dist/assets/inter-italic-latin-ext.CN1xVJS-.woff2 new file mode 100644 index 0000000..2210a89 Binary files /dev/null and b/.vitepress/dist/assets/inter-italic-latin-ext.CN1xVJS-.woff2 differ diff --git a/.vitepress/dist/assets/inter-italic-latin.C2AdPX0b.woff2 b/.vitepress/dist/assets/inter-italic-latin.C2AdPX0b.woff2 new file mode 100644 index 0000000..790d62d Binary files /dev/null and b/.vitepress/dist/assets/inter-italic-latin.C2AdPX0b.woff2 differ diff --git a/.vitepress/dist/assets/inter-italic-vietnamese.BSbpV94h.woff2 b/.vitepress/dist/assets/inter-italic-vietnamese.BSbpV94h.woff2 new file mode 100644 index 0000000..1eec077 Binary files /dev/null and b/.vitepress/dist/assets/inter-italic-vietnamese.BSbpV94h.woff2 differ diff --git a/.vitepress/dist/assets/inter-roman-cyrillic-ext.BBPuwvHQ.woff2 b/.vitepress/dist/assets/inter-roman-cyrillic-ext.BBPuwvHQ.woff2 new file mode 100644 index 0000000..2cfe615 Binary files /dev/null and b/.vitepress/dist/assets/inter-roman-cyrillic-ext.BBPuwvHQ.woff2 differ diff --git a/.vitepress/dist/assets/inter-roman-cyrillic.C5lxZ8CY.woff2 b/.vitepress/dist/assets/inter-roman-cyrillic.C5lxZ8CY.woff2 new file mode 100644 index 0000000..e3886dd Binary files /dev/null and b/.vitepress/dist/assets/inter-roman-cyrillic.C5lxZ8CY.woff2 differ diff --git a/.vitepress/dist/assets/inter-roman-greek-ext.CqjqNYQ-.woff2 b/.vitepress/dist/assets/inter-roman-greek-ext.CqjqNYQ-.woff2 new file mode 100644 index 0000000..36d6748 Binary files /dev/null and b/.vitepress/dist/assets/inter-roman-greek-ext.CqjqNYQ-.woff2 differ diff --git a/.vitepress/dist/assets/inter-roman-greek.BBVDIX6e.woff2 b/.vitepress/dist/assets/inter-roman-greek.BBVDIX6e.woff2 new file mode 100644 index 0000000..2bed1e8 Binary files /dev/null and b/.vitepress/dist/assets/inter-roman-greek.BBVDIX6e.woff2 differ diff --git a/.vitepress/dist/assets/inter-roman-latin-ext.4ZJIpNVo.woff2 b/.vitepress/dist/assets/inter-roman-latin-ext.4ZJIpNVo.woff2 new file mode 100644 index 0000000..9a8d1e2 Binary files /dev/null and b/.vitepress/dist/assets/inter-roman-latin-ext.4ZJIpNVo.woff2 differ diff --git a/.vitepress/dist/assets/inter-roman-latin.Di8DUHzh.woff2 b/.vitepress/dist/assets/inter-roman-latin.Di8DUHzh.woff2 new file mode 100644 index 0000000..07d3c53 Binary files /dev/null and b/.vitepress/dist/assets/inter-roman-latin.Di8DUHzh.woff2 differ diff --git a/.vitepress/dist/assets/inter-roman-vietnamese.BjW4sHH5.woff2 b/.vitepress/dist/assets/inter-roman-vietnamese.BjW4sHH5.woff2 new file mode 100644 index 0000000..57bdc22 Binary files /dev/null and b/.vitepress/dist/assets/inter-roman-vietnamese.BjW4sHH5.woff2 differ diff --git a/.vitepress/dist/assets/style.BwkHieaG.css b/.vitepress/dist/assets/style.BwkHieaG.css new file mode 100644 index 0000000..7d3c00e --- /dev/null +++ b/.vitepress/dist/assets/style.BwkHieaG.css @@ -0,0 +1 @@ +@font-face{font-family:Inter;font-style:normal;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-roman-cyrillic-ext.BBPuwvHQ.woff2) format("woff2");unicode-range:U+0460-052F,U+1C80-1C88,U+20B4,U+2DE0-2DFF,U+A640-A69F,U+FE2E-FE2F}@font-face{font-family:Inter;font-style:normal;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-roman-cyrillic.C5lxZ8CY.woff2) format("woff2");unicode-range:U+0301,U+0400-045F,U+0490-0491,U+04B0-04B1,U+2116}@font-face{font-family:Inter;font-style:normal;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-roman-greek-ext.CqjqNYQ-.woff2) format("woff2");unicode-range:U+1F00-1FFF}@font-face{font-family:Inter;font-style:normal;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-roman-greek.BBVDIX6e.woff2) format("woff2");unicode-range:U+0370-0377,U+037A-037F,U+0384-038A,U+038C,U+038E-03A1,U+03A3-03FF}@font-face{font-family:Inter;font-style:normal;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-roman-vietnamese.BjW4sHH5.woff2) format("woff2");unicode-range:U+0102-0103,U+0110-0111,U+0128-0129,U+0168-0169,U+01A0-01A1,U+01AF-01B0,U+0300-0301,U+0303-0304,U+0308-0309,U+0323,U+0329,U+1EA0-1EF9,U+20AB}@font-face{font-family:Inter;font-style:normal;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-roman-latin-ext.4ZJIpNVo.woff2) format("woff2");unicode-range:U+0100-02AF,U+0304,U+0308,U+0329,U+1E00-1E9F,U+1EF2-1EFF,U+2020,U+20A0-20AB,U+20AD-20C0,U+2113,U+2C60-2C7F,U+A720-A7FF}@font-face{font-family:Inter;font-style:normal;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-roman-latin.Di8DUHzh.woff2) format("woff2");unicode-range:U+0000-00FF,U+0131,U+0152-0153,U+02BB-02BC,U+02C6,U+02DA,U+02DC,U+0304,U+0308,U+0329,U+2000-206F,U+2074,U+20AC,U+2122,U+2191,U+2193,U+2212,U+2215,U+FEFF,U+FFFD}@font-face{font-family:Inter;font-style:italic;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-italic-cyrillic-ext.r48I6akx.woff2) format("woff2");unicode-range:U+0460-052F,U+1C80-1C88,U+20B4,U+2DE0-2DFF,U+A640-A69F,U+FE2E-FE2F}@font-face{font-family:Inter;font-style:italic;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-italic-cyrillic.By2_1cv3.woff2) format("woff2");unicode-range:U+0301,U+0400-045F,U+0490-0491,U+04B0-04B1,U+2116}@font-face{font-family:Inter;font-style:italic;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-italic-greek-ext.1u6EdAuj.woff2) format("woff2");unicode-range:U+1F00-1FFF}@font-face{font-family:Inter;font-style:italic;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-italic-greek.DJ8dCoTZ.woff2) format("woff2");unicode-range:U+0370-0377,U+037A-037F,U+0384-038A,U+038C,U+038E-03A1,U+03A3-03FF}@font-face{font-family:Inter;font-style:italic;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-italic-vietnamese.BSbpV94h.woff2) format("woff2");unicode-range:U+0102-0103,U+0110-0111,U+0128-0129,U+0168-0169,U+01A0-01A1,U+01AF-01B0,U+0300-0301,U+0303-0304,U+0308-0309,U+0323,U+0329,U+1EA0-1EF9,U+20AB}@font-face{font-family:Inter;font-style:italic;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-italic-latin-ext.CN1xVJS-.woff2) format("woff2");unicode-range:U+0100-02AF,U+0304,U+0308,U+0329,U+1E00-1E9F,U+1EF2-1EFF,U+2020,U+20A0-20AB,U+20AD-20C0,U+2113,U+2C60-2C7F,U+A720-A7FF}@font-face{font-family:Inter;font-style:italic;font-weight:100 900;font-display:swap;src:url(/Documentation/assets/inter-italic-latin.C2AdPX0b.woff2) format("woff2");unicode-range:U+0000-00FF,U+0131,U+0152-0153,U+02BB-02BC,U+02C6,U+02DA,U+02DC,U+0304,U+0308,U+0329,U+2000-206F,U+2074,U+20AC,U+2122,U+2191,U+2193,U+2212,U+2215,U+FEFF,U+FFFD}@font-face{font-family:Punctuation SC;font-weight:400;src:local("PingFang SC Regular"),local("Noto Sans CJK SC"),local("Microsoft YaHei");unicode-range:U+201C,U+201D,U+2018,U+2019,U+2E3A,U+2014,U+2013,U+2026,U+00B7,U+007E,U+002F}@font-face{font-family:Punctuation SC;font-weight:500;src:local("PingFang SC Medium"),local("Noto Sans CJK SC"),local("Microsoft YaHei");unicode-range:U+201C,U+201D,U+2018,U+2019,U+2E3A,U+2014,U+2013,U+2026,U+00B7,U+007E,U+002F}@font-face{font-family:Punctuation SC;font-weight:600;src:local("PingFang SC Semibold"),local("Noto Sans CJK SC Bold"),local("Microsoft YaHei Bold");unicode-range:U+201C,U+201D,U+2018,U+2019,U+2E3A,U+2014,U+2013,U+2026,U+00B7,U+007E,U+002F}@font-face{font-family:Punctuation SC;font-weight:700;src:local("PingFang SC Semibold"),local("Noto Sans CJK SC Bold"),local("Microsoft YaHei Bold");unicode-range:U+201C,U+201D,U+2018,U+2019,U+2E3A,U+2014,U+2013,U+2026,U+00B7,U+007E,U+002F}:root{--vp-c-white: #ffffff;--vp-c-black: #000000;--vp-c-neutral: var(--vp-c-black);--vp-c-neutral-inverse: var(--vp-c-white)}.dark{--vp-c-neutral: var(--vp-c-white);--vp-c-neutral-inverse: var(--vp-c-black)}:root{--vp-c-gray-1: #dddde3;--vp-c-gray-2: #e4e4e9;--vp-c-gray-3: #ebebef;--vp-c-gray-soft: rgba(142, 150, 170, .14);--vp-c-indigo-1: #3451b2;--vp-c-indigo-2: #3a5ccc;--vp-c-indigo-3: #5672cd;--vp-c-indigo-soft: rgba(100, 108, 255, .14);--vp-c-purple-1: #6f42c1;--vp-c-purple-2: #7e4cc9;--vp-c-purple-3: #8e5cd9;--vp-c-purple-soft: rgba(159, 122, 234, .14);--vp-c-green-1: #18794e;--vp-c-green-2: #299764;--vp-c-green-3: #30a46c;--vp-c-green-soft: rgba(16, 185, 129, .14);--vp-c-yellow-1: #915930;--vp-c-yellow-2: #946300;--vp-c-yellow-3: #9f6a00;--vp-c-yellow-soft: rgba(234, 179, 8, .14);--vp-c-red-1: #b8272c;--vp-c-red-2: #d5393e;--vp-c-red-3: #e0575b;--vp-c-red-soft: rgba(244, 63, 94, .14);--vp-c-sponsor: #db2777}.dark{--vp-c-gray-1: #515c67;--vp-c-gray-2: #414853;--vp-c-gray-3: #32363f;--vp-c-gray-soft: rgba(101, 117, 133, .16);--vp-c-indigo-1: #a8b1ff;--vp-c-indigo-2: #5c73e7;--vp-c-indigo-3: #3e63dd;--vp-c-indigo-soft: rgba(100, 108, 255, .16);--vp-c-purple-1: #c8abfa;--vp-c-purple-2: #a879e6;--vp-c-purple-3: #8e5cd9;--vp-c-purple-soft: rgba(159, 122, 234, .16);--vp-c-green-1: #3dd68c;--vp-c-green-2: #30a46c;--vp-c-green-3: #298459;--vp-c-green-soft: rgba(16, 185, 129, .16);--vp-c-yellow-1: #f9b44e;--vp-c-yellow-2: #da8b17;--vp-c-yellow-3: #a46a0a;--vp-c-yellow-soft: rgba(234, 179, 8, .16);--vp-c-red-1: #f66f81;--vp-c-red-2: #f14158;--vp-c-red-3: #b62a3c;--vp-c-red-soft: rgba(244, 63, 94, .16)}:root{--vp-c-bg: #ffffff;--vp-c-bg-alt: #f6f6f7;--vp-c-bg-elv: #ffffff;--vp-c-bg-soft: #f6f6f7}.dark{--vp-c-bg: #1b1b1f;--vp-c-bg-alt: #161618;--vp-c-bg-elv: #202127;--vp-c-bg-soft: #202127}:root{--vp-c-border: #c2c2c4;--vp-c-divider: #e2e2e3;--vp-c-gutter: #e2e2e3}.dark{--vp-c-border: #3c3f44;--vp-c-divider: #2e2e32;--vp-c-gutter: #000000}:root{--vp-c-text-1: #3c3c43;--vp-c-text-2: #67676c;--vp-c-text-3: #929295}.dark{--vp-c-text-1: #dfdfd6;--vp-c-text-2: #98989f;--vp-c-text-3: #6a6a71}:root{--vp-c-default-1: var(--vp-c-gray-1);--vp-c-default-2: var(--vp-c-gray-2);--vp-c-default-3: var(--vp-c-gray-3);--vp-c-default-soft: var(--vp-c-gray-soft);--vp-c-brand-1: var(--vp-c-indigo-1);--vp-c-brand-2: var(--vp-c-indigo-2);--vp-c-brand-3: var(--vp-c-indigo-3);--vp-c-brand-soft: var(--vp-c-indigo-soft);--vp-c-brand: var(--vp-c-brand-1);--vp-c-tip-1: var(--vp-c-brand-1);--vp-c-tip-2: var(--vp-c-brand-2);--vp-c-tip-3: var(--vp-c-brand-3);--vp-c-tip-soft: var(--vp-c-brand-soft);--vp-c-note-1: var(--vp-c-brand-1);--vp-c-note-2: var(--vp-c-brand-2);--vp-c-note-3: var(--vp-c-brand-3);--vp-c-note-soft: var(--vp-c-brand-soft);--vp-c-success-1: var(--vp-c-green-1);--vp-c-success-2: var(--vp-c-green-2);--vp-c-success-3: var(--vp-c-green-3);--vp-c-success-soft: var(--vp-c-green-soft);--vp-c-important-1: var(--vp-c-purple-1);--vp-c-important-2: var(--vp-c-purple-2);--vp-c-important-3: var(--vp-c-purple-3);--vp-c-important-soft: var(--vp-c-purple-soft);--vp-c-warning-1: var(--vp-c-yellow-1);--vp-c-warning-2: var(--vp-c-yellow-2);--vp-c-warning-3: var(--vp-c-yellow-3);--vp-c-warning-soft: var(--vp-c-yellow-soft);--vp-c-danger-1: var(--vp-c-red-1);--vp-c-danger-2: var(--vp-c-red-2);--vp-c-danger-3: var(--vp-c-red-3);--vp-c-danger-soft: var(--vp-c-red-soft);--vp-c-caution-1: var(--vp-c-red-1);--vp-c-caution-2: var(--vp-c-red-2);--vp-c-caution-3: var(--vp-c-red-3);--vp-c-caution-soft: var(--vp-c-red-soft)}:root{--vp-font-family-base: "Inter", ui-sans-serif, system-ui, sans-serif, "Apple Color Emoji", "Segoe UI Emoji", "Segoe UI Symbol", "Noto Color Emoji";--vp-font-family-mono: ui-monospace, "Menlo", "Monaco", "Consolas", "Liberation Mono", "Courier New", monospace;font-optical-sizing:auto}:root:where(:lang(zh)){--vp-font-family-base: "Punctuation SC", "Inter", ui-sans-serif, system-ui, sans-serif, "Apple Color Emoji", "Segoe UI Emoji", "Segoe UI Symbol", "Noto Color Emoji"}:root{--vp-shadow-1: 0 1px 2px rgba(0, 0, 0, .04), 0 1px 2px rgba(0, 0, 0, .06);--vp-shadow-2: 0 3px 12px rgba(0, 0, 0, .07), 0 1px 4px rgba(0, 0, 0, .07);--vp-shadow-3: 0 12px 32px rgba(0, 0, 0, .1), 0 2px 6px rgba(0, 0, 0, .08);--vp-shadow-4: 0 14px 44px rgba(0, 0, 0, .12), 0 3px 9px rgba(0, 0, 0, .12);--vp-shadow-5: 0 18px 56px rgba(0, 0, 0, .16), 0 4px 12px rgba(0, 0, 0, .16)}:root{--vp-z-index-footer: 10;--vp-z-index-local-nav: 20;--vp-z-index-nav: 30;--vp-z-index-layout-top: 40;--vp-z-index-backdrop: 50;--vp-z-index-sidebar: 60}@media (min-width: 960px){:root{--vp-z-index-sidebar: 25}}:root{--vp-layout-max-width: 1440px}:root{--vp-header-anchor-symbol: "#"}:root{--vp-code-line-height: 1.7;--vp-code-font-size: .875em;--vp-code-color: var(--vp-c-brand-1);--vp-code-link-color: var(--vp-c-brand-1);--vp-code-link-hover-color: var(--vp-c-brand-2);--vp-code-bg: var(--vp-c-default-soft);--vp-code-block-color: var(--vp-c-text-2);--vp-code-block-bg: var(--vp-c-bg-alt);--vp-code-block-divider-color: var(--vp-c-gutter);--vp-code-lang-color: var(--vp-c-text-3);--vp-code-line-highlight-color: var(--vp-c-default-soft);--vp-code-line-number-color: var(--vp-c-text-3);--vp-code-line-diff-add-color: var(--vp-c-success-soft);--vp-code-line-diff-add-symbol-color: var(--vp-c-success-1);--vp-code-line-diff-remove-color: var(--vp-c-danger-soft);--vp-code-line-diff-remove-symbol-color: var(--vp-c-danger-1);--vp-code-line-warning-color: var(--vp-c-warning-soft);--vp-code-line-error-color: var(--vp-c-danger-soft);--vp-code-copy-code-border-color: var(--vp-c-divider);--vp-code-copy-code-bg: var(--vp-c-bg-soft);--vp-code-copy-code-hover-border-color: var(--vp-c-divider);--vp-code-copy-code-hover-bg: var(--vp-c-bg);--vp-code-copy-code-active-text: var(--vp-c-text-2);--vp-code-copy-copied-text-content: "Copied";--vp-code-tab-divider: var(--vp-code-block-divider-color);--vp-code-tab-text-color: var(--vp-c-text-2);--vp-code-tab-bg: var(--vp-code-block-bg);--vp-code-tab-hover-text-color: var(--vp-c-text-1);--vp-code-tab-active-text-color: var(--vp-c-text-1);--vp-code-tab-active-bar-color: var(--vp-c-brand-1)}:lang(es),:lang(pt){--vp-code-copy-copied-text-content: "Copiado"}:lang(fa){--vp-code-copy-copied-text-content: "کپی شد"}:lang(ko){--vp-code-copy-copied-text-content: "복사됨"}:lang(ru){--vp-code-copy-copied-text-content: "Скопировано"}:lang(zh){--vp-code-copy-copied-text-content: "已复制"}:root{--vp-button-brand-border: transparent;--vp-button-brand-text: var(--vp-c-white);--vp-button-brand-bg: var(--vp-c-brand-3);--vp-button-brand-hover-border: transparent;--vp-button-brand-hover-text: var(--vp-c-white);--vp-button-brand-hover-bg: var(--vp-c-brand-2);--vp-button-brand-active-border: transparent;--vp-button-brand-active-text: var(--vp-c-white);--vp-button-brand-active-bg: var(--vp-c-brand-1);--vp-button-alt-border: transparent;--vp-button-alt-text: var(--vp-c-text-1);--vp-button-alt-bg: var(--vp-c-default-3);--vp-button-alt-hover-border: transparent;--vp-button-alt-hover-text: var(--vp-c-text-1);--vp-button-alt-hover-bg: var(--vp-c-default-2);--vp-button-alt-active-border: transparent;--vp-button-alt-active-text: var(--vp-c-text-1);--vp-button-alt-active-bg: var(--vp-c-default-1);--vp-button-sponsor-border: var(--vp-c-text-2);--vp-button-sponsor-text: var(--vp-c-text-2);--vp-button-sponsor-bg: transparent;--vp-button-sponsor-hover-border: var(--vp-c-sponsor);--vp-button-sponsor-hover-text: var(--vp-c-sponsor);--vp-button-sponsor-hover-bg: transparent;--vp-button-sponsor-active-border: var(--vp-c-sponsor);--vp-button-sponsor-active-text: var(--vp-c-sponsor);--vp-button-sponsor-active-bg: transparent}:root{--vp-custom-block-font-size: 14px;--vp-custom-block-code-font-size: 13px;--vp-custom-block-info-border: transparent;--vp-custom-block-info-text: var(--vp-c-text-1);--vp-custom-block-info-bg: var(--vp-c-default-soft);--vp-custom-block-info-code-bg: var(--vp-c-default-soft);--vp-custom-block-note-border: transparent;--vp-custom-block-note-text: var(--vp-c-text-1);--vp-custom-block-note-bg: var(--vp-c-default-soft);--vp-custom-block-note-code-bg: var(--vp-c-default-soft);--vp-custom-block-tip-border: transparent;--vp-custom-block-tip-text: var(--vp-c-text-1);--vp-custom-block-tip-bg: var(--vp-c-tip-soft);--vp-custom-block-tip-code-bg: var(--vp-c-tip-soft);--vp-custom-block-important-border: transparent;--vp-custom-block-important-text: var(--vp-c-text-1);--vp-custom-block-important-bg: var(--vp-c-important-soft);--vp-custom-block-important-code-bg: var(--vp-c-important-soft);--vp-custom-block-warning-border: transparent;--vp-custom-block-warning-text: var(--vp-c-text-1);--vp-custom-block-warning-bg: var(--vp-c-warning-soft);--vp-custom-block-warning-code-bg: var(--vp-c-warning-soft);--vp-custom-block-danger-border: transparent;--vp-custom-block-danger-text: var(--vp-c-text-1);--vp-custom-block-danger-bg: var(--vp-c-danger-soft);--vp-custom-block-danger-code-bg: var(--vp-c-danger-soft);--vp-custom-block-caution-border: transparent;--vp-custom-block-caution-text: