A high-performance FastCDC 2020 implementation written in Python + Cython
Supports Python 3.6+. Provides prebuilt wheels for Python 3.8+
Its core algorithm implementation is a direct port of the v2020 module from nlfiedler/fastcdc-rs, which means that the output of PyFastCDC completely matches the output of nlfiedler/fastcdc-rs
PyFastCDC is available on PyPI, with prebuilt wheels for many common platforms thanks to cibuildwheel
To install, you can use pip or any other Python package manager you prefer:
pip install pyfastcdcFor platforms without prebuilt wheels, a suitable build environment capable of compiling Python extension modules is required.
For example, on Debian, you might need to install gcc and python3-dev via apt
If the Cython extension fails to compile, the installation will fall back to a pure Python implementation, which is significantly slower (about 0.5% or less in memory chunking speed)
I only want to use the Cython implemetion, not the slow pure-Python one
You can set the environment variable PYFASTCDC_REQUIRE_CYTHON=true or PYFASTCDC_REQUIRE_CYTHON=1 for the pip installation command
to disable the pure-Python fallback on extension compilation error and make the installation fail hard.
Thus, after a successful installation, you will always have a working Cython extension
Example bash command using pip:
$ PYFASTCDC_REQUIRE_CYTHON=true pip install pyfastcdc
... some pip output ...
Building wheels for collected packages: pyfastcdc
Building wheel for pyfastcdc (pyproject.toml) ... error
... some pip output ...
###########################################################################################################
Failed to compile pyfastcdc Cython extension, fail hard since PYFASTCDC_REQUIRE_CYTHON is set to true
Unset PYFASTCDC_REQUIRE_CYTHON to allow pure-Python fallback if that's acceptable for your use case.
<class 'distutils.compilers.C.errors.CompileError'> command 'gcc' failed: No such file or directory
###########################################################################################################
error: command 'gcc' failed: No such file or directory
----------------------------------------
ERROR: Failed building wheel for pyfastcdc
... some pip output ...
The usage of PyFastCDC is simple:
- Construct a
FastCDCinstance with desired parameters - Call
FastCDC.cut_xxx()function to chunk your data- Call
cut_buf()to chunk in-memory data buffers - Call
cut_file()to chunk a regular file using mmap - Call
cut_stream()to chunk a custom file-like streaming object
- Call
Example:
import hashlib
from pyfastcdc import FastCDC
for chunk in FastCDC(16384).cut_file('archive.tar'):
print(chunk.offset, chunk.length, hashlib.sha256(chunk.data).hexdigest())See docstrings of exported objects in the pyfastcdc module for more API details
Please only import members from pyfastcdc in your application code and avoid importing inner modules (e.g. pyfastcdc.common) directly.
Only public APIs inside the pyfastcdc module are guaranteed to be stable across releases
from pyfastcdc import NormalizedChunking # GOOD
from pyfastcdc.common import NormalizedChunking # BAD, no API stability guaranteeWith the help of Cython, PyFastCDC can achieve near-native performance on chunking inputs
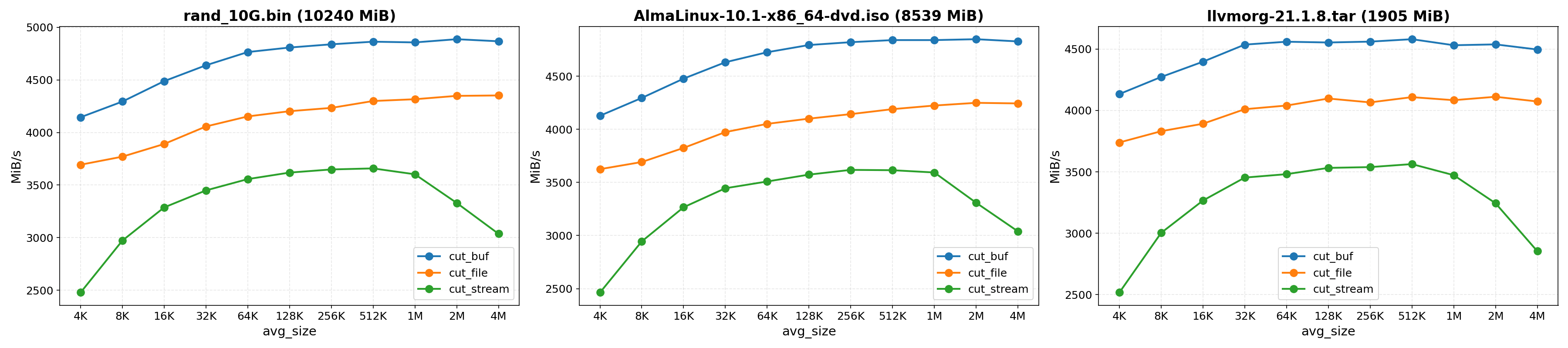
Each test was run 10 times for averaging, achieving a maximum in-memory chunking speed of about 4.8GB/s
Benchmark details
FastCDC parameters:
avg_size: Independent variablemin_size:avg_size/ 4 (default)max_size:avg_size* 4 (default)normalized_chunking: 1 (default)seed: 0 (default)
Test environment:
- PyFastCDC 0.2.0b1, precompiled wheel from Test PyPI, Cython 3.2.4
- Python 3.11.14 using docker image
python:3.11 - Ryzen 7 6800H @ 4.55GHz, NVMe SSD, Debian 13.2
Test files:
rand_10G.bin: 10GiB randomly generated binary dataAlmaLinux-10.1-x86_64-dvd.iso: the DVD ISO image of AlmaLinux 10.1llvmorg-21.1.8.tar: gzip-unzipped LLVM 21.1.8 source code
Test command:
cd scripts
python benchmark.py --test-files rand_10G.bin AlmaLinux-10.1-x86_64-dvd.iso llvmorg-21.1.8.tarThis project is inspired by iscc/fastcdc-py, but differs in the following ways:
- Based on nlfiedler/fastcdc-rs, using its FastCDC 2020 implementation aligned with the original paper, rather than the simplified ronomon implementation
- Supports multiple types of input, including in-memory data buffers, regular file using mmap, and custom streaming input
- Does not include any CLI tool. It provides only the core FastCDC functionality
Papers
- FastCDC 2016: FastCDC: A Fast and Efficient Content-Defined Chunking Approach for Data Deduplication
- FastCDC 2020: The Design of Fast Content-Defined Chunking for Data Deduplication Based Storage Systems
Other FastCDC Implementations
- HIT-HSSL/destor, the C implementation reference from the paper
- nlfiedler/fastcdc-rs, where this implementation is based on
- iscc/fastcdc-py, which provides an alternative FastCDC implementation based on ronomon/deduplication