var(--vp-c-text-1);--vp-custom-block-caution-bg: var(--vp-c-caution-soft);--vp-custom-block-caution-code-bg: var(--vp-c-caution-soft);--vp-custom-block-details-border: var(--vp-custom-block-info-border);--vp-custom-block-details-text: var(--vp-custom-block-info-text);--vp-custom-block-details-bg: var(--vp-custom-block-info-bg);--vp-custom-block-details-code-bg: var(--vp-custom-block-info-code-bg)}:root{--vp-input-border-color: var(--vp-c-border);--vp-input-bg-color: var(--vp-c-bg-alt);--vp-input-switch-bg-color: var(--vp-c-default-soft)}:root{--vp-nav-height: 64px;--vp-nav-bg-color: var(--vp-c-bg);--vp-nav-screen-bg-color: var(--vp-c-bg);--vp-nav-logo-height: 24px}.hide-nav{--vp-nav-height: 0px}.hide-nav .VPSidebar{--vp-nav-height: 22px}:root{--vp-local-nav-bg-color: var(--vp-c-bg)}:root{--vp-sidebar-width: 272px;--vp-sidebar-bg-color: var(--vp-c-bg-alt)}:root{--vp-backdrop-bg-color: rgba(0, 0, 0, .6)}:root{--vp-home-hero-name-color: var(--vp-c-brand-1);--vp-home-hero-name-background: transparent;--vp-home-hero-image-background-image: none;--vp-home-hero-image-filter: none}:root{--vp-badge-info-border: transparent;--vp-badge-info-text: var(--vp-c-text-2);--vp-badge-info-bg: var(--vp-c-default-soft);--vp-badge-tip-border: transparent;--vp-badge-tip-text: var(--vp-c-tip-1);--vp-badge-tip-bg: var(--vp-c-tip-soft);--vp-badge-warning-border: transparent;--vp-badge-warning-text: var(--vp-c-warning-1);--vp-badge-warning-bg: var(--vp-c-warning-soft);--vp-badge-danger-border: transparent;--vp-badge-danger-text: var(--vp-c-danger-1);--vp-badge-danger-bg: var(--vp-c-danger-soft)}:root{--vp-carbon-ads-text-color: var(--vp-c-text-1);--vp-carbon-ads-poweredby-color: var(--vp-c-text-2);--vp-carbon-ads-bg-color: var(--vp-c-bg-soft);--vp-carbon-ads-hover-text-color: var(--vp-c-brand-1);--vp-carbon-ads-hover-poweredby-color: var(--vp-c-text-1)}:root{--vp-local-search-bg: var(--vp-c-bg);--vp-local-search-result-bg: var(--vp-c-bg);--vp-local-search-result-border: var(--vp-c-divider);--vp-local-search-result-selected-bg: var(--vp-c-bg);--vp-local-search-result-selected-border: var(--vp-c-brand-1);--vp-local-search-highlight-bg: var(--vp-c-brand-1);--vp-local-search-highlight-text: var(--vp-c-neutral-inverse)}@media (prefers-reduced-motion: reduce){*,:before,:after{animation-delay:-1ms!important;animation-duration:1ms!important;animation-iteration-count:1!important;background-attachment:initial!important;scroll-behavior:auto!important;transition-duration:0s!important;transition-delay:0s!important}}*,:before,:after{box-sizing:border-box}html{line-height:1.4;font-size:16px;-webkit-text-size-adjust:100%}html.dark{color-scheme:dark}body{margin:0;width:100%;min-width:320px;min-height:100vh;line-height:24px;font-family:var(--vp-font-family-base);font-size:16px;font-weight:400;color:var(--vp-c-text-1);background-color:var(--vp-c-bg);font-synthesis:style;text-rendering:optimizeLegibility;-webkit-font-smoothing:antialiased;-moz-osx-font-smoothing:grayscale}main{display:block}h1,h2,h3,h4,h5,h6{margin:0;line-height:24px;font-size:16px;font-weight:400}p{margin:0}strong,b{font-weight:600}a,area,button,[role=button],input,label,select,summary,textarea{touch-action:manipulation}a{color:inherit;text-decoration:inherit}ol,ul{list-style:none;margin:0;padding:0}blockquote{margin:0}pre,code,kbd,samp{font-family:var(--vp-font-family-mono)}img,svg,video,canvas,audio,iframe,embed,object{display:block}figure{margin:0}img,video{max-width:100%;height:auto}button,input,optgroup,select,textarea{border:0;padding:0;line-height:inherit;color:inherit}button{padding:0;font-family:inherit;background-color:transparent;background-image:none}button:enabled,[role=button]:enabled{cursor:pointer}button:focus,button:focus-visible{outline:1px dotted;outline:4px auto -webkit-focus-ring-color}button:focus:not(:focus-visible){outline:none!important}input:focus,textarea:focus,select:focus{outline:none}table{border-collapse:collapse}input{background-color:transparent}input:-ms-input-placeholder,textarea:-ms-input-placeholder{color:var(--vp-c-text-3)}input::-ms-input-placeholder,textarea::-ms-input-placeholder{color:var(--vp-c-text-3)}input::placeholder,textarea::placeholder{color:var(--vp-c-text-3)}input::-webkit-outer-spin-button,input::-webkit-inner-spin-button{-webkit-appearance:none;margin:0}input[type=number]{-moz-appearance:textfield}textarea{resize:vertical}select{-webkit-appearance:none}fieldset{margin:0;padding:0}h1,h2,h3,h4,h5,h6,li,p{overflow-wrap:break-word}vite-error-overlay{z-index:9999}mjx-container{overflow-x:auto}mjx-container>svg{display:inline-block;margin:auto}[class^=vpi-],[class*=" vpi-"],.vp-icon{width:1em;height:1em}[class^=vpi-].bg,[class*=" vpi-"].bg,.vp-icon.bg{background-size:100% 100%;background-color:transparent}[class^=vpi-]:not(.bg),[class*=" vpi-"]:not(.bg),.vp-icon:not(.bg){-webkit-mask:var(--icon) no-repeat;mask:var(--icon) no-repeat;-webkit-mask-size:100% 100%;mask-size:100% 100%;background-color:currentColor;color:inherit}.vpi-align-left{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='M21 6H3M15 12H3M17 18H3'/%3E%3C/svg%3E")}.vpi-arrow-right,.vpi-arrow-down,.vpi-arrow-left,.vpi-arrow-up{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='M5 12h14M12 5l7 7-7 7'/%3E%3C/svg%3E")}.vpi-chevron-right,.vpi-chevron-down,.vpi-chevron-left,.vpi-chevron-up{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='m9 18 6-6-6-6'/%3E%3C/svg%3E")}.vpi-chevron-down,.vpi-arrow-down{transform:rotate(90deg)}.vpi-chevron-left,.vpi-arrow-left{transform:rotate(180deg)}.vpi-chevron-up,.vpi-arrow-up{transform:rotate(-90deg)}.vpi-square-pen{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='M12 3H5a2 2 0 0 0-2 2v14a2 2 0 0 0 2 2h14a2 2 0 0 0 2-2v-7'/%3E%3Cpath d='M18.375 2.625a2.121 2.121 0 1 1 3 3L12 15l-4 1 1-4Z'/%3E%3C/svg%3E")}.vpi-plus{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='M5 12h14M12 5v14'/%3E%3C/svg%3E")}.vpi-sun{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Ccircle cx='12' cy='12' r='4'/%3E%3Cpath d='M12 2v2M12 20v2M4.93 4.93l1.41 1.41M17.66 17.66l1.41 1.41M2 12h2M20 12h2M6.34 17.66l-1.41 1.41M19.07 4.93l-1.41 1.41'/%3E%3C/svg%3E")}.vpi-moon{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='M12 3a6 6 0 0 0 9 9 9 9 0 1 1-9-9Z'/%3E%3C/svg%3E")}.vpi-more-horizontal{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Ccircle cx='12' cy='12' r='1'/%3E%3Ccircle cx='19' cy='12' r='1'/%3E%3Ccircle cx='5' cy='12' r='1'/%3E%3C/svg%3E")}.vpi-languages{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='m5 8 6 6M4 14l6-6 2-3M2 5h12M7 2h1M22 22l-5-10-5 10M14 18h6'/%3E%3C/svg%3E")}.vpi-heart{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='M19 14c1.49-1.46 3-3.21 3-5.5A5.5 5.5 0 0 0 16.5 3c-1.76 0-3 .5-4.5 2-1.5-1.5-2.74-2-4.5-2A5.5 5.5 0 0 0 2 8.5c0 2.3 1.5 4.05 3 5.5l7 7Z'/%3E%3C/svg%3E")}.vpi-search{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Ccircle cx='11' cy='11' r='8'/%3E%3Cpath d='m21 21-4.3-4.3'/%3E%3C/svg%3E")}.vpi-layout-list{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Crect width='7' height='7' x='3' y='3' rx='1'/%3E%3Crect width='7' height='7' x='3' y='14' rx='1'/%3E%3Cpath d='M14 4h7M14 9h7M14 15h7M14 20h7'/%3E%3C/svg%3E")}.vpi-delete{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='M20 5H9l-7 7 7 7h11a2 2 0 0 0 2-2V7a2 2 0 0 0-2-2ZM18 9l-6 6M12 9l6 6'/%3E%3C/svg%3E")}.vpi-corner-down-left{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Cpath d='m9 10-5 5 5 5'/%3E%3Cpath d='M20 4v7a4 4 0 0 1-4 4H4'/%3E%3C/svg%3E")}:root{--vp-icon-copy: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='rgba(128,128,128,1)' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Crect width='8' height='4' x='8' y='2' rx='1' ry='1'/%3E%3Cpath d='M16 4h2a2 2 0 0 1 2 2v14a2 2 0 0 1-2 2H6a2 2 0 0 1-2-2V6a2 2 0 0 1 2-2h2'/%3E%3C/svg%3E");--vp-icon-copied: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' fill='none' stroke='rgba(128,128,128,1)' stroke-linecap='round' stroke-linejoin='round' stroke-width='2' viewBox='0 0 24 24'%3E%3Crect width='8' height='4' x='8' y='2' rx='1' ry='1'/%3E%3Cpath d='M16 4h2a2 2 0 0 1 2 2v14a2 2 0 0 1-2 2H6a2 2 0 0 1-2-2V6a2 2 0 0 1 2-2h2'/%3E%3Cpath d='m9 14 2 2 4-4'/%3E%3C/svg%3E")}.visually-hidden{position:absolute;width:1px;height:1px;white-space:nowrap;clip:rect(0 0 0 0);clip-path:inset(50%);overflow:hidden}.custom-block{border:1px solid transparent;border-radius:8px;padding:16px 16px 8px;line-height:24px;font-size:var(--vp-custom-block-font-size);color:var(--vp-c-text-2)}.custom-block.info{border-color:var(--vp-custom-block-info-border);color:var(--vp-custom-block-info-text);background-color:var(--vp-custom-block-info-bg)}.custom-block.info a,.custom-block.info code{color:var(--vp-c-brand-1)}.custom-block.info a:hover,.custom-block.info a:hover>code{color:var(--vp-c-brand-2)}.custom-block.info code{background-color:var(--vp-custom-block-info-code-bg)}.custom-block.note{border-color:var(--vp-custom-block-note-border);color:var(--vp-custom-block-note-text);background-color:var(--vp-custom-block-note-bg)}.custom-block.note a,.custom-block.note code{color:var(--vp-c-brand-1)}.custom-block.note a:hover,.custom-block.note a:hover>code{color:var(--vp-c-brand-2)}.custom-block.note code{background-color:var(--vp-custom-block-note-code-bg)}.custom-block.tip{border-color:var(--vp-custom-block-tip-border);color:var(--vp-custom-block-tip-text);background-color:var(--vp-custom-block-tip-bg)}.custom-block.tip a,.custom-block.tip code{color:var(--vp-c-tip-1)}.custom-block.tip a:hover,.custom-block.tip a:hover>code{color:var(--vp-c-tip-2)}.custom-block.tip code{background-color:var(--vp-custom-block-tip-code-bg)}.custom-block.important{border-color:var(--vp-custom-block-important-border);color:var(--vp-custom-block-important-text);background-color:var(--vp-custom-block-important-bg)}.custom-block.important a,.custom-block.important code{color:var(--vp-c-important-1)}.custom-block.important a:hover,.custom-block.important a:hover>code{color:var(--vp-c-important-2)}.custom-block.important code{background-color:var(--vp-custom-block-important-code-bg)}.custom-block.warning{border-color:var(--vp-custom-block-warning-border);color:var(--vp-custom-block-warning-text);background-color:var(--vp-custom-block-warning-bg)}.custom-block.warning a,.custom-block.warning code{color:var(--vp-c-warning-1)}.custom-block.warning a:hover,.custom-block.warning a:hover>code{color:var(--vp-c-warning-2)}.custom-block.warning code{background-color:var(--vp-custom-block-warning-code-bg)}.custom-block.danger{border-color:var(--vp-custom-block-danger-border);color:var(--vp-custom-block-danger-text);background-color:var(--vp-custom-block-danger-bg)}.custom-block.danger a,.custom-block.danger code{color:var(--vp-c-danger-1)}.custom-block.danger a:hover,.custom-block.danger a:hover>code{color:var(--vp-c-danger-2)}.custom-block.danger code{background-color:var(--vp-custom-block-danger-code-bg)}.custom-block.caution{border-color:var(--vp-custom-block-caution-border);color:var(--vp-custom-block-caution-text);background-color:var(--vp-custom-block-caution-bg)}.custom-block.caution a,.custom-block.caution code{color:var(--vp-c-caution-1)}.custom-block.caution a:hover,.custom-block.caution a:hover>code{color:var(--vp-c-caution-2)}.custom-block.caution code{background-color:var(--vp-custom-block-caution-code-bg)}.custom-block.details{border-color:var(--vp-custom-block-details-border);color:var(--vp-custom-block-details-text);background-color:var(--vp-custom-block-details-bg)}.custom-block.details a{color:var(--vp-c-brand-1)}.custom-block.details a:hover,.custom-block.details a:hover>code{color:var(--vp-c-brand-2)}.custom-block.details code{background-color:var(--vp-custom-block-details-code-bg)}.custom-block-title{font-weight:600}.custom-block p+p{margin:8px 0}.custom-block.details summary{margin:0 0 8px;font-weight:700;cursor:pointer;-webkit-user-select:none;user-select:none}.custom-block.details summary+p{margin:8px 0}.custom-block a{color:inherit;font-weight:600;text-decoration:underline;text-underline-offset:2px;transition:opacity .25s}.custom-block a:hover{opacity:.75}.custom-block code{font-size:var(--vp-custom-block-code-font-size)}.custom-block.custom-block th,.custom-block.custom-block blockquote>p{font-size:var(--vp-custom-block-font-size);color:inherit}.dark .vp-code span{color:var(--shiki-dark, inherit)}html:not(.dark) .vp-code span{color:var(--shiki-light, inherit)}.vp-code-group{margin-top:16px}.vp-code-group .tabs{position:relative;display:flex;margin-right:-24px;margin-left:-24px;padding:0 12px;background-color:var(--vp-code-tab-bg);overflow-x:auto;overflow-y:hidden;box-shadow:inset 0 -1px var(--vp-code-tab-divider)}@media (min-width: 640px){.vp-code-group .tabs{margin-right:0;margin-left:0;border-radius:8px 8px 0 0}}.vp-code-group .tabs input{position:fixed;opacity:0;pointer-events:none}.vp-code-group .tabs label{position:relative;display:inline-block;border-bottom:1px solid transparent;padding:0 12px;line-height:48px;font-size:14px;font-weight:500;color:var(--vp-code-tab-text-color);white-space:nowrap;cursor:pointer;transition:color .25s}.vp-code-group .tabs label:after{position:absolute;right:8px;bottom:-1px;left:8px;z-index:1;height:2px;border-radius:2px;content:"";background-color:transparent;transition:background-color .25s}.vp-code-group label:hover{color:var(--vp-code-tab-hover-text-color)}.vp-code-group input:checked+label{color:var(--vp-code-tab-active-text-color)}.vp-code-group input:checked+label:after{background-color:var(--vp-code-tab-active-bar-color)}.vp-code-group div[class*=language-],.vp-block{display:none;margin-top:0!important;border-top-left-radius:0!important;border-top-right-radius:0!important}.vp-code-group div[class*=language-].active,.vp-block.active{display:block}.vp-block{padding:20px 24px}.vp-doc h1,.vp-doc h2,.vp-doc h3,.vp-doc h4,.vp-doc h5,.vp-doc h6{position:relative;font-weight:600;outline:none}.vp-doc h1{letter-spacing:-.02em;line-height:40px;font-size:28px}.vp-doc h2{margin:48px 0 16px;border-top:1px solid var(--vp-c-divider);padding-top:24px;letter-spacing:-.02em;line-height:32px;font-size:24px}.vp-doc h3{margin:32px 0 0;letter-spacing:-.01em;line-height:28px;font-size:20px}.vp-doc h4{margin:24px 0 0;letter-spacing:-.01em;line-height:24px;font-size:18px}.vp-doc .header-anchor{position:absolute;top:0;left:0;margin-left:-.87em;font-weight:500;-webkit-user-select:none;user-select:none;opacity:0;text-decoration:none;transition:color .25s,opacity .25s}.vp-doc .header-anchor:before{content:var(--vp-header-anchor-symbol)}.vp-doc h1:hover .header-anchor,.vp-doc h1 .header-anchor:focus,.vp-doc h2:hover .header-anchor,.vp-doc h2 .header-anchor:focus,.vp-doc h3:hover .header-anchor,.vp-doc h3 .header-anchor:focus,.vp-doc h4:hover .header-anchor,.vp-doc h4 .header-anchor:focus,.vp-doc h5:hover .header-anchor,.vp-doc h5 .header-anchor:focus,.vp-doc h6:hover .header-anchor,.vp-doc h6 .header-anchor:focus{opacity:1}@media (min-width: 768px){.vp-doc h1{letter-spacing:-.02em;line-height:40px;font-size:32px}}.vp-doc h2 .header-anchor{top:24px}.vp-doc p,.vp-doc summary{margin:16px 0}.vp-doc p{line-height:28px}.vp-doc blockquote{margin:16px 0;border-left:2px solid var(--vp-c-divider);padding-left:16px;transition:border-color .5s;color:var(--vp-c-text-2)}.vp-doc blockquote>p{margin:0;font-size:16px;transition:color .5s}.vp-doc a{font-weight:500;color:var(--vp-c-brand-1);text-decoration:underline;text-underline-offset:2px;transition:color .25s,opacity .25s}.vp-doc a:hover{color:var(--vp-c-brand-2)}.vp-doc strong{font-weight:600}.vp-doc ul,.vp-doc ol{padding-left:1.25rem;margin:16px 0}.vp-doc ul{list-style:disc}.vp-doc ol{list-style:decimal}.vp-doc li+li{margin-top:8px}.vp-doc li>ol,.vp-doc li>ul{margin:8px 0 0}.vp-doc table{display:block;border-collapse:collapse;margin:20px 0;overflow-x:auto}.vp-doc tr{background-color:var(--vp-c-bg);border-top:1px solid var(--vp-c-divider);transition:background-color .5s}.vp-doc tr:nth-child(2n){background-color:var(--vp-c-bg-soft)}.vp-doc th,.vp-doc td{border:1px solid var(--vp-c-divider);padding:8px 16px}.vp-doc th{text-align:left;font-size:14px;font-weight:600;color:var(--vp-c-text-2);background-color:var(--vp-c-bg-soft)}.vp-doc td{font-size:14px}.vp-doc hr{margin:16px 0;border:none;border-top:1px solid var(--vp-c-divider)}.vp-doc .custom-block{margin:16px 0}.vp-doc .custom-block p{margin:8px 0;line-height:24px}.vp-doc .custom-block p:first-child{margin:0}.vp-doc .custom-block div[class*=language-]{margin:8px 0;border-radius:8px}.vp-doc .custom-block div[class*=language-] code{font-weight:400;background-color:transparent}.vp-doc .custom-block .vp-code-group .tabs{margin:0;border-radius:8px 8px 0 0}.vp-doc :not(pre,h1,h2,h3,h4,h5,h6)>code{font-size:var(--vp-code-font-size);color:var(--vp-code-color)}.vp-doc :not(pre)>code{border-radius:4px;padding:3px 6px;background-color:var(--vp-code-bg);transition:color .25s,background-color .5s}.vp-doc a>code{color:var(--vp-code-link-color)}.vp-doc a:hover>code{color:var(--vp-code-link-hover-color)}.vp-doc h1>code,.vp-doc h2>code,.vp-doc h3>code,.vp-doc h4>code{font-size:.9em}.vp-doc div[class*=language-],.vp-block{position:relative;margin:16px -24px;background-color:var(--vp-code-block-bg);overflow-x:auto;transition:background-color .5s}@media (min-width: 640px){.vp-doc div[class*=language-],.vp-block{border-radius:8px;margin:16px 0}}@media (max-width: 639px){.vp-doc li div[class*=language-]{border-radius:8px 0 0 8px}}.vp-doc div[class*=language-]+div[class*=language-],.vp-doc div[class$=-api]+div[class*=language-],.vp-doc div[class*=language-]+div[class$=-api]>div[class*=language-]{margin-top:-8px}.vp-doc [class*=language-] pre,.vp-doc [class*=language-] code{direction:ltr;text-align:left;white-space:pre;word-spacing:normal;word-break:normal;word-wrap:normal;-moz-tab-size:4;-o-tab-size:4;tab-size:4;-webkit-hyphens:none;-moz-hyphens:none;-ms-hyphens:none;hyphens:none}.vp-doc [class*=language-] pre{position:relative;z-index:1;margin:0;padding:20px 0;background:transparent;overflow-x:auto}.vp-doc [class*=language-] code{display:block;padding:0 24px;width:fit-content;min-width:100%;line-height:var(--vp-code-line-height);font-size:var(--vp-code-font-size);color:var(--vp-code-block-color);transition:color .5s}.vp-doc [class*=language-] code .highlighted{background-color:var(--vp-code-line-highlight-color);transition:background-color .5s;margin:0 -24px;padding:0 24px;width:calc(100% + 48px);display:inline-block}.vp-doc [class*=language-] code .highlighted.error{background-color:var(--vp-code-line-error-color)}.vp-doc [class*=language-] code .highlighted.warning{background-color:var(--vp-code-line-warning-color)}.vp-doc [class*=language-] code .diff{transition:background-color .5s;margin:0 -24px;padding:0 24px;width:calc(100% + 48px);display:inline-block}.vp-doc [class*=language-] code .diff:before{position:absolute;left:10px}.vp-doc [class*=language-] .has-focused-lines .line:not(.has-focus){filter:blur(.095rem);opacity:.4;transition:filter .35s,opacity .35s}.vp-doc [class*=language-] .has-focused-lines .line:not(.has-focus){opacity:.7;transition:filter .35s,opacity .35s}.vp-doc [class*=language-]:hover .has-focused-lines .line:not(.has-focus){filter:blur(0);opacity:1}.vp-doc [class*=language-] code .diff.remove{background-color:var(--vp-code-line-diff-remove-color);opacity:.7}.vp-doc [class*=language-] code .diff.remove:before{content:"-";color:var(--vp-code-line-diff-remove-symbol-color)}.vp-doc [class*=language-] code .diff.add{background-color:var(--vp-code-line-diff-add-color)}.vp-doc [class*=language-] code .diff.add:before{content:"+";color:var(--vp-code-line-diff-add-symbol-color)}.vp-doc div[class*=language-].line-numbers-mode{padding-left:32px}.vp-doc .line-numbers-wrapper{position:absolute;top:0;bottom:0;left:0;z-index:3;border-right:1px solid var(--vp-code-block-divider-color);padding-top:20px;width:32px;text-align:center;font-family:var(--vp-font-family-mono);line-height:var(--vp-code-line-height);font-size:var(--vp-code-font-size);color:var(--vp-code-line-number-color);transition:border-color .5s,color .5s}.vp-doc [class*=language-]>button.copy{direction:ltr;position:absolute;top:12px;right:12px;z-index:3;border:1px solid var(--vp-code-copy-code-border-color);border-radius:4px;width:40px;height:40px;background-color:var(--vp-code-copy-code-bg);opacity:0;cursor:pointer;background-image:var(--vp-icon-copy);background-position:50%;background-size:20px;background-repeat:no-repeat;transition:border-color .25s,background-color .25s,opacity .25s}.vp-doc [class*=language-]:hover>button.copy,.vp-doc [class*=language-]>button.copy:focus{opacity:1}.vp-doc [class*=language-]>button.copy:hover,.vp-doc [class*=language-]>button.copy.copied{border-color:var(--vp-code-copy-code-hover-border-color);background-color:var(--vp-code-copy-code-hover-bg)}.vp-doc [class*=language-]>button.copy.copied,.vp-doc [class*=language-]>button.copy:hover.copied{border-radius:0 4px 4px 0;background-color:var(--vp-code-copy-code-hover-bg);background-image:var(--vp-icon-copied)}.vp-doc [class*=language-]>button.copy.copied:before,.vp-doc [class*=language-]>button.copy:hover.copied:before{position:relative;top:-1px;transform:translate(calc(-100% - 1px));display:flex;justify-content:center;align-items:center;border:1px solid var(--vp-code-copy-code-hover-border-color);border-right:0;border-radius:4px 0 0 4px;padding:0 10px;width:fit-content;height:40px;text-align:center;font-size:12px;font-weight:500;color:var(--vp-code-copy-code-active-text);background-color:var(--vp-code-copy-code-hover-bg);white-space:nowrap;content:var(--vp-code-copy-copied-text-content)}.vp-doc [class*=language-]>span.lang{position:absolute;top:2px;right:8px;z-index:2;font-size:12px;font-weight:500;-webkit-user-select:none;user-select:none;color:var(--vp-code-lang-color);transition:color .4s,opacity .4s}.vp-doc [class*=language-]:hover>button.copy+span.lang,.vp-doc [class*=language-]>button.copy:focus+span.lang{opacity:0}.vp-doc .VPTeamMembers{margin-top:24px}.vp-doc .VPTeamMembers.small.count-1 .container{margin:0!important;max-width:calc((100% - 24px)/2)!important}.vp-doc .VPTeamMembers.small.count-2 .container,.vp-doc .VPTeamMembers.small.count-3 .container{max-width:100%!important}.vp-doc .VPTeamMembers.medium.count-1 .container{margin:0!important;max-width:calc((100% - 24px)/2)!important}:is(.vp-external-link-icon,.vp-doc a[href*="://"],.vp-doc a[target=_blank]):not(:is(.no-icon,svg a,:has(img,svg))):after{display:inline-block;margin-top:-1px;margin-left:4px;width:11px;height:11px;background:currentColor;color:var(--vp-c-text-3);flex-shrink:0;--icon: url("data:image/svg+xml, %3Csvg xmlns='http://www.w3.org/2000/svg' viewBox='0 0 24 24' %3E%3Cpath d='M0 0h24v24H0V0z' fill='none' /%3E%3Cpath d='M9 5v2h6.59L4 18.59 5.41 20 17 8.41V15h2V5H9z' /%3E%3C/svg%3E");-webkit-mask-image:var(--icon);mask-image:var(--icon)}.vp-external-link-icon:after{content:""}.external-link-icon-enabled :is(.vp-doc a[href*="://"],.vp-doc a[target=_blank]):not(:is(.no-icon,svg a,:has(img,svg))):after{content:"";color:currentColor}.vp-sponsor{border-radius:16px;overflow:hidden}.vp-sponsor.aside{border-radius:12px}.vp-sponsor-section+.vp-sponsor-section{margin-top:4px}.vp-sponsor-tier{margin:0 0 4px!important;text-align:center;letter-spacing:1px!important;line-height:24px;width:100%;font-weight:600;color:var(--vp-c-text-2);background-color:var(--vp-c-bg-soft)}.vp-sponsor.normal .vp-sponsor-tier{padding:13px 0 11px;font-size:14px}.vp-sponsor.aside .vp-sponsor-tier{padding:9px 0 7px;font-size:12px}.vp-sponsor-grid+.vp-sponsor-tier{margin-top:4px}.vp-sponsor-grid{display:flex;flex-wrap:wrap;gap:4px}.vp-sponsor-grid.xmini .vp-sponsor-grid-link{height:64px}.vp-sponsor-grid.xmini .vp-sponsor-grid-image{max-width:64px;max-height:22px}.vp-sponsor-grid.mini .vp-sponsor-grid-link{height:72px}.vp-sponsor-grid.mini .vp-sponsor-grid-image{max-width:96px;max-height:24px}.vp-sponsor-grid.small .vp-sponsor-grid-link{height:96px}.vp-sponsor-grid.small .vp-sponsor-grid-image{max-width:96px;max-height:24px}.vp-sponsor-grid.medium .vp-sponsor-grid-link{height:112px}.vp-sponsor-grid.medium .vp-sponsor-grid-image{max-width:120px;max-height:36px}.vp-sponsor-grid.big .vp-sponsor-grid-link{height:184px}.vp-sponsor-grid.big .vp-sponsor-grid-image{max-width:192px;max-height:56px}.vp-sponsor-grid[data-vp-grid="2"] .vp-sponsor-grid-item{width:calc((100% - 4px)/2)}.vp-sponsor-grid[data-vp-grid="3"] .vp-sponsor-grid-item{width:calc((100% - 4px * 2) / 3)}.vp-sponsor-grid[data-vp-grid="4"] .vp-sponsor-grid-item{width:calc((100% - 12px)/4)}.vp-sponsor-grid[data-vp-grid="5"] .vp-sponsor-grid-item{width:calc((100% - 16px)/5)}.vp-sponsor-grid[data-vp-grid="6"] .vp-sponsor-grid-item{width:calc((100% - 4px * 5) / 6)}.vp-sponsor-grid-item{flex-shrink:0;width:100%;background-color:var(--vp-c-bg-soft);transition:background-color .25s}.vp-sponsor-grid-item:hover{background-color:var(--vp-c-default-soft)}.vp-sponsor-grid-item:hover .vp-sponsor-grid-image{filter:grayscale(0) invert(0)}.vp-sponsor-grid-item.empty:hover{background-color:var(--vp-c-bg-soft)}.dark .vp-sponsor-grid-item:hover{background-color:var(--vp-c-white)}.dark .vp-sponsor-grid-item.empty:hover{background-color:var(--vp-c-bg-soft)}.vp-sponsor-grid-link{display:flex}.vp-sponsor-grid-box{display:flex;justify-content:center;align-items:center;width:100%}.vp-sponsor-grid-image{max-width:100%;filter:grayscale(1);transition:filter .25s}.dark .vp-sponsor-grid-image{filter:grayscale(1) invert(1)}.VPBadge{display:inline-block;margin-left:2px;border:1px solid transparent;border-radius:12px;padding:0 10px;line-height:22px;font-size:12px;font-weight:500;transform:translateY(-2px)}.VPBadge.small{padding:0 6px;line-height:18px;font-size:10px;transform:translateY(-8px)}.VPDocFooter .VPBadge{display:none}.vp-doc h1>.VPBadge{margin-top:4px;vertical-align:top}.vp-doc h2>.VPBadge{margin-top:3px;padding:0 8px;vertical-align:top}.vp-doc h3>.VPBadge{vertical-align:middle}.vp-doc h4>.VPBadge,.vp-doc h5>.VPBadge,.vp-doc h6>.VPBadge{vertical-align:middle;line-height:18px}.VPBadge.info{border-color:var(--vp-badge-info-border);color:var(--vp-badge-info-text);background-color:var(--vp-badge-info-bg)}.VPBadge.tip{border-color:var(--vp-badge-tip-border);color:var(--vp-badge-tip-text);background-color:var(--vp-badge-tip-bg)}.VPBadge.warning{border-color:var(--vp-badge-warning-border);color:var(--vp-badge-warning-text);background-color:var(--vp-badge-warning-bg)}.VPBadge.danger{border-color:var(--vp-badge-danger-border);color:var(--vp-badge-danger-text);background-color:var(--vp-badge-danger-bg)}.VPBackdrop[data-v-54a304ca]{position:fixed;top:0;right:0;bottom:0;left:0;z-index:var(--vp-z-index-backdrop);background:var(--vp-backdrop-bg-color);transition:opacity .5s}.VPBackdrop.fade-enter-from[data-v-54a304ca],.VPBackdrop.fade-leave-to[data-v-54a304ca]{opacity:0}.VPBackdrop.fade-leave-active[data-v-54a304ca]{transition-duration:.25s}@media (min-width: 1280px){.VPBackdrop[data-v-54a304ca]{display:none}}.NotFound[data-v-6ff51ddd]{padding:64px 24px 96px;text-align:center}@media (min-width: 768px){.NotFound[data-v-6ff51ddd]{padding:96px 32px 168px}}.code[data-v-6ff51ddd]{line-height:64px;font-size:64px;font-weight:600}.title[data-v-6ff51ddd]{padding-top:12px;letter-spacing:2px;line-height:20px;font-size:20px;font-weight:700}.divider[data-v-6ff51ddd]{margin:24px auto 18px;width:64px;height:1px;background-color:var(--vp-c-divider)}.quote[data-v-6ff51ddd]{margin:0 auto;max-width:256px;font-size:14px;font-weight:500;color:var(--vp-c-text-2)}.action[data-v-6ff51ddd]{padding-top:20px}.link[data-v-6ff51ddd]{display:inline-block;border:1px solid var(--vp-c-brand-1);border-radius:16px;padding:3px 16px;font-size:14px;font-weight:500;color:var(--vp-c-brand-1);transition:border-color .25s,color .25s}.link[data-v-6ff51ddd]:hover{border-color:var(--vp-c-brand-2);color:var(--vp-c-brand-2)}.root[data-v-53c99d69]{position:relative;z-index:1}.nested[data-v-53c99d69]{padding-right:16px;padding-left:16px}.outline-link[data-v-53c99d69]{display:block;line-height:32px;font-size:14px;font-weight:400;color:var(--vp-c-text-2);white-space:nowrap;overflow:hidden;text-overflow:ellipsis;transition:color .5s}.outline-link[data-v-53c99d69]:hover,.outline-link.active[data-v-53c99d69]{color:var(--vp-c-text-1);transition:color .25s}.outline-link.nested[data-v-53c99d69]{padding-left:13px}.VPDocAsideOutline[data-v-f610f197]{display:none}.VPDocAsideOutline.has-outline[data-v-f610f197]{display:block}.content[data-v-f610f197]{position:relative;border-left:1px solid var(--vp-c-divider);padding-left:16px;font-size:13px;font-weight:500}.outline-marker[data-v-f610f197]{position:absolute;top:32px;left:-1px;z-index:0;opacity:0;width:2px;border-radius:2px;height:18px;background-color:var(--vp-c-brand-1);transition:top .25s cubic-bezier(0,1,.5,1),background-color .5s,opacity .25s}.outline-title[data-v-f610f197]{line-height:32px;font-size:14px;font-weight:600}.VPDocAside[data-v-cb998dce]{display:flex;flex-direction:column;flex-grow:1}.spacer[data-v-cb998dce]{flex-grow:1}.VPDocAside[data-v-cb998dce] .spacer+.VPDocAsideSponsors,.VPDocAside[data-v-cb998dce] .spacer+.VPDocAsideCarbonAds{margin-top:24px}.VPDocAside[data-v-cb998dce] .VPDocAsideSponsors+.VPDocAsideCarbonAds{margin-top:16px}.VPLastUpdated[data-v-1bb0c8a8]{line-height:24px;font-size:14px;font-weight:500;color:var(--vp-c-text-2)}@media (min-width: 640px){.VPLastUpdated[data-v-1bb0c8a8]{line-height:32px;font-size:14px;font-weight:500}}.VPDocFooter[data-v-1bcd8184]{margin-top:64px}.edit-info[data-v-1bcd8184]{padding-bottom:18px}@media (min-width: 640px){.edit-info[data-v-1bcd8184]{display:flex;justify-content:space-between;align-items:center;padding-bottom:14px}}.edit-link-button[data-v-1bcd8184]{display:flex;align-items:center;border:0;line-height:32px;font-size:14px;font-weight:500;color:var(--vp-c-brand-1);transition:color .25s}.edit-link-button[data-v-1bcd8184]:hover{color:var(--vp-c-brand-2)}.edit-link-icon[data-v-1bcd8184]{margin-right:8px}.prev-next[data-v-1bcd8184]{border-top:1px solid var(--vp-c-divider);padding-top:24px;display:grid;grid-row-gap:8px}@media (min-width: 640px){.prev-next[data-v-1bcd8184]{grid-template-columns:repeat(2,1fr);grid-column-gap:16px}}.pager-link[data-v-1bcd8184]{display:block;border:1px solid var(--vp-c-divider);border-radius:8px;padding:11px 16px 13px;width:100%;height:100%;transition:border-color .25s}.pager-link[data-v-1bcd8184]:hover{border-color:var(--vp-c-brand-1)}.pager-link.next[data-v-1bcd8184]{margin-left:auto;text-align:right}.desc[data-v-1bcd8184]{display:block;line-height:20px;font-size:12px;font-weight:500;color:var(--vp-c-text-2)}.title[data-v-1bcd8184]{display:block;line-height:20px;font-size:14px;font-weight:500;color:var(--vp-c-brand-1);transition:color .25s}.VPDoc[data-v-e6f2a212]{padding:32px 24px 96px;width:100%}@media (min-width: 768px){.VPDoc[data-v-e6f2a212]{padding:48px 32px 128px}}@media (min-width: 960px){.VPDoc[data-v-e6f2a212]{padding:48px 32px 0}.VPDoc:not(.has-sidebar) .container[data-v-e6f2a212]{display:flex;justify-content:center;max-width:992px}.VPDoc:not(.has-sidebar) .content[data-v-e6f2a212]{max-width:752px}}@media (min-width: 1280px){.VPDoc .container[data-v-e6f2a212]{display:flex;justify-content:center}.VPDoc .aside[data-v-e6f2a212]{display:block}}@media (min-width: 1440px){.VPDoc:not(.has-sidebar) .content[data-v-e6f2a212]{max-width:784px}.VPDoc:not(.has-sidebar) .container[data-v-e6f2a212]{max-width:1104px}}.container[data-v-e6f2a212]{margin:0 auto;width:100%}.aside[data-v-e6f2a212]{position:relative;display:none;order:2;flex-grow:1;padding-left:32px;width:100%;max-width:256px}.left-aside[data-v-e6f2a212]{order:1;padding-left:unset;padding-right:32px}.aside-container[data-v-e6f2a212]{position:fixed;top:0;padding-top:calc(var(--vp-nav-height) + var(--vp-layout-top-height, 0px) + var(--vp-doc-top-height, 0px) + 48px);width:224px;height:100vh;overflow-x:hidden;overflow-y:auto;scrollbar-width:none}.aside-container[data-v-e6f2a212]::-webkit-scrollbar{display:none}.aside-curtain[data-v-e6f2a212]{position:fixed;bottom:0;z-index:10;width:224px;height:32px;background:linear-gradient(transparent,var(--vp-c-bg) 70%)}.aside-content[data-v-e6f2a212]{display:flex;flex-direction:column;min-height:calc(100vh - (var(--vp-nav-height) + var(--vp-layout-top-height, 0px) + 48px));padding-bottom:32px}.content[data-v-e6f2a212]{position:relative;margin:0 auto;width:100%}@media (min-width: 960px){.content[data-v-e6f2a212]{padding:0 32px 128px}}@media (min-width: 1280px){.content[data-v-e6f2a212]{order:1;margin:0;min-width:640px}}.content-container[data-v-e6f2a212]{margin:0 auto}.VPDoc.has-aside .content-container[data-v-e6f2a212]{max-width:688px}.VPButton[data-v-93dc4167]{display:inline-block;border:1px solid transparent;text-align:center;font-weight:600;white-space:nowrap;transition:color .25s,border-color .25s,background-color .25s}.VPButton[data-v-93dc4167]:active{transition:color .1s,border-color .1s,background-color .1s}.VPButton.medium[data-v-93dc4167]{border-radius:20px;padding:0 20px;line-height:38px;font-size:14px}.VPButton.big[data-v-93dc4167]{border-radius:24px;padding:0 24px;line-height:46px;font-size:16px}.VPButton.brand[data-v-93dc4167]{border-color:var(--vp-button-brand-border);color:var(--vp-button-brand-text);background-color:var(--vp-button-brand-bg)}.VPButton.brand[data-v-93dc4167]:hover{border-color:var(--vp-button-brand-hover-border);color:var(--vp-button-brand-hover-text);background-color:var(--vp-button-brand-hover-bg)}.VPButton.brand[data-v-93dc4167]:active{border-color:var(--vp-button-brand-active-border);color:var(--vp-button-brand-active-text);background-color:var(--vp-button-brand-active-bg)}.VPButton.alt[data-v-93dc4167]{border-color:var(--vp-button-alt-border);color:var(--vp-button-alt-text);background-color:var(--vp-button-alt-bg)}.VPButton.alt[data-v-93dc4167]:hover{border-color:var(--vp-button-alt-hover-border);color:var(--vp-button-alt-hover-text);background-color:var(--vp-button-alt-hover-bg)}.VPButton.alt[data-v-93dc4167]:active{border-color:var(--vp-button-alt-active-border);color:var(--vp-button-alt-active-text);background-color:var(--vp-button-alt-active-bg)}.VPButton.sponsor[data-v-93dc4167]{border-color:var(--vp-button-sponsor-border);color:var(--vp-button-sponsor-text);background-color:var(--vp-button-sponsor-bg)}.VPButton.sponsor[data-v-93dc4167]:hover{border-color:var(--vp-button-sponsor-hover-border);color:var(--vp-button-sponsor-hover-text);background-color:var(--vp-button-sponsor-hover-bg)}.VPButton.sponsor[data-v-93dc4167]:active{border-color:var(--vp-button-sponsor-active-border);color:var(--vp-button-sponsor-active-text);background-color:var(--vp-button-sponsor-active-bg)}html:not(.dark) .VPImage.dark[data-v-ab19afbb]{display:none}.dark .VPImage.light[data-v-ab19afbb]{display:none}.VPHero[data-v-dd8814ff]{margin-top:calc((var(--vp-nav-height) + var(--vp-layout-top-height, 0px)) * -1);padding:calc(var(--vp-nav-height) + var(--vp-layout-top-height, 0px) + 48px) 24px 48px}@media (min-width: 640px){.VPHero[data-v-dd8814ff]{padding:calc(var(--vp-nav-height) + var(--vp-layout-top-height, 0px) + 80px) 48px 64px}}@media (min-width: 960px){.VPHero[data-v-dd8814ff]{padding:calc(var(--vp-nav-height) + var(--vp-layout-top-height, 0px) + 80px) 64px 64px}}.container[data-v-dd8814ff]{display:flex;flex-direction:column;margin:0 auto;max-width:1152px}@media (min-width: 960px){.container[data-v-dd8814ff]{flex-direction:row}}.main[data-v-dd8814ff]{position:relative;z-index:10;order:2;flex-grow:1;flex-shrink:0}.VPHero.has-image .container[data-v-dd8814ff]{text-align:center}@media (min-width: 960px){.VPHero.has-image .container[data-v-dd8814ff]{text-align:left}}@media (min-width: 960px){.main[data-v-dd8814ff]{order:1;width:calc((100% / 3) * 2)}.VPHero.has-image .main[data-v-dd8814ff]{max-width:592px}}.heading[data-v-dd8814ff]{display:flex;flex-direction:column}.name[data-v-dd8814ff],.text[data-v-dd8814ff]{width:fit-content;max-width:392px;letter-spacing:-.4px;line-height:40px;font-size:32px;font-weight:700;white-space:pre-wrap}.VPHero.has-image .name[data-v-dd8814ff],.VPHero.has-image .text[data-v-dd8814ff]{margin:0 auto}.name[data-v-dd8814ff]{color:var(--vp-home-hero-name-color)}.clip[data-v-dd8814ff]{background:var(--vp-home-hero-name-background);-webkit-background-clip:text;background-clip:text;-webkit-text-fill-color:var(--vp-home-hero-name-color)}@media (min-width: 640px){.name[data-v-dd8814ff],.text[data-v-dd8814ff]{max-width:576px;line-height:56px;font-size:48px}}@media (min-width: 960px){.name[data-v-dd8814ff],.text[data-v-dd8814ff]{line-height:64px;font-size:56px}.VPHero.has-image .name[data-v-dd8814ff],.VPHero.has-image .text[data-v-dd8814ff]{margin:0}}.tagline[data-v-dd8814ff]{padding-top:8px;max-width:392px;line-height:28px;font-size:18px;font-weight:500;white-space:pre-wrap;color:var(--vp-c-text-2)}.VPHero.has-image .tagline[data-v-dd8814ff]{margin:0 auto}@media (min-width: 640px){.tagline[data-v-dd8814ff]{padding-top:12px;max-width:576px;line-height:32px;font-size:20px}}@media (min-width: 960px){.tagline[data-v-dd8814ff]{line-height:36px;font-size:24px}.VPHero.has-image .tagline[data-v-dd8814ff]{margin:0}}.actions[data-v-dd8814ff]{display:flex;flex-wrap:wrap;margin:-6px;padding-top:24px}.VPHero.has-image .actions[data-v-dd8814ff]{justify-content:center}@media (min-width: 640px){.actions[data-v-dd8814ff]{padding-top:32px}}@media (min-width: 960px){.VPHero.has-image .actions[data-v-dd8814ff]{justify-content:flex-start}}.action[data-v-dd8814ff]{flex-shrink:0;padding:6px}.image[data-v-dd8814ff]{order:1;margin:-76px -24px -48px}@media (min-width: 640px){.image[data-v-dd8814ff]{margin:-108px -24px -48px}}@media (min-width: 960px){.image[data-v-dd8814ff]{flex-grow:1;order:2;margin:0;min-height:100%}}.image-container[data-v-dd8814ff]{position:relative;margin:0 auto;width:320px;height:320px}@media (min-width: 640px){.image-container[data-v-dd8814ff]{width:392px;height:392px}}@media (min-width: 960px){.image-container[data-v-dd8814ff]{display:flex;justify-content:center;align-items:center;width:100%;height:100%;transform:translate(-32px,-32px)}}.image-bg[data-v-dd8814ff]{position:absolute;top:50%;left:50%;border-radius:50%;width:192px;height:192px;background-image:var(--vp-home-hero-image-background-image);filter:var(--vp-home-hero-image-filter);transform:translate(-50%,-50%)}@media (min-width: 640px){.image-bg[data-v-dd8814ff]{width:256px;height:256px}}@media (min-width: 960px){.image-bg[data-v-dd8814ff]{width:320px;height:320px}}[data-v-dd8814ff] .image-src{position:absolute;top:50%;left:50%;max-width:192px;max-height:192px;transform:translate(-50%,-50%)}@media (min-width: 640px){[data-v-dd8814ff] .image-src{max-width:256px;max-height:256px}}@media (min-width: 960px){[data-v-dd8814ff] .image-src{max-width:320px;max-height:320px}}.VPFeature[data-v-bd37d1a2]{display:block;border:1px solid var(--vp-c-bg-soft);border-radius:12px;height:100%;background-color:var(--vp-c-bg-soft);transition:border-color .25s,background-color .25s}.VPFeature.link[data-v-bd37d1a2]:hover{border-color:var(--vp-c-brand-1)}.box[data-v-bd37d1a2]{display:flex;flex-direction:column;padding:24px;height:100%}.box[data-v-bd37d1a2]>.VPImage{margin-bottom:20px}.icon[data-v-bd37d1a2]{display:flex;justify-content:center;align-items:center;margin-bottom:20px;border-radius:6px;background-color:var(--vp-c-default-soft);width:48px;height:48px;font-size:24px;transition:background-color .25s}.title[data-v-bd37d1a2]{line-height:24px;font-size:16px;font-weight:600}.details[data-v-bd37d1a2]{flex-grow:1;padding-top:8px;line-height:24px;font-size:14px;font-weight:500;color:var(--vp-c-text-2)}.link-text[data-v-bd37d1a2]{padding-top:8px}.link-text-value[data-v-bd37d1a2]{display:flex;align-items:center;font-size:14px;font-weight:500;color:var(--vp-c-brand-1)}.link-text-icon[data-v-bd37d1a2]{margin-left:6px}.VPFeatures[data-v-b1eea84a]{position:relative;padding:0 24px}@media (min-width: 640px){.VPFeatures[data-v-b1eea84a]{padding:0 48px}}@media (min-width: 960px){.VPFeatures[data-v-b1eea84a]{padding:0 64px}}.container[data-v-b1eea84a]{margin:0 auto;max-width:1152px}.items[data-v-b1eea84a]{display:flex;flex-wrap:wrap;margin:-8px}.item[data-v-b1eea84a]{padding:8px;width:100%}@media (min-width: 640px){.item.grid-2[data-v-b1eea84a],.item.grid-4[data-v-b1eea84a],.item.grid-6[data-v-b1eea84a]{width:50%}}@media (min-width: 768px){.item.grid-2[data-v-b1eea84a],.item.grid-4[data-v-b1eea84a]{width:50%}.item.grid-3[data-v-b1eea84a],.item.grid-6[data-v-b1eea84a]{width:calc(100% / 3)}}@media (min-width: 960px){.item.grid-4[data-v-b1eea84a]{width:25%}}.container[data-v-c141a4bd]{margin:auto;width:100%;max-width:1280px;padding:0 24px}@media (min-width: 640px){.container[data-v-c141a4bd]{padding:0 48px}}@media (min-width: 960px){.container[data-v-c141a4bd]{width:100%;padding:0 64px}}.vp-doc[data-v-c141a4bd] .VPHomeSponsors,.vp-doc[data-v-c141a4bd] .VPTeamPage{margin-left:var(--vp-offset, calc(50% - 50vw) );margin-right:var(--vp-offset, calc(50% - 50vw) )}.vp-doc[data-v-c141a4bd] .VPHomeSponsors h2{border-top:none;letter-spacing:normal}.vp-doc[data-v-c141a4bd] .VPHomeSponsors a,.vp-doc[data-v-c141a4bd] .VPTeamPage a{text-decoration:none}.VPHome[data-v-e07eaea7]{margin-bottom:96px}@media (min-width: 768px){.VPHome[data-v-e07eaea7]{margin-bottom:128px}}.VPContent[data-v-9a6c75ad]{flex-grow:1;flex-shrink:0;margin:var(--vp-layout-top-height, 0px) auto 0;width:100%}.VPContent.is-home[data-v-9a6c75ad]{width:100%;max-width:100%}.VPContent.has-sidebar[data-v-9a6c75ad]{margin:0}@media (min-width: 960px){.VPContent[data-v-9a6c75ad]{padding-top:var(--vp-nav-height)}.VPContent.has-sidebar[data-v-9a6c75ad]{margin:var(--vp-layout-top-height, 0px) 0 0;padding-left:var(--vp-sidebar-width)}}@media (min-width: 1440px){.VPContent.has-sidebar[data-v-9a6c75ad]{padding-right:calc((100vw - var(--vp-layout-max-width)) / 2);padding-left:calc((100vw - var(--vp-layout-max-width)) / 2 + var(--vp-sidebar-width))}}.VPFooter[data-v-566314d4]{position:relative;z-index:var(--vp-z-index-footer);border-top:1px solid var(--vp-c-gutter);padding:32px 24px;background-color:var(--vp-c-bg)}.VPFooter.has-sidebar[data-v-566314d4]{display:none}.VPFooter[data-v-566314d4] a{text-decoration-line:underline;text-underline-offset:2px;transition:color .25s}.VPFooter[data-v-566314d4] a:hover{color:var(--vp-c-text-1)}@media (min-width: 768px){.VPFooter[data-v-566314d4]{padding:32px}}.container[data-v-566314d4]{margin:0 auto;max-width:var(--vp-layout-max-width);text-align:center}.message[data-v-566314d4],.copyright[data-v-566314d4]{line-height:24px;font-size:14px;font-weight:500;color:var(--vp-c-text-2)}.VPLocalNavOutlineDropdown[data-v-6b867909]{padding:12px 20px 11px}@media (min-width: 960px){.VPLocalNavOutlineDropdown[data-v-6b867909]{padding:12px 36px 11px}}.VPLocalNavOutlineDropdown button[data-v-6b867909]{display:block;font-size:12px;font-weight:500;line-height:24px;color:var(--vp-c-text-2);transition:color .5s;position:relative}.VPLocalNavOutlineDropdown button[data-v-6b867909]:hover{color:var(--vp-c-text-1);transition:color .25s}.VPLocalNavOutlineDropdown button.open[data-v-6b867909]{color:var(--vp-c-text-1)}.icon[data-v-6b867909]{display:inline-block;vertical-align:middle;margin-left:2px;font-size:14px;transform:rotate(0);transition:transform .25s}@media (min-width: 960px){.VPLocalNavOutlineDropdown button[data-v-6b867909]{font-size:14px}.icon[data-v-6b867909]{font-size:16px}}.open>.icon[data-v-6b867909]{transform:rotate(90deg)}.items[data-v-6b867909]{position:absolute;top:40px;right:16px;left:16px;display:grid;gap:1px;border:1px solid var(--vp-c-border);border-radius:8px;background-color:var(--vp-c-gutter);max-height:calc(var(--vp-vh, 100vh) - 86px);overflow:hidden auto;box-shadow:var(--vp-shadow-3)}@media (min-width: 960px){.items[data-v-6b867909]{right:auto;left:calc(var(--vp-sidebar-width) + 32px);width:320px}}.header[data-v-6b867909]{background-color:var(--vp-c-bg-soft)}.top-link[data-v-6b867909]{display:block;padding:0 16px;line-height:48px;font-size:14px;font-weight:500;color:var(--vp-c-brand-1)}.outline[data-v-6b867909]{padding:8px 0;background-color:var(--vp-c-bg-soft)}.flyout-enter-active[data-v-6b867909]{transition:all .2s ease-out}.flyout-leave-active[data-v-6b867909]{transition:all .15s ease-in}.flyout-enter-from[data-v-6b867909],.flyout-leave-to[data-v-6b867909]{opacity:0;transform:translateY(-16px)}.VPLocalNav[data-v-2488c25a]{position:sticky;top:0;left:0;z-index:var(--vp-z-index-local-nav);border-bottom:1px solid var(--vp-c-gutter);padding-top:var(--vp-layout-top-height, 0px);width:100%;background-color:var(--vp-local-nav-bg-color)}.VPLocalNav.fixed[data-v-2488c25a]{position:fixed}@media (min-width: 960px){.VPLocalNav[data-v-2488c25a]{top:var(--vp-nav-height)}.VPLocalNav.has-sidebar[data-v-2488c25a]{padding-left:var(--vp-sidebar-width)}.VPLocalNav.empty[data-v-2488c25a]{display:none}}@media (min-width: 1280px){.VPLocalNav[data-v-2488c25a]{display:none}}@media (min-width: 1440px){.VPLocalNav.has-sidebar[data-v-2488c25a]{padding-left:calc((100vw - var(--vp-layout-max-width)) / 2 + var(--vp-sidebar-width))}}.container[data-v-2488c25a]{display:flex;justify-content:space-between;align-items:center}.menu[data-v-2488c25a]{display:flex;align-items:center;padding:12px 24px 11px;line-height:24px;font-size:12px;font-weight:500;color:var(--vp-c-text-2);transition:color .5s}.menu[data-v-2488c25a]:hover{color:var(--vp-c-text-1);transition:color .25s}@media (min-width: 768px){.menu[data-v-2488c25a]{padding:0 32px}}@media (min-width: 960px){.menu[data-v-2488c25a]{display:none}}.menu-icon[data-v-2488c25a]{margin-right:8px;font-size:14px}.VPOutlineDropdown[data-v-2488c25a]{padding:12px 24px 11px}@media (min-width: 768px){.VPOutlineDropdown[data-v-2488c25a]{padding:12px 32px 11px}}.VPSwitch[data-v-b4ccac88]{position:relative;border-radius:11px;display:block;width:40px;height:22px;flex-shrink:0;border:1px solid var(--vp-input-border-color);background-color:var(--vp-input-switch-bg-color);transition:border-color .25s!important}.VPSwitch[data-v-b4ccac88]:hover{border-color:var(--vp-c-brand-1)}.check[data-v-b4ccac88]{position:absolute;top:1px;left:1px;width:18px;height:18px;border-radius:50%;background-color:var(--vp-c-neutral-inverse);box-shadow:var(--vp-shadow-1);transition:transform .25s!important}.icon[data-v-b4ccac88]{position:relative;display:block;width:18px;height:18px;border-radius:50%;overflow:hidden}.icon[data-v-b4ccac88] [class^=vpi-]{position:absolute;top:3px;left:3px;width:12px;height:12px;color:var(--vp-c-text-2)}.dark .icon[data-v-b4ccac88] [class^=vpi-]{color:var(--vp-c-text-1);transition:opacity .25s!important}.sun[data-v-be9742d9]{opacity:1}.moon[data-v-be9742d9],.dark .sun[data-v-be9742d9]{opacity:0}.dark .moon[data-v-be9742d9]{opacity:1}.dark .VPSwitchAppearance[data-v-be9742d9] .check{transform:translate(18px)}.VPNavBarAppearance[data-v-3f90c1a5]{display:none}@media (min-width: 1280px){.VPNavBarAppearance[data-v-3f90c1a5]{display:flex;align-items:center}}.VPMenuGroup+.VPMenuLink[data-v-7eeeb2dc]{margin:12px -12px 0;border-top:1px solid var(--vp-c-divider);padding:12px 12px 0}.link[data-v-7eeeb2dc]{display:block;border-radius:6px;padding:0 12px;line-height:32px;font-size:14px;font-weight:500;color:var(--vp-c-text-1);white-space:nowrap;transition:background-color .25s,color .25s}.link[data-v-7eeeb2dc]:hover{color:var(--vp-c-brand-1);background-color:var(--vp-c-default-soft)}.link.active[data-v-7eeeb2dc]{color:var(--vp-c-brand-1)}.VPMenuGroup[data-v-a6b0397c]{margin:12px -12px 0;border-top:1px solid var(--vp-c-divider);padding:12px 12px 0}.VPMenuGroup[data-v-a6b0397c]:first-child{margin-top:0;border-top:0;padding-top:0}.VPMenuGroup+.VPMenuGroup[data-v-a6b0397c]{margin-top:12px;border-top:1px solid var(--vp-c-divider)}.title[data-v-a6b0397c]{padding:0 12px;line-height:32px;font-size:14px;font-weight:600;color:var(--vp-c-text-2);white-space:nowrap;transition:color .25s}.VPMenu[data-v-20ed86d6]{border-radius:12px;padding:12px;min-width:128px;border:1px solid var(--vp-c-divider);background-color:var(--vp-c-bg-elv);box-shadow:var(--vp-shadow-3);transition:background-color .5s;max-height:calc(100vh - var(--vp-nav-height));overflow-y:auto}.VPMenu[data-v-20ed86d6] .group{margin:0 -12px;padding:0 12px 12px}.VPMenu[data-v-20ed86d6] .group+.group{border-top:1px solid var(--vp-c-divider);padding:11px 12px 12px}.VPMenu[data-v-20ed86d6] .group:last-child{padding-bottom:0}.VPMenu[data-v-20ed86d6] .group+.item{border-top:1px solid var(--vp-c-divider);padding:11px 16px 0}.VPMenu[data-v-20ed86d6] .item{padding:0 16px;white-space:nowrap}.VPMenu[data-v-20ed86d6] .label{flex-grow:1;line-height:28px;font-size:12px;font-weight:500;color:var(--vp-c-text-2);transition:color .5s}.VPMenu[data-v-20ed86d6] .action{padding-left:24px}.VPFlyout[data-v-bfe7971f]{position:relative}.VPFlyout[data-v-bfe7971f]:hover{color:var(--vp-c-brand-1);transition:color .25s}.VPFlyout:hover .text[data-v-bfe7971f]{color:var(--vp-c-text-2)}.VPFlyout:hover .icon[data-v-bfe7971f]{fill:var(--vp-c-text-2)}.VPFlyout.active .text[data-v-bfe7971f]{color:var(--vp-c-brand-1)}.VPFlyout.active:hover .text[data-v-bfe7971f]{color:var(--vp-c-brand-2)}.button[aria-expanded=false]+.menu[data-v-bfe7971f]{opacity:0;visibility:hidden;transform:translateY(0)}.VPFlyout:hover .menu[data-v-bfe7971f],.button[aria-expanded=true]+.menu[data-v-bfe7971f]{opacity:1;visibility:visible;transform:translateY(0)}.button[data-v-bfe7971f]{display:flex;align-items:center;padding:0 12px;height:var(--vp-nav-height);color:var(--vp-c-text-1);transition:color .5s}.text[data-v-bfe7971f]{display:flex;align-items:center;line-height:var(--vp-nav-height);font-size:14px;font-weight:500;color:var(--vp-c-text-1);transition:color .25s}.option-icon[data-v-bfe7971f]{margin-right:0;font-size:16px}.text-icon[data-v-bfe7971f]{margin-left:4px;font-size:14px}.icon[data-v-bfe7971f]{font-size:20px;transition:fill .25s}.menu[data-v-bfe7971f]{position:absolute;top:calc(var(--vp-nav-height) / 2 + 20px);right:0;opacity:0;visibility:hidden;transition:opacity .25s,visibility .25s,transform .25s}.VPSocialLink[data-v-60a9a2d3]{display:flex;justify-content:center;align-items:center;width:36px;height:36px;color:var(--vp-c-text-2);transition:color .5s}.VPSocialLink[data-v-60a9a2d3]:hover{color:var(--vp-c-text-1);transition:color .25s}.VPSocialLink[data-v-60a9a2d3]>svg,.VPSocialLink[data-v-60a9a2d3]>[class^=vpi-social-]{width:20px;height:20px;fill:currentColor}.VPSocialLinks[data-v-e71e869c]{display:flex;justify-content:center}.VPNavBarExtra[data-v-f953d92f]{display:none;margin-right:-12px}@media (min-width: 768px){.VPNavBarExtra[data-v-f953d92f]{display:block}}@media (min-width: 1280px){.VPNavBarExtra[data-v-f953d92f]{display:none}}.trans-title[data-v-f953d92f]{padding:0 24px 0 12px;line-height:32px;font-size:14px;font-weight:700;color:var(--vp-c-text-1)}.item.appearance[data-v-f953d92f],.item.social-links[data-v-f953d92f]{display:flex;align-items:center;padding:0 12px}.item.appearance[data-v-f953d92f]{min-width:176px}.appearance-action[data-v-f953d92f]{margin-right:-2px}.social-links-list[data-v-f953d92f]{margin:-4px -8px}.VPNavBarHamburger[data-v-6bee1efd]{display:flex;justify-content:center;align-items:center;width:48px;height:var(--vp-nav-height)}@media (min-width: 768px){.VPNavBarHamburger[data-v-6bee1efd]{display:none}}.container[data-v-6bee1efd]{position:relative;width:16px;height:14px;overflow:hidden}.VPNavBarHamburger:hover .top[data-v-6bee1efd]{top:0;left:0;transform:translate(4px)}.VPNavBarHamburger:hover .middle[data-v-6bee1efd]{top:6px;left:0;transform:translate(0)}.VPNavBarHamburger:hover .bottom[data-v-6bee1efd]{top:12px;left:0;transform:translate(8px)}.VPNavBarHamburger.active .top[data-v-6bee1efd]{top:6px;transform:translate(0) rotate(225deg)}.VPNavBarHamburger.active .middle[data-v-6bee1efd]{top:6px;transform:translate(16px)}.VPNavBarHamburger.active .bottom[data-v-6bee1efd]{top:6px;transform:translate(0) rotate(135deg)}.VPNavBarHamburger.active:hover .top[data-v-6bee1efd],.VPNavBarHamburger.active:hover .middle[data-v-6bee1efd],.VPNavBarHamburger.active:hover .bottom[data-v-6bee1efd]{background-color:var(--vp-c-text-2);transition:top .25s,background-color .25s,transform .25s}.top[data-v-6bee1efd],.middle[data-v-6bee1efd],.bottom[data-v-6bee1efd]{position:absolute;width:16px;height:2px;background-color:var(--vp-c-text-1);transition:top .25s,background-color .5s,transform .25s}.top[data-v-6bee1efd]{top:0;left:0;transform:translate(0)}.middle[data-v-6bee1efd]{top:6px;left:0;transform:translate(8px)}.bottom[data-v-6bee1efd]{top:12px;left:0;transform:translate(4px)}.VPNavBarMenuLink[data-v-815115f5]{display:flex;align-items:center;padding:0 12px;line-height:var(--vp-nav-height);font-size:14px;font-weight:500;color:var(--vp-c-text-1);transition:color .25s}.VPNavBarMenuLink.active[data-v-815115f5],.VPNavBarMenuLink[data-v-815115f5]:hover{color:var(--vp-c-brand-1)}.VPNavBarMenu[data-v-afb2845e]{display:none}@media (min-width: 768px){.VPNavBarMenu[data-v-afb2845e]{display:flex}}/*! @docsearch/css 3.8.2 | MIT License | © Algolia, Inc. and contributors | https://docsearch.algolia.com */:root{--docsearch-primary-color:#5468ff;--docsearch-text-color:#1c1e21;--docsearch-spacing:12px;--docsearch-icon-stroke-width:1.4;--docsearch-highlight-color:var(--docsearch-primary-color);--docsearch-muted-color:#969faf;--docsearch-container-background:rgba(101,108,133,.8);--docsearch-logo-color:#5468ff;--docsearch-modal-width:560px;--docsearch-modal-height:600px;--docsearch-modal-background:#f5f6f7;--docsearch-modal-shadow:inset 1px 1px 0 0 hsla(0,0%,100%,.5),0 3px 8px 0 #555a64;--docsearch-searchbox-height:56px;--docsearch-searchbox-background:#ebedf0;--docsearch-searchbox-focus-background:#fff;--docsearch-searchbox-shadow:inset 0 0 0 2px var(--docsearch-primary-color);--docsearch-hit-height:56px;--docsearch-hit-color:#444950;--docsearch-hit-active-color:#fff;--docsearch-hit-background:#fff;--docsearch-hit-shadow:0 1px 3px 0 #d4d9e1;--docsearch-key-gradient:linear-gradient(-225deg,#d5dbe4,#f8f8f8);--docsearch-key-shadow:inset 0 -2px 0 0 #cdcde6,inset 0 0 1px 1px #fff,0 1px 2px 1px rgba(30,35,90,.4);--docsearch-key-pressed-shadow:inset 0 -2px 0 0 #cdcde6,inset 0 0 1px 1px #fff,0 1px 1px 0 rgba(30,35,90,.4);--docsearch-footer-height:44px;--docsearch-footer-background:#fff;--docsearch-footer-shadow:0 -1px 0 0 #e0e3e8,0 -3px 6px 0 rgba(69,98,155,.12)}html[data-theme=dark]{--docsearch-text-color:#f5f6f7;--docsearch-container-background:rgba(9,10,17,.8);--docsearch-modal-background:#15172a;--docsearch-modal-shadow:inset 1px 1px 0 0 #2c2e40,0 3px 8px 0 #000309;--docsearch-searchbox-background:#090a11;--docsearch-searchbox-focus-background:#000;--docsearch-hit-color:#bec3c9;--docsearch-hit-shadow:none;--docsearch-hit-background:#090a11;--docsearch-key-gradient:linear-gradient(-26.5deg,#565872,#31355b);--docsearch-key-shadow:inset 0 -2px 0 0 #282d55,inset 0 0 1px 1px #51577d,0 2px 2px 0 rgba(3,4,9,.3);--docsearch-key-pressed-shadow:inset 0 -2px 0 0 #282d55,inset 0 0 1px 1px #51577d,0 1px 1px 0 #0304094d;--docsearch-footer-background:#1e2136;--docsearch-footer-shadow:inset 0 1px 0 0 rgba(73,76,106,.5),0 -4px 8px 0 rgba(0,0,0,.2);--docsearch-logo-color:#fff;--docsearch-muted-color:#7f8497}.DocSearch-Button{align-items:center;background:var(--docsearch-searchbox-background);border:0;border-radius:40px;color:var(--docsearch-muted-color);cursor:pointer;display:flex;font-weight:500;height:36px;justify-content:space-between;margin:0 0 0 16px;padding:0 8px;-webkit-user-select:none;user-select:none}.DocSearch-Button:active,.DocSearch-Button:focus,.DocSearch-Button:hover{background:var(--docsearch-searchbox-focus-background);box-shadow:var(--docsearch-searchbox-shadow);color:var(--docsearch-text-color);outline:none}.DocSearch-Button-Container{align-items:center;display:flex}.DocSearch-Search-Icon{stroke-width:1.6}.DocSearch-Button .DocSearch-Search-Icon{color:var(--docsearch-text-color)}.DocSearch-Button-Placeholder{font-size:1rem;padding:0 12px 0 6px}.DocSearch-Button-Keys{display:flex;min-width:calc(40px + .8em)}.DocSearch-Button-Key{align-items:center;background:var(--docsearch-key-gradient);border:0;border-radius:3px;box-shadow:var(--docsearch-key-shadow);color:var(--docsearch-muted-color);display:flex;height:18px;justify-content:center;margin-right:.4em;padding:0 0 2px;position:relative;top:-1px;width:20px}.DocSearch-Button-Key--pressed{box-shadow:var(--docsearch-key-pressed-shadow);transform:translate3d(0,1px,0)}@media (max-width:768px){.DocSearch-Button-Keys,.DocSearch-Button-Placeholder{display:none}}.DocSearch--active{overflow:hidden!important}.DocSearch-Container,.DocSearch-Container *{box-sizing:border-box}.DocSearch-Container{background-color:var(--docsearch-container-background);height:100vh;left:0;position:fixed;top:0;width:100vw;z-index:200}.DocSearch-Container a{text-decoration:none}.DocSearch-Link{-webkit-appearance:none;-moz-appearance:none;appearance:none;background:none;border:0;color:var(--docsearch-highlight-color);cursor:pointer;font:inherit;margin:0;padding:0}.DocSearch-Modal{background:var(--docsearch-modal-background);border-radius:6px;box-shadow:var(--docsearch-modal-shadow);flex-direction:column;margin:60px auto auto;max-width:var(--docsearch-modal-width);position:relative}.DocSearch-SearchBar{display:flex;padding:var(--docsearch-spacing) var(--docsearch-spacing) 0}.DocSearch-Form{align-items:center;background:var(--docsearch-searchbox-focus-background);border-radius:4px;box-shadow:var(--docsearch-searchbox-shadow);display:flex;height:var(--docsearch-searchbox-height);margin:0;padding:0 var(--docsearch-spacing);position:relative;width:100%}.DocSearch-Input{-webkit-appearance:none;-moz-appearance:none;appearance:none;background:transparent;border:0;color:var(--docsearch-text-color);flex:1;font:inherit;font-size:1.2em;height:100%;outline:none;padding:0 0 0 8px;width:80%}.DocSearch-Input::placeholder{color:var(--docsearch-muted-color);opacity:1}.DocSearch-Input::-webkit-search-cancel-button,.DocSearch-Input::-webkit-search-decoration,.DocSearch-Input::-webkit-search-results-button,.DocSearch-Input::-webkit-search-results-decoration{display:none}.DocSearch-LoadingIndicator,.DocSearch-MagnifierLabel,.DocSearch-Reset{margin:0;padding:0}.DocSearch-MagnifierLabel,.DocSearch-Reset{align-items:center;color:var(--docsearch-highlight-color);display:flex;justify-content:center}.DocSearch-Container--Stalled .DocSearch-MagnifierLabel,.DocSearch-LoadingIndicator{display:none}.DocSearch-Container--Stalled .DocSearch-LoadingIndicator{align-items:center;color:var(--docsearch-highlight-color);display:flex;justify-content:center}@media screen and (prefers-reduced-motion:reduce){.DocSearch-Reset{animation:none;-webkit-appearance:none;-moz-appearance:none;appearance:none;background:none;border:0;border-radius:50%;color:var(--docsearch-icon-color);cursor:pointer;right:0;stroke-width:var(--docsearch-icon-stroke-width)}}.DocSearch-Reset{animation:fade-in .1s ease-in forwards;-webkit-appearance:none;-moz-appearance:none;appearance:none;background:none;border:0;border-radius:50%;color:var(--docsearch-icon-color);cursor:pointer;padding:2px;right:0;stroke-width:var(--docsearch-icon-stroke-width)}.DocSearch-Reset[hidden]{display:none}.DocSearch-Reset:hover{color:var(--docsearch-highlight-color)}.DocSearch-LoadingIndicator svg,.DocSearch-MagnifierLabel svg{height:24px;width:24px}.DocSearch-Cancel{display:none}.DocSearch-Dropdown{max-height:calc(var(--docsearch-modal-height) - var(--docsearch-searchbox-height) - var(--docsearch-spacing) - var(--docsearch-footer-height));min-height:var(--docsearch-spacing);overflow-y:auto;overflow-y:overlay;padding:0 var(--docsearch-spacing);scrollbar-color:var(--docsearch-muted-color) var(--docsearch-modal-background);scrollbar-width:thin}.DocSearch-Dropdown::-webkit-scrollbar{width:12px}.DocSearch-Dropdown::-webkit-scrollbar-track{background:transparent}.DocSearch-Dropdown::-webkit-scrollbar-thumb{background-color:var(--docsearch-muted-color);border:3px solid var(--docsearch-modal-background);border-radius:20px}.DocSearch-Dropdown ul{list-style:none;margin:0;padding:0}.DocSearch-Label{font-size:.75em;line-height:1.6em}.DocSearch-Help,.DocSearch-Label{color:var(--docsearch-muted-color)}.DocSearch-Help{font-size:.9em;margin:0;-webkit-user-select:none;user-select:none}.DocSearch-Title{font-size:1.2em}.DocSearch-Logo a{display:flex}.DocSearch-Logo svg{color:var(--docsearch-logo-color);margin-left:8px}.DocSearch-Hits:last-of-type{margin-bottom:24px}.DocSearch-Hits mark{background:none;color:var(--docsearch-highlight-color)}.DocSearch-HitsFooter{color:var(--docsearch-muted-color);display:flex;font-size:.85em;justify-content:center;margin-bottom:var(--docsearch-spacing);padding:var(--docsearch-spacing)}.DocSearch-HitsFooter a{border-bottom:1px solid;color:inherit}.DocSearch-Hit{border-radius:4px;display:flex;padding-bottom:4px;position:relative}@media screen and (prefers-reduced-motion:reduce){.DocSearch-Hit--deleting{transition:none}}.DocSearch-Hit--deleting{opacity:0;transition:all .25s linear}@media screen and (prefers-reduced-motion:reduce){.DocSearch-Hit--favoriting{transition:none}}.DocSearch-Hit--favoriting{transform:scale(0);transform-origin:top center;transition:all .25s linear;transition-delay:.25s}.DocSearch-Hit a{background:var(--docsearch-hit-background);border-radius:4px;box-shadow:var(--docsearch-hit-shadow);display:block;padding-left:var(--docsearch-spacing);width:100%}.DocSearch-Hit-source{background:var(--docsearch-modal-background);color:var(--docsearch-highlight-color);font-size:.85em;font-weight:600;line-height:32px;margin:0 -4px;padding:8px 4px 0;position:sticky;top:0;z-index:10}.DocSearch-Hit-Tree{color:var(--docsearch-muted-color);height:var(--docsearch-hit-height);opacity:.5;stroke-width:var(--docsearch-icon-stroke-width);width:24px}.DocSearch-Hit[aria-selected=true] a{background-color:var(--docsearch-highlight-color)}.DocSearch-Hit[aria-selected=true] mark{text-decoration:underline}.DocSearch-Hit-Container{align-items:center;color:var(--docsearch-hit-color);display:flex;flex-direction:row;height:var(--docsearch-hit-height);padding:0 var(--docsearch-spacing) 0 0}.DocSearch-Hit-icon{height:20px;width:20px}.DocSearch-Hit-action,.DocSearch-Hit-icon{color:var(--docsearch-muted-color);stroke-width:var(--docsearch-icon-stroke-width)}.DocSearch-Hit-action{align-items:center;display:flex;height:22px;width:22px}.DocSearch-Hit-action svg{display:block;height:18px;width:18px}.DocSearch-Hit-action+.DocSearch-Hit-action{margin-left:6px}.DocSearch-Hit-action-button{-webkit-appearance:none;-moz-appearance:none;appearance:none;background:none;border:0;border-radius:50%;color:inherit;cursor:pointer;padding:2px}svg.DocSearch-Hit-Select-Icon{display:none}.DocSearch-Hit[aria-selected=true] .DocSearch-Hit-Select-Icon{display:block}.DocSearch-Hit-action-button:focus,.DocSearch-Hit-action-button:hover{background:#0003;transition:background-color .1s ease-in}@media screen and (prefers-reduced-motion:reduce){.DocSearch-Hit-action-button:focus,.DocSearch-Hit-action-button:hover{transition:none}}.DocSearch-Hit-action-button:focus path,.DocSearch-Hit-action-button:hover path{fill:#fff}.DocSearch-Hit-content-wrapper{display:flex;flex:1 1 auto;flex-direction:column;font-weight:500;justify-content:center;line-height:1.2em;margin:0 8px;overflow-x:hidden;position:relative;text-overflow:ellipsis;white-space:nowrap;width:80%}.DocSearch-Hit-title{font-size:.9em}.DocSearch-Hit-path{color:var(--docsearch-muted-color);font-size:.75em}.DocSearch-Hit[aria-selected=true] .DocSearch-Hit-Tree,.DocSearch-Hit[aria-selected=true] .DocSearch-Hit-action,.DocSearch-Hit[aria-selected=true] .DocSearch-Hit-icon,.DocSearch-Hit[aria-selected=true] .DocSearch-Hit-path,.DocSearch-Hit[aria-selected=true] .DocSearch-Hit-text,.DocSearch-Hit[aria-selected=true] .DocSearch-Hit-title,.DocSearch-Hit[aria-selected=true] mark{color:var(--docsearch-hit-active-color)!important}@media screen and (prefers-reduced-motion:reduce){.DocSearch-Hit-action-button:focus,.DocSearch-Hit-action-button:hover{background:#0003;transition:none}}.DocSearch-ErrorScreen,.DocSearch-NoResults,.DocSearch-StartScreen{font-size:.9em;margin:0 auto;padding:36px 0;text-align:center;width:80%}.DocSearch-Screen-Icon{color:var(--docsearch-muted-color);padding-bottom:12px}.DocSearch-NoResults-Prefill-List{display:inline-block;padding-bottom:24px;text-align:left}.DocSearch-NoResults-Prefill-List ul{display:inline-block;padding:8px 0 0}.DocSearch-NoResults-Prefill-List li{list-style-position:inside;list-style-type:"» "}.DocSearch-Prefill{-webkit-appearance:none;-moz-appearance:none;appearance:none;background:none;border:0;border-radius:1em;color:var(--docsearch-highlight-color);cursor:pointer;display:inline-block;font-size:1em;font-weight:700;padding:0}.DocSearch-Prefill:focus,.DocSearch-Prefill:hover{outline:none;text-decoration:underline}.DocSearch-Footer{align-items:center;background:var(--docsearch-footer-background);border-radius:0 0 8px 8px;box-shadow:var(--docsearch-footer-shadow);display:flex;flex-direction:row-reverse;flex-shrink:0;height:var(--docsearch-footer-height);justify-content:space-between;padding:0 var(--docsearch-spacing);position:relative;-webkit-user-select:none;user-select:none;width:100%;z-index:300}.DocSearch-Commands{color:var(--docsearch-muted-color);display:flex;list-style:none;margin:0;padding:0}.DocSearch-Commands li{align-items:center;display:flex}.DocSearch-Commands li:not(:last-of-type){margin-right:.8em}.DocSearch-Commands-Key{align-items:center;background:var(--docsearch-key-gradient);border:0;border-radius:2px;box-shadow:var(--docsearch-key-shadow);color:var(--docsearch-muted-color);display:flex;height:18px;justify-content:center;margin-right:.4em;padding:0 0 1px;width:20px}.DocSearch-VisuallyHiddenForAccessibility{clip:rect(0 0 0 0);clip-path:inset(50%);height:1px;overflow:hidden;position:absolute;white-space:nowrap;width:1px}@media (max-width:768px){:root{--docsearch-spacing:10px;--docsearch-footer-height:40px}.DocSearch-Dropdown{height:100%}.DocSearch-Container{height:100vh;height:-webkit-fill-available;height:calc(var(--docsearch-vh, 1vh)*100);position:absolute}.DocSearch-Footer{border-radius:0;bottom:0;position:absolute}.DocSearch-Hit-content-wrapper{display:flex;position:relative;width:80%}.DocSearch-Modal{border-radius:0;box-shadow:none;height:100vh;height:-webkit-fill-available;height:calc(var(--docsearch-vh, 1vh)*100);margin:0;max-width:100%;width:100%}.DocSearch-Dropdown{max-height:calc(var(--docsearch-vh, 1vh)*100 - var(--docsearch-searchbox-height) - var(--docsearch-spacing) - var(--docsearch-footer-height))}.DocSearch-Cancel{-webkit-appearance:none;-moz-appearance:none;appearance:none;background:none;border:0;color:var(--docsearch-highlight-color);cursor:pointer;display:inline-block;flex:none;font:inherit;font-size:1em;font-weight:500;margin-left:var(--docsearch-spacing);outline:none;overflow:hidden;padding:0;-webkit-user-select:none;user-select:none;white-space:nowrap}.DocSearch-Commands,.DocSearch-Hit-Tree{display:none}}@keyframes fade-in{0%{opacity:0}to{opacity:1}}[class*=DocSearch]{--docsearch-primary-color: var(--vp-c-brand-1);--docsearch-highlight-color: var(--docsearch-primary-color);--docsearch-text-color: var(--vp-c-text-1);--docsearch-muted-color: var(--vp-c-text-2);--docsearch-searchbox-shadow: none;--docsearch-searchbox-background: transparent;--docsearch-searchbox-focus-background: transparent;--docsearch-key-gradient: transparent;--docsearch-key-shadow: none;--docsearch-modal-background: var(--vp-c-bg-soft);--docsearch-footer-background: var(--vp-c-bg)}.dark [class*=DocSearch]{--docsearch-modal-shadow: none;--docsearch-footer-shadow: none;--docsearch-logo-color: var(--vp-c-text-2);--docsearch-hit-background: var(--vp-c-default-soft);--docsearch-hit-color: var(--vp-c-text-2);--docsearch-hit-shadow: none}.DocSearch-Button{display:flex;justify-content:center;align-items:center;margin:0;padding:0;width:48px;height:55px;background:transparent;transition:border-color .25s}.DocSearch-Button:hover{background:transparent}.DocSearch-Button:focus{outline:1px dotted;outline:5px auto -webkit-focus-ring-color}.DocSearch-Button-Key--pressed{transform:none;box-shadow:none}.DocSearch-Button:focus:not(:focus-visible){outline:none!important}@media (min-width: 768px){.DocSearch-Button{justify-content:flex-start;border:1px solid transparent;border-radius:8px;padding:0 10px 0 12px;width:100%;height:40px;background-color:var(--vp-c-bg-alt)}.DocSearch-Button:hover{border-color:var(--vp-c-brand-1);background:var(--vp-c-bg-alt)}}.DocSearch-Button .DocSearch-Button-Container{display:flex;align-items:center}.DocSearch-Button .DocSearch-Search-Icon{position:relative;width:16px;height:16px;color:var(--vp-c-text-1);fill:currentColor;transition:color .5s}.DocSearch-Button:hover .DocSearch-Search-Icon{color:var(--vp-c-text-1)}@media (min-width: 768px){.DocSearch-Button .DocSearch-Search-Icon{top:1px;margin-right:8px;width:14px;height:14px;color:var(--vp-c-text-2)}}.DocSearch-Button .DocSearch-Button-Placeholder{display:none;margin-top:2px;padding:0 16px 0 0;font-size:13px;font-weight:500;color:var(--vp-c-text-2);transition:color .5s}.DocSearch-Button:hover .DocSearch-Button-Placeholder{color:var(--vp-c-text-1)}@media (min-width: 768px){.DocSearch-Button .DocSearch-Button-Placeholder{display:inline-block}}.DocSearch-Button .DocSearch-Button-Keys{direction:ltr;display:none;min-width:auto}@media (min-width: 768px){.DocSearch-Button .DocSearch-Button-Keys{display:flex;align-items:center}}.DocSearch-Button .DocSearch-Button-Key{display:block;margin:2px 0 0;border:1px solid var(--vp-c-divider);border-right:none;border-radius:4px 0 0 4px;padding-left:6px;min-width:0;width:auto;height:22px;line-height:22px;font-family:var(--vp-font-family-base);font-size:12px;font-weight:500;transition:color .5s,border-color .5s}.DocSearch-Button .DocSearch-Button-Key+.DocSearch-Button-Key{border-right:1px solid var(--vp-c-divider);border-left:none;border-radius:0 4px 4px 0;padding-left:2px;padding-right:6px}.DocSearch-Button .DocSearch-Button-Key:first-child{font-size:0!important}.DocSearch-Button .DocSearch-Button-Key:first-child:after{content:"Ctrl";font-size:12px;letter-spacing:normal;color:var(--docsearch-muted-color)}.mac .DocSearch-Button .DocSearch-Button-Key:first-child:after{content:"⌘"}.DocSearch-Button .DocSearch-Button-Key:first-child>*{display:none}.DocSearch-Search-Icon{--icon: url("data:image/svg+xml,%3Csvg xmlns='http://www.w3.org/2000/svg' stroke-width='1.6' viewBox='0 0 20 20'%3E%3Cpath fill='none' stroke='currentColor' stroke-linecap='round' stroke-linejoin='round' d='m14.386 14.386 4.088 4.088-4.088-4.088A7.533 7.533 0 1 1 3.733 3.733a7.533 7.533 0 0 1 10.653 10.653z'/%3E%3C/svg%3E")}.VPNavBarSearch{display:flex;align-items:center}@media (min-width: 768px){.VPNavBarSearch{flex-grow:1;padding-left:24px}}@media (min-width: 960px){.VPNavBarSearch{padding-left:32px}}.dark .DocSearch-Footer{border-top:1px solid var(--vp-c-divider)}.DocSearch-Form{border:1px solid var(--vp-c-brand-1);background-color:var(--vp-c-white)}.dark .DocSearch-Form{background-color:var(--vp-c-default-soft)}.DocSearch-Screen-Icon>svg{margin:auto}.VPNavBarSocialLinks[data-v-ef6192dc]{display:none}@media (min-width: 1280px){.VPNavBarSocialLinks[data-v-ef6192dc]{display:flex;align-items:center}}.title[data-v-9f43907a]{display:flex;align-items:center;border-bottom:1px solid transparent;width:100%;height:var(--vp-nav-height);font-size:16px;font-weight:600;color:var(--vp-c-text-1);transition:opacity .25s}@media (min-width: 960px){.title[data-v-9f43907a]{flex-shrink:0}.VPNavBarTitle.has-sidebar .title[data-v-9f43907a]{border-bottom-color:var(--vp-c-divider)}}[data-v-9f43907a] .logo{margin-right:8px;height:var(--vp-nav-logo-height)}.VPNavBarTranslations[data-v-acee064b]{display:none}@media (min-width: 1280px){.VPNavBarTranslations[data-v-acee064b]{display:flex;align-items:center}}.title[data-v-acee064b]{padding:0 24px 0 12px;line-height:32px;font-size:14px;font-weight:700;color:var(--vp-c-text-1)}.VPNavBar[data-v-9fd4d1dd]{position:relative;height:var(--vp-nav-height);pointer-events:none;white-space:nowrap;transition:background-color .25s}.VPNavBar.screen-open[data-v-9fd4d1dd]{transition:none;background-color:var(--vp-nav-bg-color);border-bottom:1px solid var(--vp-c-divider)}.VPNavBar[data-v-9fd4d1dd]:not(.home){background-color:var(--vp-nav-bg-color)}@media (min-width: 960px){.VPNavBar[data-v-9fd4d1dd]:not(.home){background-color:transparent}.VPNavBar[data-v-9fd4d1dd]:not(.has-sidebar):not(.home.top){background-color:var(--vp-nav-bg-color)}}.wrapper[data-v-9fd4d1dd]{padding:0 8px 0 24px}@media (min-width: 768px){.wrapper[data-v-9fd4d1dd]{padding:0 32px}}@media (min-width: 960px){.VPNavBar.has-sidebar .wrapper[data-v-9fd4d1dd]{padding:0}}.container[data-v-9fd4d1dd]{display:flex;justify-content:space-between;margin:0 auto;max-width:calc(var(--vp-layout-max-width) - 64px);height:var(--vp-nav-height);pointer-events:none}.container>.title[data-v-9fd4d1dd],.container>.content[data-v-9fd4d1dd]{pointer-events:none}.container[data-v-9fd4d1dd] *{pointer-events:auto}@media (min-width: 960px){.VPNavBar.has-sidebar .container[data-v-9fd4d1dd]{max-width:100%}}.title[data-v-9fd4d1dd]{flex-shrink:0;height:calc(var(--vp-nav-height) - 1px);transition:background-color .5s}@media (min-width: 960px){.VPNavBar.has-sidebar .title[data-v-9fd4d1dd]{position:absolute;top:0;left:0;z-index:2;padding:0 32px;width:var(--vp-sidebar-width);height:var(--vp-nav-height);background-color:transparent}}@media (min-width: 1440px){.VPNavBar.has-sidebar .title[data-v-9fd4d1dd]{padding-left:max(32px,calc((100% - (var(--vp-layout-max-width) - 64px)) / 2));width:calc((100% - (var(--vp-layout-max-width) - 64px)) / 2 + var(--vp-sidebar-width) - 32px)}}.content[data-v-9fd4d1dd]{flex-grow:1}@media (min-width: 960px){.VPNavBar.has-sidebar .content[data-v-9fd4d1dd]{position:relative;z-index:1;padding-right:32px;padding-left:var(--vp-sidebar-width)}}@media (min-width: 1440px){.VPNavBar.has-sidebar .content[data-v-9fd4d1dd]{padding-right:calc((100vw - var(--vp-layout-max-width)) / 2 + 32px);padding-left:calc((100vw - var(--vp-layout-max-width)) / 2 + var(--vp-sidebar-width))}}.content-body[data-v-9fd4d1dd]{display:flex;justify-content:flex-end;align-items:center;height:var(--vp-nav-height);transition:background-color .5s}@media (min-width: 960px){.VPNavBar:not(.home.top) .content-body[data-v-9fd4d1dd]{position:relative;background-color:var(--vp-nav-bg-color)}.VPNavBar:not(.has-sidebar):not(.home.top) .content-body[data-v-9fd4d1dd]{background-color:transparent}}@media (max-width: 767px){.content-body[data-v-9fd4d1dd]{column-gap:.5rem}}.menu+.translations[data-v-9fd4d1dd]:before,.menu+.appearance[data-v-9fd4d1dd]:before,.menu+.social-links[data-v-9fd4d1dd]:before,.translations+.appearance[data-v-9fd4d1dd]:before,.appearance+.social-links[data-v-9fd4d1dd]:before{margin-right:8px;margin-left:8px;width:1px;height:24px;background-color:var(--vp-c-divider);content:""}.menu+.appearance[data-v-9fd4d1dd]:before,.translations+.appearance[data-v-9fd4d1dd]:before{margin-right:16px}.appearance+.social-links[data-v-9fd4d1dd]:before{margin-left:16px}.social-links[data-v-9fd4d1dd]{margin-right:-8px}.divider[data-v-9fd4d1dd]{width:100%;height:1px}@media (min-width: 960px){.VPNavBar.has-sidebar .divider[data-v-9fd4d1dd]{padding-left:var(--vp-sidebar-width)}}@media (min-width: 1440px){.VPNavBar.has-sidebar .divider[data-v-9fd4d1dd]{padding-left:calc((100vw - var(--vp-layout-max-width)) / 2 + var(--vp-sidebar-width))}}.divider-line[data-v-9fd4d1dd]{width:100%;height:1px;transition:background-color .5s}.VPNavBar:not(.home) .divider-line[data-v-9fd4d1dd]{background-color:var(--vp-c-gutter)}@media (min-width: 960px){.VPNavBar:not(.home.top) .divider-line[data-v-9fd4d1dd]{background-color:var(--vp-c-gutter)}.VPNavBar:not(.has-sidebar):not(.home.top) .divider[data-v-9fd4d1dd]{background-color:var(--vp-c-gutter)}}.VPNavScreenAppearance[data-v-a3e2920d]{display:flex;justify-content:space-between;align-items:center;border-radius:8px;padding:12px 14px 12px 16px;background-color:var(--vp-c-bg-soft)}.text[data-v-a3e2920d]{line-height:24px;font-size:12px;font-weight:500;color:var(--vp-c-text-2)}.VPNavScreenMenuLink[data-v-fa963d97]{display:block;border-bottom:1px solid var(--vp-c-divider);padding:12px 0 11px;line-height:24px;font-size:14px;font-weight:500;color:var(--vp-c-text-1);transition:border-color .25s,color .25s}.VPNavScreenMenuLink[data-v-fa963d97]:hover{color:var(--vp-c-brand-1)}.VPNavScreenMenuGroupLink[data-v-e04f3e85]{display:block;margin-left:12px;line-height:32px;font-size:14px;font-weight:400;color:var(--vp-c-text-1);transition:color .25s}.VPNavScreenMenuGroupLink[data-v-e04f3e85]:hover{color:var(--vp-c-brand-1)}.VPNavScreenMenuGroupSection[data-v-f60dbfa7]{display:block}.title[data-v-f60dbfa7]{line-height:32px;font-size:13px;font-weight:700;color:var(--vp-c-text-2);transition:color .25s}.VPNavScreenMenuGroup[data-v-d99bfeec]{border-bottom:1px solid var(--vp-c-divider);height:48px;overflow:hidden;transition:border-color .5s}.VPNavScreenMenuGroup .items[data-v-d99bfeec]{visibility:hidden}.VPNavScreenMenuGroup.open .items[data-v-d99bfeec]{visibility:visible}.VPNavScreenMenuGroup.open[data-v-d99bfeec]{padding-bottom:10px;height:auto}.VPNavScreenMenuGroup.open .button[data-v-d99bfeec]{padding-bottom:6px;color:var(--vp-c-brand-1)}.VPNavScreenMenuGroup.open .button-icon[data-v-d99bfeec]{transform:rotate(45deg)}.button[data-v-d99bfeec]{display:flex;justify-content:space-between;align-items:center;padding:12px 4px 11px 0;width:100%;line-height:24px;font-size:14px;font-weight:500;color:var(--vp-c-text-1);transition:color .25s}.button[data-v-d99bfeec]:hover{color:var(--vp-c-brand-1)}.button-icon[data-v-d99bfeec]{transition:transform .25s}.group[data-v-d99bfeec]:first-child{padding-top:0}.group+.group[data-v-d99bfeec],.group+.item[data-v-d99bfeec]{padding-top:4px}.VPNavScreenTranslations[data-v-516e4bc3]{height:24px;overflow:hidden}.VPNavScreenTranslations.open[data-v-516e4bc3]{height:auto}.title[data-v-516e4bc3]{display:flex;align-items:center;font-size:14px;font-weight:500;color:var(--vp-c-text-1)}.icon[data-v-516e4bc3]{font-size:16px}.icon.lang[data-v-516e4bc3]{margin-right:8px}.icon.chevron[data-v-516e4bc3]{margin-left:4px}.list[data-v-516e4bc3]{padding:4px 0 0 24px}.link[data-v-516e4bc3]{line-height:32px;font-size:13px;color:var(--vp-c-text-1)}.VPNavScreen[data-v-2dd6d0c7]{position:fixed;top:calc(var(--vp-nav-height) + var(--vp-layout-top-height, 0px));right:0;bottom:0;left:0;padding:0 32px;width:100%;background-color:var(--vp-nav-screen-bg-color);overflow-y:auto;transition:background-color .25s;pointer-events:auto}.VPNavScreen.fade-enter-active[data-v-2dd6d0c7],.VPNavScreen.fade-leave-active[data-v-2dd6d0c7]{transition:opacity .25s}.VPNavScreen.fade-enter-active .container[data-v-2dd6d0c7],.VPNavScreen.fade-leave-active .container[data-v-2dd6d0c7]{transition:transform .25s ease}.VPNavScreen.fade-enter-from[data-v-2dd6d0c7],.VPNavScreen.fade-leave-to[data-v-2dd6d0c7]{opacity:0}.VPNavScreen.fade-enter-from .container[data-v-2dd6d0c7],.VPNavScreen.fade-leave-to .container[data-v-2dd6d0c7]{transform:translateY(-8px)}@media (min-width: 768px){.VPNavScreen[data-v-2dd6d0c7]{display:none}}.container[data-v-2dd6d0c7]{margin:0 auto;padding:24px 0 96px;max-width:288px}.menu+.translations[data-v-2dd6d0c7],.menu+.appearance[data-v-2dd6d0c7],.translations+.appearance[data-v-2dd6d0c7]{margin-top:24px}.menu+.social-links[data-v-2dd6d0c7]{margin-top:16px}.appearance+.social-links[data-v-2dd6d0c7]{margin-top:16px}.VPNav[data-v-7ad780c2]{position:relative;top:var(--vp-layout-top-height, 0px);left:0;z-index:var(--vp-z-index-nav);width:100%;pointer-events:none;transition:background-color .5s}@media (min-width: 960px){.VPNav[data-v-7ad780c2]{position:fixed}}.VPSidebarItem.level-0[data-v-0009425e]{padding-bottom:24px}.VPSidebarItem.collapsed.level-0[data-v-0009425e]{padding-bottom:10px}.item[data-v-0009425e]{position:relative;display:flex;width:100%}.VPSidebarItem.collapsible>.item[data-v-0009425e]{cursor:pointer}.indicator[data-v-0009425e]{position:absolute;top:6px;bottom:6px;left:-17px;width:2px;border-radius:2px;transition:background-color .25s}.VPSidebarItem.level-2.is-active>.item>.indicator[data-v-0009425e],.VPSidebarItem.level-3.is-active>.item>.indicator[data-v-0009425e],.VPSidebarItem.level-4.is-active>.item>.indicator[data-v-0009425e],.VPSidebarItem.level-5.is-active>.item>.indicator[data-v-0009425e]{background-color:var(--vp-c-brand-1)}.link[data-v-0009425e]{display:flex;align-items:center;flex-grow:1}.text[data-v-0009425e]{flex-grow:1;padding:4px 0;line-height:24px;font-size:14px;transition:color .25s}.VPSidebarItem.level-0 .text[data-v-0009425e]{font-weight:700;color:var(--vp-c-text-1)}.VPSidebarItem.level-1 .text[data-v-0009425e],.VPSidebarItem.level-2 .text[data-v-0009425e],.VPSidebarItem.level-3 .text[data-v-0009425e],.VPSidebarItem.level-4 .text[data-v-0009425e],.VPSidebarItem.level-5 .text[data-v-0009425e]{font-weight:500;color:var(--vp-c-text-2)}.VPSidebarItem.level-0.is-link>.item>.link:hover .text[data-v-0009425e],.VPSidebarItem.level-1.is-link>.item>.link:hover .text[data-v-0009425e],.VPSidebarItem.level-2.is-link>.item>.link:hover .text[data-v-0009425e],.VPSidebarItem.level-3.is-link>.item>.link:hover .text[data-v-0009425e],.VPSidebarItem.level-4.is-link>.item>.link:hover .text[data-v-0009425e],.VPSidebarItem.level-5.is-link>.item>.link:hover .text[data-v-0009425e]{color:var(--vp-c-brand-1)}.VPSidebarItem.level-0.has-active>.item>.text[data-v-0009425e],.VPSidebarItem.level-1.has-active>.item>.text[data-v-0009425e],.VPSidebarItem.level-2.has-active>.item>.text[data-v-0009425e],.VPSidebarItem.level-3.has-active>.item>.text[data-v-0009425e],.VPSidebarItem.level-4.has-active>.item>.text[data-v-0009425e],.VPSidebarItem.level-5.has-active>.item>.text[data-v-0009425e],.VPSidebarItem.level-0.has-active>.item>.link>.text[data-v-0009425e],.VPSidebarItem.level-1.has-active>.item>.link>.text[data-v-0009425e],.VPSidebarItem.level-2.has-active>.item>.link>.text[data-v-0009425e],.VPSidebarItem.level-3.has-active>.item>.link>.text[data-v-0009425e],.VPSidebarItem.level-4.has-active>.item>.link>.text[data-v-0009425e],.VPSidebarItem.level-5.has-active>.item>.link>.text[data-v-0009425e]{color:var(--vp-c-text-1)}.VPSidebarItem.level-0.is-active>.item .link>.text[data-v-0009425e],.VPSidebarItem.level-1.is-active>.item .link>.text[data-v-0009425e],.VPSidebarItem.level-2.is-active>.item .link>.text[data-v-0009425e],.VPSidebarItem.level-3.is-active>.item .link>.text[data-v-0009425e],.VPSidebarItem.level-4.is-active>.item .link>.text[data-v-0009425e],.VPSidebarItem.level-5.is-active>.item .link>.text[data-v-0009425e]{color:var(--vp-c-brand-1)}.caret[data-v-0009425e]{display:flex;justify-content:center;align-items:center;margin-right:-7px;width:32px;height:32px;color:var(--vp-c-text-3);cursor:pointer;transition:color .25s;flex-shrink:0}.item:hover .caret[data-v-0009425e]{color:var(--vp-c-text-2)}.item:hover .caret[data-v-0009425e]:hover{color:var(--vp-c-text-1)}.caret-icon[data-v-0009425e]{font-size:18px;transform:rotate(90deg);transition:transform .25s}.VPSidebarItem.collapsed .caret-icon[data-v-0009425e]{transform:rotate(0)}.VPSidebarItem.level-1 .items[data-v-0009425e],.VPSidebarItem.level-2 .items[data-v-0009425e],.VPSidebarItem.level-3 .items[data-v-0009425e],.VPSidebarItem.level-4 .items[data-v-0009425e],.VPSidebarItem.level-5 .items[data-v-0009425e]{border-left:1px solid var(--vp-c-divider);padding-left:16px}.VPSidebarItem.collapsed .items[data-v-0009425e]{display:none}.no-transition[data-v-51288d80] .caret-icon{transition:none}.group+.group[data-v-51288d80]{border-top:1px solid var(--vp-c-divider);padding-top:10px}@media (min-width: 960px){.group[data-v-51288d80]{padding-top:10px;width:calc(var(--vp-sidebar-width) - 64px)}}.VPSidebar[data-v-42c4c606]{position:fixed;top:var(--vp-layout-top-height, 0px);bottom:0;left:0;z-index:var(--vp-z-index-sidebar);padding:32px 32px 96px;width:calc(100vw - 64px);max-width:320px;background-color:var(--vp-sidebar-bg-color);opacity:0;box-shadow:var(--vp-c-shadow-3);overflow-x:hidden;overflow-y:auto;transform:translate(-100%);transition:opacity .5s,transform .25s ease;overscroll-behavior:contain}.VPSidebar.open[data-v-42c4c606]{opacity:1;visibility:visible;transform:translate(0);transition:opacity .25s,transform .5s cubic-bezier(.19,1,.22,1)}.dark .VPSidebar[data-v-42c4c606]{box-shadow:var(--vp-shadow-1)}@media (min-width: 960px){.VPSidebar[data-v-42c4c606]{padding-top:var(--vp-nav-height);width:var(--vp-sidebar-width);max-width:100%;background-color:var(--vp-sidebar-bg-color);opacity:1;visibility:visible;box-shadow:none;transform:translate(0)}}@media (min-width: 1440px){.VPSidebar[data-v-42c4c606]{padding-left:max(32px,calc((100% - (var(--vp-layout-max-width) - 64px)) / 2));width:calc((100% - (var(--vp-layout-max-width) - 64px)) / 2 + var(--vp-sidebar-width) - 32px)}}@media (min-width: 960px){.curtain[data-v-42c4c606]{position:sticky;top:-64px;left:0;z-index:1;margin-top:calc(var(--vp-nav-height) * -1);margin-right:-32px;margin-left:-32px;height:var(--vp-nav-height);background-color:var(--vp-sidebar-bg-color)}}.nav[data-v-42c4c606]{outline:0}.VPSkipLink[data-v-fcbfc0e0]{top:8px;left:8px;padding:8px 16px;z-index:999;border-radius:8px;font-size:12px;font-weight:700;text-decoration:none;color:var(--vp-c-brand-1);box-shadow:var(--vp-shadow-3);background-color:var(--vp-c-bg)}.VPSkipLink[data-v-fcbfc0e0]:focus{height:auto;width:auto;clip:auto;clip-path:none}@media (min-width: 1280px){.VPSkipLink[data-v-fcbfc0e0]{top:14px;left:16px}}.Layout[data-v-d8b57b2d]{display:flex;flex-direction:column;min-height:100vh}.VPHomeSponsors[data-v-3dc26e1d]{border-top:1px solid var(--vp-c-gutter);padding-top:88px!important}.VPHomeSponsors[data-v-3dc26e1d]{margin:96px 0}@media (min-width: 768px){.VPHomeSponsors[data-v-3dc26e1d]{margin:128px 0}}.VPHomeSponsors[data-v-3dc26e1d]{padding:0 24px}@media (min-width: 768px){.VPHomeSponsors[data-v-3dc26e1d]{padding:0 48px}}@media (min-width: 960px){.VPHomeSponsors[data-v-3dc26e1d]{padding:0 64px}}.container[data-v-3dc26e1d]{margin:0 auto;max-width:1152px}.love[data-v-3dc26e1d]{margin:0 auto;width:fit-content;font-size:28px;color:var(--vp-c-text-3)}.icon[data-v-3dc26e1d]{display:inline-block}.message[data-v-3dc26e1d]{margin:0 auto;padding-top:10px;max-width:320px;text-align:center;line-height:24px;font-size:16px;font-weight:500;color:var(--vp-c-text-2)}.sponsors[data-v-3dc26e1d]{padding-top:32px}.action[data-v-3dc26e1d]{padding-top:40px;text-align:center}.VPTeamMembersItem[data-v-acff304e]{display:flex;flex-direction:column;gap:2px;border-radius:12px;width:100%;height:100%;overflow:hidden}.VPTeamMembersItem.small .profile[data-v-acff304e]{padding:32px}.VPTeamMembersItem.small .data[data-v-acff304e]{padding-top:20px}.VPTeamMembersItem.small .avatar[data-v-acff304e]{width:64px;height:64px}.VPTeamMembersItem.small .name[data-v-acff304e]{line-height:24px;font-size:16px}.VPTeamMembersItem.small .affiliation[data-v-acff304e]{padding-top:4px;line-height:20px;font-size:14px}.VPTeamMembersItem.small .desc[data-v-acff304e]{padding-top:12px;line-height:20px;font-size:14px}.VPTeamMembersItem.small .links[data-v-acff304e]{margin:0 -16px -20px;padding:10px 0 0}.VPTeamMembersItem.medium .profile[data-v-acff304e]{padding:48px 32px}.VPTeamMembersItem.medium .data[data-v-acff304e]{padding-top:24px;text-align:center}.VPTeamMembersItem.medium .avatar[data-v-acff304e]{width:96px;height:96px}.VPTeamMembersItem.medium .name[data-v-acff304e]{letter-spacing:.15px;line-height:28px;font-size:20px}.VPTeamMembersItem.medium .affiliation[data-v-acff304e]{padding-top:4px;font-size:16px}.VPTeamMembersItem.medium .desc[data-v-acff304e]{padding-top:16px;max-width:288px;font-size:16px}.VPTeamMembersItem.medium .links[data-v-acff304e]{margin:0 -16px -12px;padding:16px 12px 0}.profile[data-v-acff304e]{flex-grow:1;background-color:var(--vp-c-bg-soft)}.data[data-v-acff304e]{text-align:center}.avatar[data-v-acff304e]{position:relative;flex-shrink:0;margin:0 auto;border-radius:50%;box-shadow:var(--vp-shadow-3)}.avatar-img[data-v-acff304e]{position:absolute;top:0;right:0;bottom:0;left:0;border-radius:50%;object-fit:cover}.name[data-v-acff304e]{margin:0;font-weight:600}.affiliation[data-v-acff304e]{margin:0;font-weight:500;color:var(--vp-c-text-2)}.org.link[data-v-acff304e]{color:var(--vp-c-text-2);transition:color .25s}.org.link[data-v-acff304e]:hover{color:var(--vp-c-brand-1)}.desc[data-v-acff304e]{margin:0 auto}.desc[data-v-acff304e] a{font-weight:500;color:var(--vp-c-brand-1);text-decoration-style:dotted;transition:color .25s}.links[data-v-acff304e]{display:flex;justify-content:center;height:56px}.sp-link[data-v-acff304e]{display:flex;justify-content:center;align-items:center;text-align:center;padding:16px;font-size:14px;font-weight:500;color:var(--vp-c-sponsor);background-color:var(--vp-c-bg-soft);transition:color .25s,background-color .25s}.sp .sp-link.link[data-v-acff304e]:hover,.sp .sp-link.link[data-v-acff304e]:focus{outline:none;color:var(--vp-c-white);background-color:var(--vp-c-sponsor)}.sp-icon[data-v-acff304e]{margin-right:8px;font-size:16px}.VPTeamMembers.small .container[data-v-bf782009]{grid-template-columns:repeat(auto-fit,minmax(224px,1fr))}.VPTeamMembers.small.count-1 .container[data-v-bf782009]{max-width:276px}.VPTeamMembers.small.count-2 .container[data-v-bf782009]{max-width:576px}.VPTeamMembers.small.count-3 .container[data-v-bf782009]{max-width:876px}.VPTeamMembers.medium .container[data-v-bf782009]{grid-template-columns:repeat(auto-fit,minmax(256px,1fr))}@media (min-width: 375px){.VPTeamMembers.medium .container[data-v-bf782009]{grid-template-columns:repeat(auto-fit,minmax(288px,1fr))}}.VPTeamMembers.medium.count-1 .container[data-v-bf782009]{max-width:368px}.VPTeamMembers.medium.count-2 .container[data-v-bf782009]{max-width:760px}.container[data-v-bf782009]{display:grid;gap:24px;margin:0 auto;max-width:1152px}.VPTeamPage[data-v-a5329171]{margin:96px 0}@media (min-width: 768px){.VPTeamPage[data-v-a5329171]{margin:128px 0}}.VPHome .VPTeamPageTitle[data-v-a5329171-s]{border-top:1px solid var(--vp-c-gutter);padding-top:88px!important}.VPTeamPageSection+.VPTeamPageSection[data-v-a5329171-s],.VPTeamMembers+.VPTeamPageSection[data-v-a5329171-s]{margin-top:64px}.VPTeamMembers+.VPTeamMembers[data-v-a5329171-s]{margin-top:24px}@media (min-width: 768px){.VPTeamPageTitle+.VPTeamPageSection[data-v-a5329171-s]{margin-top:16px}.VPTeamPageSection+.VPTeamPageSection[data-v-a5329171-s],.VPTeamMembers+.VPTeamPageSection[data-v-a5329171-s]{margin-top:96px}}.VPTeamMembers[data-v-a5329171-s]{padding:0 24px}@media (min-width: 768px){.VPTeamMembers[data-v-a5329171-s]{padding:0 48px}}@media (min-width: 960px){.VPTeamMembers[data-v-a5329171-s]{padding:0 64px}}.VPTeamPageSection[data-v-3bf2e850]{padding:0 32px}@media (min-width: 768px){.VPTeamPageSection[data-v-3bf2e850]{padding:0 48px}}@media (min-width: 960px){.VPTeamPageSection[data-v-3bf2e850]{padding:0 64px}}.title[data-v-3bf2e850]{position:relative;margin:0 auto;max-width:1152px;text-align:center;color:var(--vp-c-text-2)}.title-line[data-v-3bf2e850]{position:absolute;top:16px;left:0;width:100%;height:1px;background-color:var(--vp-c-divider)}.title-text[data-v-3bf2e850]{position:relative;display:inline-block;padding:0 24px;letter-spacing:0;line-height:32px;font-size:20px;font-weight:500;background-color:var(--vp-c-bg)}.lead[data-v-3bf2e850]{margin:0 auto;max-width:480px;padding-top:12px;text-align:center;line-height:24px;font-size:16px;font-weight:500;color:var(--vp-c-text-2)}.members[data-v-3bf2e850]{padding-top:40px}.VPTeamPageTitle[data-v-46c5e327]{padding:48px 32px;text-align:center}@media (min-width: 768px){.VPTeamPageTitle[data-v-46c5e327]{padding:64px 48px 48px}}@media (min-width: 960px){.VPTeamPageTitle[data-v-46c5e327]{padding:80px 64px 48px}}.title[data-v-46c5e327]{letter-spacing:0;line-height:44px;font-size:36px;font-weight:500}@media (min-width: 768px){.title[data-v-46c5e327]{letter-spacing:-.5px;line-height:56px;font-size:48px}}.lead[data-v-46c5e327]{margin:0 auto;max-width:512px;padding-top:12px;line-height:24px;font-size:16px;font-weight:500;color:var(--vp-c-text-2)}@media (min-width: 768px){.lead[data-v-46c5e327]{max-width:592px;letter-spacing:.15px;line-height:28px;font-size:20px}}:root{--vp-home-hero-name-color: transparent;--vp-home-hero-name-background: -webkit-linear-gradient(120deg, #4E004D, #C1298D)} diff --git a/.vitepress/dist/assets/tutorials.md.ATW1haj-.js b/.vitepress/dist/assets/tutorials.md.ATW1haj-.js new file mode 100644 index 0000000..d2edec8 --- /dev/null +++ b/.vitepress/dist/assets/tutorials.md.ATW1haj-.js @@ -0,0 +1 @@ +import{_ as a,o,c as n,j as t,a as i,ae as l}from"./chunks/framework.DIXg7aqb.js";const g=JSON.parse('{"title":"Tutorials","description":"","frontmatter":{},"headers":[],"relativePath":"tutorials.md","filePath":"tutorials.md"}'),s={name:"tutorials.md"};function r(m,e,c,d,u,h){return o(),n("div",null,[...e[0]||(e[0]=[t("h1",{id:"tutorials",tabindex:"-1"},[i("Tutorials "),t("a",{class:"header-anchor",href:"#tutorials","aria-label":'Permalink to "Tutorials"'},"")],-1),t("p",null,"Comprehensive tutorials and guides for using XMapTools effectively.",-1),t("h2",{id:"tutorial-1-xmaptools-for-epma-2023",tabindex:"-1"},[i("Tutorial 1: XMapTools for EPMA (2023) "),t("a",{class:"header-anchor",href:"#tutorial-1-xmaptools-for-epma-2023","aria-label":'Permalink to "Tutorial 1: XMapTools for EPMA (2023)"'},"")],-1),t("p",null,"Follow the steps in this tutorial to familiarise yourself with the EPMA data reduction procedure and discover the main features of XMapTools.",-1),t("p",null,"A video tutorial is also available on YouTube and Bilibili:",-1),t("div",{style:{position:"relative","padding-bottom":"56.25%",height:"0",overflow:"hidden","max-width":"100%",margin:"16px 0"}},[t("iframe",{style:{position:"absolute",top:"0",left:"0",width:"100%",height:"100%"},src:"https://www.youtube.com/embed/SRXio2szh3c",title:"XMapTools Course 1 – How to get started (EPMA)?",frameborder:"0",allow:"accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture; web-share",allowfullscreen:""})],-1),l('Located in China? Watch this video on Bilibili
The dataset used in this tutorial comes from the study of Duesterhoeft & Lanari (2020). It consists of EPMA maps of the sample HAL104 from West Guilford (Ontario) of the Canadian Grenville Province. It was analysed using 15 KeV accelerating voltage, 100 nA specimen current, 130 ms dwell time, 20 µm step size, ~1 µm beam size with 1000*1000 pixels. The mapping time was ~75 h.
Please do not use these data for any other purpose than this tutorial without the explicit authorisation of the author(s).
| File | Type | Comment |
|---|---|---|
| _Ce.txt | EDS map | Mostly below detection limit |
| _Cr.txt | EDS map | Mostly below detection limit |
| _La.txt | EDS map | Mostly below detection limit |
| _Ni.txt | EDS map | Below detection limit |
| _P.txt | EDS map | Overlap with Zr |
| _S.txt | EDS map | |
| Al.txt | WDS map | |
| Ca.txt | WDS map | |
| Fe.txt | WDS map | |
| K.txt | WDS map | |
| Mg.txt | WDS map | |
| Mn.txt | WDS map | |
| Na.txt | WDS map | |
| SEI.txt | Other | Secondary electron image map |
| Si.txt | WDS map | |
| Ti.txt | WDS map | |
| TOPO.txt | Other | Topographic image map (obtained by BSE) |
| Standards.txt | Standard file | File containing the spot analyses used for map standardisation |
Reference: Duesterhoeft, E. & Lanari, P. (2020). Iterative thermodynamic modelling – Part 1: A theoretical scoring technique and a computer program (BINGO-ANTIDOTE). Journal of Metamorphic Geology, 38, 527-55.
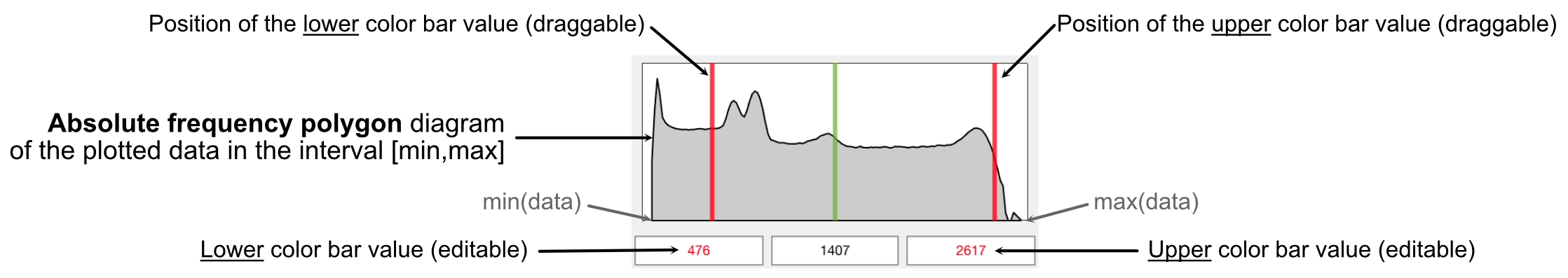
Classification is the step used to create a classified image in which each pixels are attributed to a class (mineral, glass, epoxy, etc.). In this tutorial we are using the random forest algorithm. The classification in XMapTools is supervised. You need to create a "training set" containing training data for each class.
Additional information about the classification in XMapTools can be found in : Lanari, P., Tedeschi, M., (2025). Chemical map classification in XMapTools. Applied Computing and Geosciences, 25, 100230 [Download].
The analytical standardisation consists of transforming the intensity maps of elements into weight percent maps of oxides. XMapTools 4 includes a calibration module that performs an automated calibration of all minerals. This black box algorithm works ok when all phases have spot analyses to be used as internal standard and when intensities are heigh enough. However, you need to carefully check the results of the calibration for each mineral.
The calibration method used by XMapTools is described in Lanari, P., Vho, A., Bovay, T., Airaghi, L., Centrella, S., (2019). Quantitative compositional mapping of mineral phases by electron probe micro-analyser. Geological Society of London, Special Publication, 478, 39-63 [Download].
Note: The video tutorial (see above) provides more tricks and explanations about the calibration module and how to check the quality of the calibration curves.
The method to extract local bulk composition is described in Lanari, P., & Engi, M. (2017). Local bulk composition effects on metamorphic mineral assemblages, Reviews in Mineralogy and Geochemistry, 83, 55-102 [Download].
Watch our videos to find out more and learn how to use XMapTools.
Free and versatile software solution for quantitative chemical maps analysis.

XMapTools is an advanced analysis software for quantitative chemical analysis of solids in 1D, 2D and 3D.
It provides numerical tools and packages implemented in a guided and versatile environment that allows you to explore and visualise data in your own way.
For example, XMapTools includes a wide range of data processing options including routines for classification, segmentation, calibration and visualisation via single and multi-channel maps or via binary, ternary and spider diagrams.
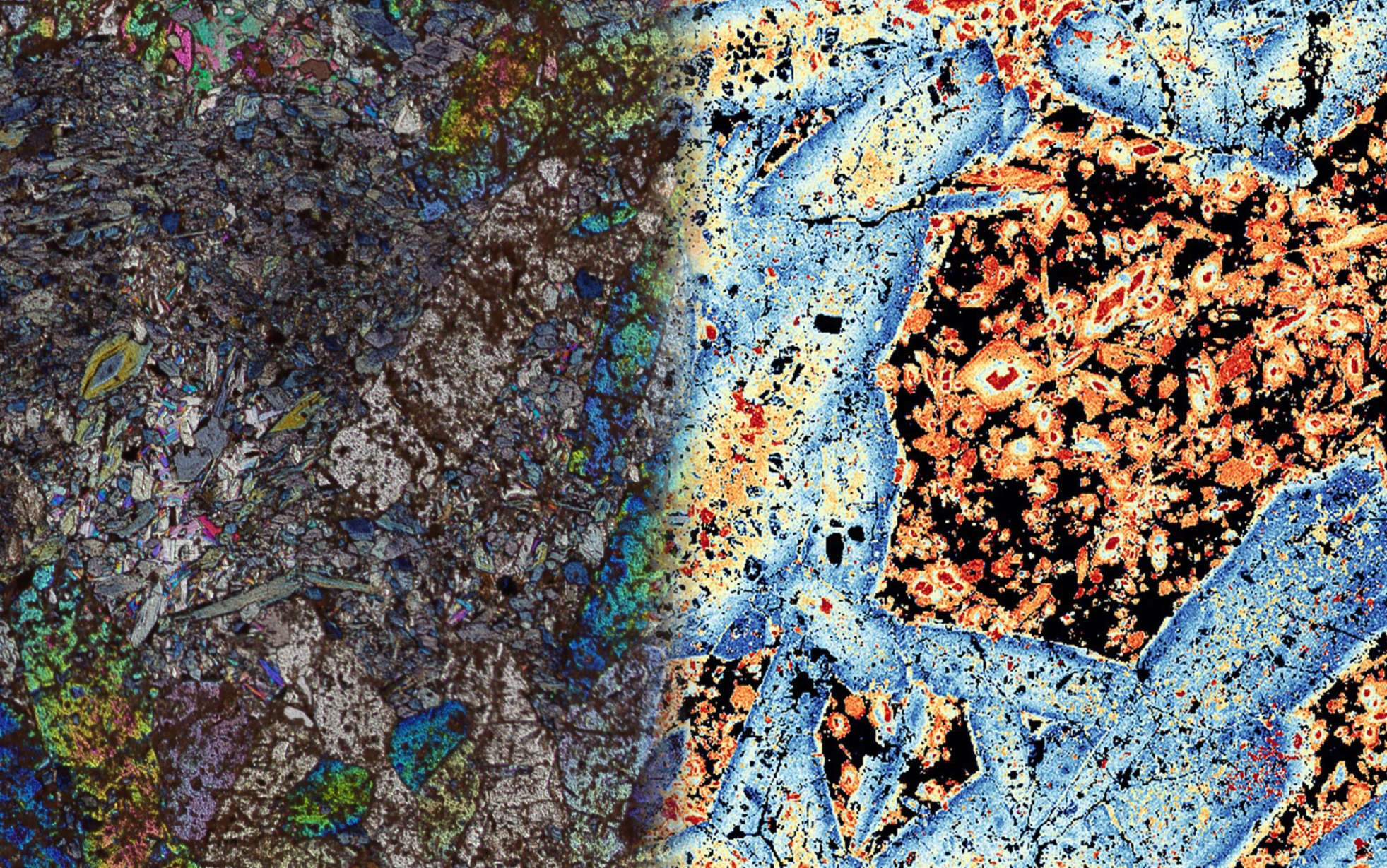


 +
+Bingo-Antidote is a petrological software originally developed by Pierre Lanari and Erik Duesterhoeft that offers an alternative modelling strategy based on iterative thermodynamic models integrated with quantitative compositional mapping. This second version, distributed as an XMapTools add-on, comes with a redesigned graphical user interface and improved features.
+
+
+## Iterative thermodynamic models
+
+Bingo-Antidote proposes a hybrid strategy combining the strengths of Gibbs energy minimisation (GEM) and inverse thermobarometry models, based on the comparison between modelled and observed mineral assemblages, modes and compositions. The overall technique relies on quantitative compositional maps obtained by electron probe microanalysis to provide a mutually consistent set of observed data such as bulk rock and mineral compositions.
+
+
+## Theriak-Domino
+
+Bingo-Antidote uses the Gibbs energy minimiser Theriak developed by Christian de Capitani. The latest version of Theriak-Domino, provided by Doug Tinkham, is required to use Bingo-Antidote.
+
+[Download Theriak-Domino from Github](https://github.com/Theriak-Domino/theriak-domino/releases/)
+
+
+
+Bingo-Antidote is a petrological software originally developed by Pierre Lanari and Erik Duesterhoeft that offers an alternative modelling strategy based on iterative thermodynamic models integrated with quantitative compositional mapping. This second version, distributed as an XMapTools add-on, comes with a redesigned graphical user interface and improved features.
+
+
+## Iterative thermodynamic models
+
+Bingo-Antidote proposes a hybrid strategy combining the strengths of Gibbs energy minimisation (GEM) and inverse thermobarometry models, based on the comparison between modelled and observed mineral assemblages, modes and compositions. The overall technique relies on quantitative compositional maps obtained by electron probe microanalysis to provide a mutually consistent set of observed data such as bulk rock and mineral compositions.
+
+
+## Theriak-Domino
+
+Bingo-Antidote uses the Gibbs energy minimiser Theriak developed by Christian de Capitani. The latest version of Theriak-Domino, provided by Doug Tinkham, is required to use Bingo-Antidote.
+
+[Download Theriak-Domino from Github](https://github.com/Theriak-Domino/theriak-domino/releases/)
+
+ +
+
+
+ +
+
+## References
+
+There are two technical papers, the first by Duesterhoeft & Lanari (2020) and the second by Lanari & Hermann (2021), which includes a proof of concept and an application example.
+
+The philosophy of iterative thermodynamic modelling behind Bingo-Antidote is also described in older publications: Lanari & Engi (2017) and Lanari & Duesterhoeft (2019). Note that the Lanari & Duesterhoeft (2019) paper gives a general overview of thermodynamic modelling and also provides an example of an application of Bingo-Antidote.
+
+- Duesterhoeft, E. & Lanari, P. (2020). Iterative thermodynamic modelling – Part 1: A theoretical scoring technique and a computer program (BINGO-ANTIDOTE). *Journal of Metamorphic Geology*, 38, 527-551. [Download pdf](https://pierrelanari.com/wp-content/uploads/2021/05/2020_DuesterhoeftLanari_ITM.pdf)
+- Lanari, P. & Hermann, J. (2021). Iterative thermodynamic modelling—Part 2: tracing equilibrium relationships between minerals in metamorphic rocks. *Journal of Metamorphic Geology*, 39, 651-674. [Download pdf](https://pierrelanari.com/wp-content/uploads/2021/07/2021_LanariHermann_JMG.pdf)
+- Lanari, P. & Duesterhoeft, E. (2019). Modelling metamorphic rocks using equilibrium thermodynamics and internally consistent databases: past achievements, problems and perspectives. *Journal of Petrology*, 60, 19-56. [Download pdf](https://pierrelanari.com/wp-content/uploads/2021/05/2019_LanariDuesterhoeft2019_JPet.pdf)
+- Lanari, P., & Engi, M. (2017). Local bulk composition effects on metamorphic mineral assemblages, *Reviews in Mineralogy and Geochemistry*, 83, 55-102. [Download pdf](https://pierrelanari.com/wp-content/uploads/2021/05/2017_LanariEngi_RiMG.pdf)
+
+
+## Bingo-Antidote Tutorial
+
+This short tutorial on Bingo-Antidote is from the Goldschmidt Workshop, which took place in Lyon on 8–9 July 2023 and was entitled 'Modern Applications of Phase Equilibrium Modelling'.
+
+### How to get started
+
+1. Download **Tutorial_Bingo-Antidote_1_Sikkim.zip** [using this link](https://xmaptools.ch/tutorial/Tutorial_Bingo-Antidote_1_Sikkim.zip).
+2. Unzip the folder in a suitable working directory (e.g. Documents/Bingo-Antidote/**Tutorial_Bingo-Antidote_1_Sikkim**/)
+3. Open XMapTools. *Note that starting XMapTools can take up to a minute; restarting the program is slightly faster*
+4. When the program is ready, a dialog box opens; pick the working directory **Tutorial_Bingo-Antidote_1_Sikkim** and click the button "Open". *You can change the working directory later, but you need to pick a directory when you start the program. A log file is automatically created in the selected working directory.*
+
+### Example 1 (TG8C-03) – Lanari & Duesterhoeft (2019), JPET
+
+1. In XMapTools, load the 'aaa4.mat' project file provided in the 'Tutorial_Bingo-Antidote_1_Sikkim' folder.
+2. Select the 'Add-ons' tab and click the 'Bingo-Antidote' button to open Bingo-Antidote.
+3. In the Bingo-Antidote launcher, you will need to provide Bingo-Antidote with the location of your Theriak setup directory. Then click 'Launch Bingo-Antidote'.
+4. Select the chemical system: SiO₂-Al₂O₃-FeO-MgO-Na₂O-CaO-K₂O (exclude TiO₂ and MnO).
+5. Select the database JUN92.bs.
+6. Select the *P*–*T* range: 600–900 °C and 0.4–1.1 GPa.
+7. Guess your initial *P*–*T* for this sample (e.g. 680 °C and 0.7 GPa).
+8. Add a ROI for Local Bulk Composition (LBC) by selecting a large area in the map (>90 surf%). Any change in the shape of the ROI updates the LBC.
+9. Activate the H₂O fluid specie and set the amount of H to 0.5 mol and then click "next".
+10. Select a solution model and add a ROI for the following phases: "biotite", "garnet", "kfeldspar", "plagioclase", "quartz" (b-quartz) and "sillimanite". Exclude the accessory minerals for BA and LBC and click "next". *Note that excluding a phase from the BA means that it will not be selected as part of the observed mineral assemblage. Excluding a phase from LBC means that the pixels of this phase will not be used when Bingo-Antidote calculate a local bulk composition.*
+11. Use Bingo to find the conditions of the peak mineral assemblage by manually changing *P* and/or *T* until Qasm is 100%.
+12. Use Recipe 1 of Antidote "Find Optimal *P*–*T*–*X*" to obtain the optimal *P*–*T*. Save the results of the last Bingo call and evaluate the quality of the model at the optimal conditions. What are the main differences between model and observations? *Note that you can check the Qcmp values for each mineral to identify those that are not modelled correctly at these conditions.*
+13. Save the current Bingo-Antidote project as BA_project_JUN92 using the button "save" available in the menu bar.
+14. Calculate the *P*–*T* maps of quality factor. *Note that the data is saved in an Antidote folder in the working directory.*
+15. In LBC, activate the option "Optimize *N*" for the fluid and set the range of H between 0.001 and 0.05 mol. *In this case the amount of H in the bulk composition will be optimised by Antidote.*
+16. In Antidote, select "Optimization of compositional and activity variables" and "Scanning *H* (fixed *PT*)" and press the button "Antidote". *Check the paper Lanari & Duesterhoeft (2019) for a description of the results of this method.*
+17. Adjust the value of H and unselect the option "Optimize *N*".
+18. Recalculate the *P*–*T* maps of quality factor and compare the results with the H₂O-saturated case.
+19. Calculate optimal P–T conditions for garnet core and rim using the recipe "single-phase thermobarometry".
+20. Use a floating window to calculate the changes between the sillimanite-absent and sillimanite-quartz domains.
+21. Run a scanning window optimization using a grid resolution of 320 pixels.
+22. Calculate a map of chemical potentials with different resolutions.
+
diff --git a/doc-addons.md b/doc-addons.md
new file mode 100644
index 0000000..cb3a6db
--- /dev/null
+++ b/doc-addons.md
@@ -0,0 +1,50 @@
+# Add-ons
+
+*Based on XMapTools 4.5 embedded documentation – Help file version 09.06.2026*
+
+---
+
+This page describes the tools available in the **Add-ons** section. Contact Pierre Lanari if you want to develop an add-on for XMapTools.
+
+## Export
+
+This module allows XMapTools data to be exported. It is available for both Map Data and Spot Data.
+
+The export format is defined by the user and can be used to export data for a specific software (e.g. MinPlot). Select the Data source, a maskfile and the type of data to be exported (e.g. Median, Mean or random selection, see Fig. 1).
+
+The BRC correction is a correction from XMapTools 4.3 (Lanari et al. 2019) that eliminates mixing pixels using the maskfile data.
+
+
+
+
+## References
+
+There are two technical papers, the first by Duesterhoeft & Lanari (2020) and the second by Lanari & Hermann (2021), which includes a proof of concept and an application example.
+
+The philosophy of iterative thermodynamic modelling behind Bingo-Antidote is also described in older publications: Lanari & Engi (2017) and Lanari & Duesterhoeft (2019). Note that the Lanari & Duesterhoeft (2019) paper gives a general overview of thermodynamic modelling and also provides an example of an application of Bingo-Antidote.
+
+- Duesterhoeft, E. & Lanari, P. (2020). Iterative thermodynamic modelling – Part 1: A theoretical scoring technique and a computer program (BINGO-ANTIDOTE). *Journal of Metamorphic Geology*, 38, 527-551. [Download pdf](https://pierrelanari.com/wp-content/uploads/2021/05/2020_DuesterhoeftLanari_ITM.pdf)
+- Lanari, P. & Hermann, J. (2021). Iterative thermodynamic modelling—Part 2: tracing equilibrium relationships between minerals in metamorphic rocks. *Journal of Metamorphic Geology*, 39, 651-674. [Download pdf](https://pierrelanari.com/wp-content/uploads/2021/07/2021_LanariHermann_JMG.pdf)
+- Lanari, P. & Duesterhoeft, E. (2019). Modelling metamorphic rocks using equilibrium thermodynamics and internally consistent databases: past achievements, problems and perspectives. *Journal of Petrology*, 60, 19-56. [Download pdf](https://pierrelanari.com/wp-content/uploads/2021/05/2019_LanariDuesterhoeft2019_JPet.pdf)
+- Lanari, P., & Engi, M. (2017). Local bulk composition effects on metamorphic mineral assemblages, *Reviews in Mineralogy and Geochemistry*, 83, 55-102. [Download pdf](https://pierrelanari.com/wp-content/uploads/2021/05/2017_LanariEngi_RiMG.pdf)
+
+
+## Bingo-Antidote Tutorial
+
+This short tutorial on Bingo-Antidote is from the Goldschmidt Workshop, which took place in Lyon on 8–9 July 2023 and was entitled 'Modern Applications of Phase Equilibrium Modelling'.
+
+### How to get started
+
+1. Download **Tutorial_Bingo-Antidote_1_Sikkim.zip** [using this link](https://xmaptools.ch/tutorial/Tutorial_Bingo-Antidote_1_Sikkim.zip).
+2. Unzip the folder in a suitable working directory (e.g. Documents/Bingo-Antidote/**Tutorial_Bingo-Antidote_1_Sikkim**/)
+3. Open XMapTools. *Note that starting XMapTools can take up to a minute; restarting the program is slightly faster*
+4. When the program is ready, a dialog box opens; pick the working directory **Tutorial_Bingo-Antidote_1_Sikkim** and click the button "Open". *You can change the working directory later, but you need to pick a directory when you start the program. A log file is automatically created in the selected working directory.*
+
+### Example 1 (TG8C-03) – Lanari & Duesterhoeft (2019), JPET
+
+1. In XMapTools, load the 'aaa4.mat' project file provided in the 'Tutorial_Bingo-Antidote_1_Sikkim' folder.
+2. Select the 'Add-ons' tab and click the 'Bingo-Antidote' button to open Bingo-Antidote.
+3. In the Bingo-Antidote launcher, you will need to provide Bingo-Antidote with the location of your Theriak setup directory. Then click 'Launch Bingo-Antidote'.
+4. Select the chemical system: SiO₂-Al₂O₃-FeO-MgO-Na₂O-CaO-K₂O (exclude TiO₂ and MnO).
+5. Select the database JUN92.bs.
+6. Select the *P*–*T* range: 600–900 °C and 0.4–1.1 GPa.
+7. Guess your initial *P*–*T* for this sample (e.g. 680 °C and 0.7 GPa).
+8. Add a ROI for Local Bulk Composition (LBC) by selecting a large area in the map (>90 surf%). Any change in the shape of the ROI updates the LBC.
+9. Activate the H₂O fluid specie and set the amount of H to 0.5 mol and then click "next".
+10. Select a solution model and add a ROI for the following phases: "biotite", "garnet", "kfeldspar", "plagioclase", "quartz" (b-quartz) and "sillimanite". Exclude the accessory minerals for BA and LBC and click "next". *Note that excluding a phase from the BA means that it will not be selected as part of the observed mineral assemblage. Excluding a phase from LBC means that the pixels of this phase will not be used when Bingo-Antidote calculate a local bulk composition.*
+11. Use Bingo to find the conditions of the peak mineral assemblage by manually changing *P* and/or *T* until Qasm is 100%.
+12. Use Recipe 1 of Antidote "Find Optimal *P*–*T*–*X*" to obtain the optimal *P*–*T*. Save the results of the last Bingo call and evaluate the quality of the model at the optimal conditions. What are the main differences between model and observations? *Note that you can check the Qcmp values for each mineral to identify those that are not modelled correctly at these conditions.*
+13. Save the current Bingo-Antidote project as BA_project_JUN92 using the button "save" available in the menu bar.
+14. Calculate the *P*–*T* maps of quality factor. *Note that the data is saved in an Antidote folder in the working directory.*
+15. In LBC, activate the option "Optimize *N*" for the fluid and set the range of H between 0.001 and 0.05 mol. *In this case the amount of H in the bulk composition will be optimised by Antidote.*
+16. In Antidote, select "Optimization of compositional and activity variables" and "Scanning *H* (fixed *PT*)" and press the button "Antidote". *Check the paper Lanari & Duesterhoeft (2019) for a description of the results of this method.*
+17. Adjust the value of H and unselect the option "Optimize *N*".
+18. Recalculate the *P*–*T* maps of quality factor and compare the results with the H₂O-saturated case.
+19. Calculate optimal P–T conditions for garnet core and rim using the recipe "single-phase thermobarometry".
+20. Use a floating window to calculate the changes between the sillimanite-absent and sillimanite-quartz domains.
+21. Run a scanning window optimization using a grid resolution of 320 pixels.
+22. Calculate a map of chemical potentials with different resolutions.
+
diff --git a/doc-addons.md b/doc-addons.md
new file mode 100644
index 0000000..cb3a6db
--- /dev/null
+++ b/doc-addons.md
@@ -0,0 +1,50 @@
+# Add-ons
+
+*Based on XMapTools 4.5 embedded documentation – Help file version 09.06.2026*
+
+---
+
+This page describes the tools available in the **Add-ons** section. Contact Pierre Lanari if you want to develop an add-on for XMapTools.
+
+## Export
+
+This module allows XMapTools data to be exported. It is available for both Map Data and Spot Data.
+
+The export format is defined by the user and can be used to export data for a specific software (e.g. MinPlot). Select the Data source, a maskfile and the type of data to be exported (e.g. Median, Mean or random selection, see Fig. 1).
+
+The BRC correction is a correction from XMapTools 4.3 (Lanari et al. 2019) that eliminates mixing pixels using the maskfile data.
+
+
+ +
+
+ +
+
+ +
+
 -
-_Figure: This is an example of a mosaic grid for 13 maps, arranged in three columns (map: Pb in counts). This allows the maps to be merged and displayed with the same colour scale._
-
-### Real mosaic
-
-The Generate Mosaic in Original Coordinate System
-
-_Figure: This is an example of a mosaic grid for 13 maps, arranged in three columns (map: Pb in counts). This allows the maps to be merged and displayed with the same colour scale._
-
-### Real mosaic
-
-The Generate Mosaic in Original Coordinate System  -
-_Figure: This is an example of a real mosaic for 3 maps, arranged using the map coordinates (map: Al2O3 in et%)._
-
-
## Importing data using the import module
-To import map data in XMapTools, select the 'Project and Import' tab and press the 'Import Maps'
-
-_Figure: This is an example of a real mosaic for 3 maps, arranged using the map coordinates (map: Al2O3 in et%)._
-
-
## Importing data using the import module
-To import map data in XMapTools, select the 'Project and Import' tab and press the 'Import Maps'
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
 +
+_Figure: This is an example of a mosaic grid for 13 maps, arranged in three columns (map: Pb in counts). This allows the maps to be merged and displayed with the same colour scale._
+
+#### Real mosaic
+
+- The **Generate Mosaic in Original Coordinate System**
+
+_Figure: This is an example of a mosaic grid for 13 maps, arranged in three columns (map: Pb in counts). This allows the maps to be merged and displayed with the same colour scale._
+
+#### Real mosaic
+
+- The **Generate Mosaic in Original Coordinate System**  +
+_Figure: This is an example of a real mosaic for 3 maps, arranged using the map coordinates (map: Al2O3 in wt%)._
+
+
+## 3. Importing Maps & Images
+
+This section describes the tools available for importing maps and other images into XMapTools. See also the dedicated [Import Maps Tool](/doc-import-maps) documentation.
+
+- The **Import (Import Maps)**
+
+_Figure: This is an example of a real mosaic for 3 maps, arranged using the map coordinates (map: Al2O3 in wt%)._
+
+
+## 3. Importing Maps & Images
+
+This section describes the tools available for importing maps and other images into XMapTools. See also the dedicated [Import Maps Tool](/doc-import-maps) documentation.
+
+- The **Import (Import Maps)**
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
+ +
+
 @@ -77,12 +77,12 @@ Classification is the step used to create a classified image in which each pixel
Additional information about the classification in XMapTools can be found in : Lanari, P., Tedeschi, M., (2025). Chemical map classification in XMapTools. _Applied Computing and Geosciences_, __25__, 100230 [Download].
1. Set the active workspace to CLASSIFY.
-2. Create a new training set. In the secondary tree menu, select the category "Training Set (Classification)" and press the "Add"
@@ -77,12 +77,12 @@ Classification is the step used to create a classified image in which each pixel
Additional information about the classification in XMapTools can be found in : Lanari, P., Tedeschi, M., (2025). Chemical map classification in XMapTools. _Applied Computing and Geosciences_, __25__, 100230 [Download].
1. Set the active workspace to CLASSIFY.
-2. Create a new training set. In the secondary tree menu, select the category "Training Set (Classification)" and press the "Add"